Extraction, Isolation, and Structural Elucidation of Lignans from Natural Resources
Abstract
Lignans are a diverse class of plant-derived polyphenols that have attracted considerable interest for their broad biological activities and potential as functional ingredients in human health. Efficient extraction, purification, and accurate analysis are essential for advancing both basic research and applications of lignans. This review comprehensively summarizes recent methodological advancements in the isolation, separation, and characterization of lignans, emphasizing chromatographic and spectroscopic techniques. By integrating theoretical frameworks with practical protocols, the review provides a foundation for future studies and industrial utilization of lignans as valuable bioactive compounds.
Keywords:
Lignan, Extraction, Purification, Chromatography, Structural elucidationIntroduction
Lignans are a prominent group of plant-derived polyphenols, widely distributed across various species and organs and occurring in both free and glycosylated forms. They are found in knots, bark, roots, leaves, flowers, and seeds, attesting to their broad botanical presence and ecological roles, including defense and adaptation to environmental stress.1–3 Hundreds of distinct lignan structures have been identified, and substantial evidence links dietary lignans—especially those from whole grains and seeds—to beneficial physiological effects such as antioxidant and phytoestrogenic activity.4–6
Recent studies have highlighted the multifaceted roles of lignans in human health, demonstrating their ability to prevent cardiovascular diseases, improve blood lipid profiles, regulate glycemia, exert anti-inflammatory and antimicrobial effects, display anticancer properties, and alleviate menopausal symptoms.4,7–9 These findings underscore the nutritional and pharmacological significance of lignans and have broadened their appeal from traditional dietary contexts to pharmaceutical and nutraceutical development.
As lignans gain recognition for their roles in disease prevention and health promotion, there is increasing scientific and industrial interest in obtaining high-purity lignans for biological evaluation. Progress in lignan research and application relies not only on the capacity to secure sufficient quantities of well-defined compounds, but also on the use of robust and precise analytical methods.
Accordingly, this review presents a comprehensive overview of lignans, covering their definition, biosynthesis, and structural classification. It systematically examines state-of-the-art methodologies for extraction, separation, purification, and structural characterization, with particular emphasis on chromatographic and spectroscopic techniques that ensure analytical reliability and reproducibility. By integrating theoretical perspectives with practical strategies for lignan identification and quantification, this review aims to provide a solid scientific foundation for advancing future research and industrial applications of lignans.
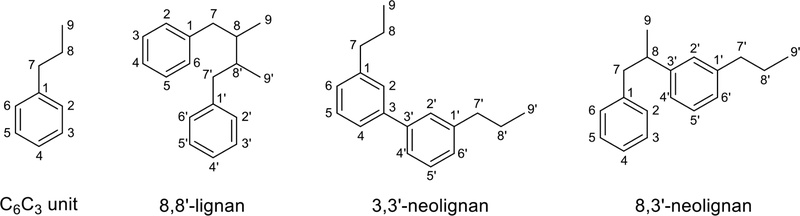
Definition – Lignans are natural polyphenolic compounds widely distributed throughout the plant kingdom. They are derived from the shikimic acid biosynthetic pathway and primarily exist as dimers resulting from the coupling of two phenylpropanoid (C6C3) derivatives in various ways. The International Union of Pure and Applied Chemistry (IUPAC) identifies lignans as dimeric C6C3 coupled motifs linked at carbon 8 and 8′. The IUPAC identifies compounds with the coupling of the two C6C3 units at positions different from C8-C8′ as neolignans (Fig. 1).10
Lignans can further be categorized into “norlignans” and “polymeric lignans” based on different polymerization positions and modes of dehydration condensation of oxygen-containing groups in their side chains.
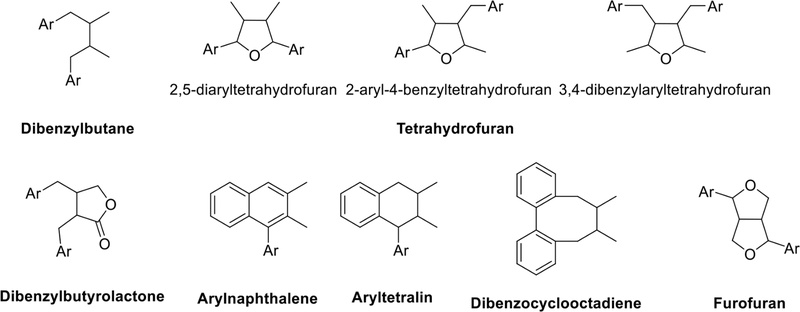
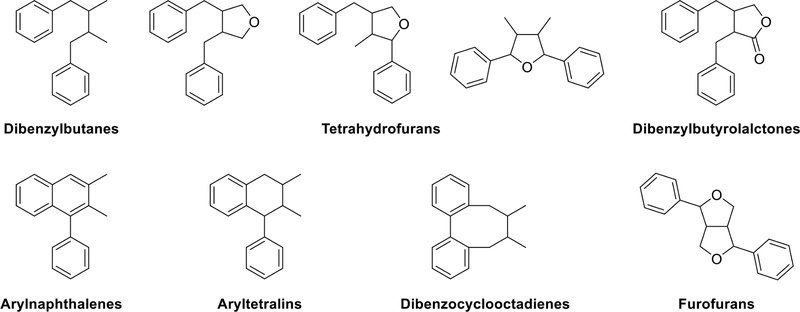
As depicted in Fig. 2, lignans are biosynthesized through the phenylpropanoid pathway, and they are classified into six groups based on their molecular structures. These are dibenzylbutane, tetrahydrofuran, dibenzylbutyrolactone, arylnaphthalene and aryltetralin, dibenzocyclooctadiene, and furofuran.
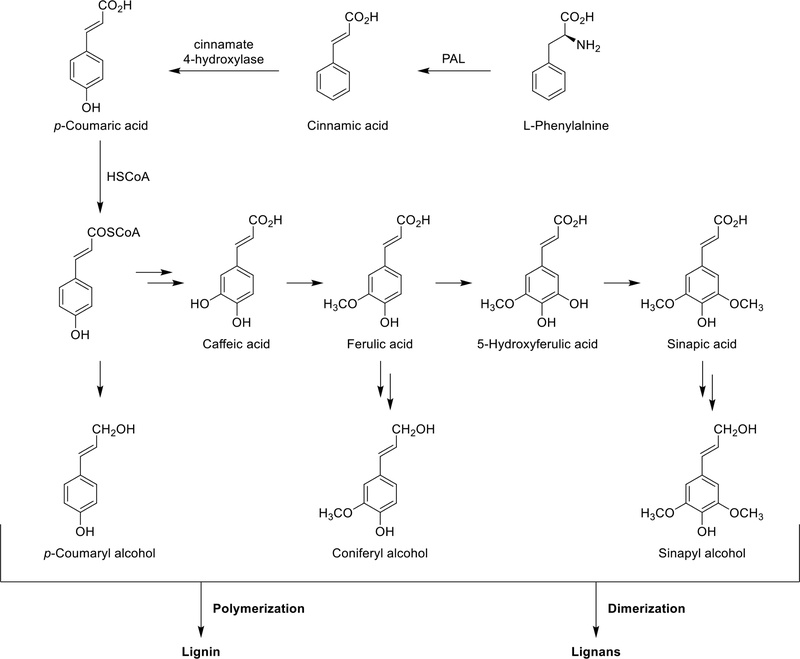
Biosynthesis – Lignans are plant secondary metabolites originating from the shikimic acid biosynthetic pathway. The biosynthetic pathway of lignans begins with the deamination of L-phenylalanine, a precursor with a C6C3 structural unit, by phenylalanine ammonia-lyase (PAL), resulting in the formation of trans-cinnamic acid (Scheme 1). Subsequently, cinnamic acid undergoes hydroxylation via P450 enzymes and cinnamate 4-hydroxylase leads to p-coumaric acid. This is activated by coenzyme A (CoA) to form p-coumaroyl-CoA, a thioester derivative. Finally, in the presence of NADPH, it undergoes reduction to form the corresponding alcohol. The formation of coenzyme A ester facilitates the first reduction step by introducing a more effective leaving group (CoAS) for the NADPH-dependent reaction (Scheme 1). This series of enzymatic transformations constitutes the early steps in the biosynthetic pathway leading to the production of lignans in plants.
The biosynthesis of lignans with 9(9′)-oxygen is most extensively studied. This type of lignan is produced through enantioselective dimerization of two coniferyl alcohol units, facilitated by a dirigent protein (DIR), resulting in the formation of pinoresinol (furofuran) (Scheme 2). In diverse plant species including Forsythia, Linum, and Podophyllum, pinoresinol is subsequently reduced by pinoresinol/lariciresinol reductase (PLR), utilizing lariciresinol (furan) as an intermediate, to form secoisolariciresinol (dibenzylbutane).11–13 Additionally, pinoresinol undergoes glucosylation by a putative pinoresinol glucosyltransferase (PNGT). Pinoresinol also undergoes glucosylation by a putative pinoresinol glucosyltranferase (PNGT). This glycosylation process is highly likely to inhibit the chemical reactivity of a phenolic hydroxyl group in pinoresinol and enhance the water solubility of the pinoresinol aglycone. As a result, substantial and stable amounts of pinoresinol are generated. In fact, approximately 90% of pinoresinol is found in its glucosylated form in Forsythia spp..14,15 Similarly, both lariciresinol and secoisolariciresinol can undergo glycosylation, facilitated by lariciresinol glycosyltransferase (LRGT) and secoisolariciresinol glycosyltransferase (SIRGT), respectively. Secoisolariciresinol is converted into matairesinol (dibenzylbutyrolactone) by secoisolariciresinol dehydrogenase (SIRD) (Scheme 2).
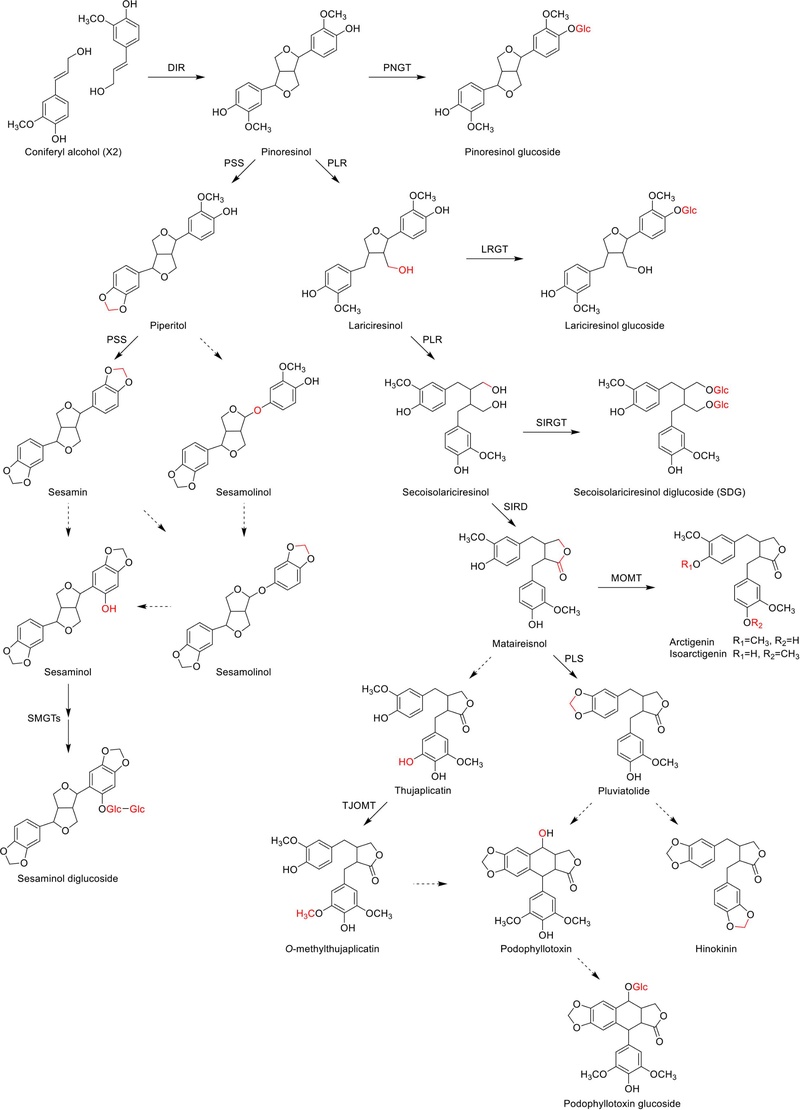
Biosynthetic pathways of major lignans. Pinoresinol sesamin, podpphyllotoxin arctigenin and their glycosides.
Matairesinol undergoes metabolism to arctigenin through the action of matairesinol O-methyltransferase (MOMT), which involves the methylation of a phenolic hydroxyl group (Scheme 2). This process occurs in various plants, including Forsythia koreana, Carthamus tinctorius, and Anthriscus sylvestris.16,17 Moreover, in Forsythia leaves, 70–90% of matairesinol is found to be glucosylated throughout the year, although the enzymes responsible for matairesinol glucosylation have not been identified.14 In Linum, Anthriscus, and Podophyllum plants, matairesinol is also transformed into hinokinin, yatein, or podophyllotoxin through multiple biosynthetic pathways, although all relevant enzymes have not yet been identified.11,18 AsTJOMT, a thujaplicatin-specific plant O-methyltransferase isolated from A. sylvestris, catalyzes the methylation of the 5-hydroxyl group of thujaplicatin, leading to the formation of 5-O-methylthujaplicatin. This compound serves as an intermediate in the biosynthesis of podophyllotoxin.19
Sesamin, another commercially significant lignan, is synthesized through a pathway starting from coniferyl alcohol and involving intermediates such as pinoresinol and piperitol (Scheme 2). Pinoresinol undergoes metabolic conversion into piperitol, and subsequently, piperitol is further transformed into (+)-sesamin by piperitol/sesamin synthase (PSS), an enzyme belonging to the cytochrome P450 family, CYP81Q1, which is responsible for via formation of two methylenedioxy bridges. Although sesamin is anticipated to be metabolized into sesaminol and sesamolin, the detailed molecular mechanisms of this conversion have not yet been fully elucidated. In some sesame plant species such as Sesamum radiatum, Sesamum indicium, and Sesamum alatum, sesaminol undergoes glucosylation at its 2-hydroxyl position catalyzed by sesaminol 2-O-glucosyltransferase (SMGT). Additionally, there is further glucosylation of sesaminol 2-O-monoglucoside at the 6-position of the conjugated glucose.11,13,20
Structure Classification
Lignans – As previously mentioned, “lignans”, formed through various positions of polymerization and modes of dehydration condensation of oxygen-containing groups of side chains, can be further categorized into six different carbon skeletons (Fig. 3). These are classified as dibenzylbutane, tetrahydrofuran, arylnaphthalene and aryltetralin, dibenzocyclooctadiene, and furofuran lignans.
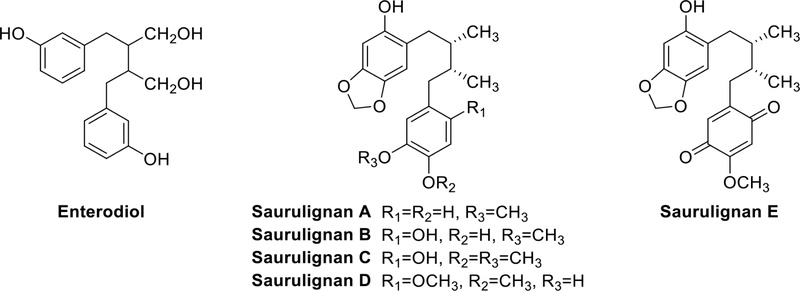
Dibenzylbutane lignans – Dibenzylbutane lignans are composed of two phenylpropanoids connected by carbons C8 and C8′, often serving as precursors for other types. Their differences lie in the variations of substituents at the benzene rings and C-8/C-8′. Sometimes, C9 and C9′ can also be -OH groups. Enterodiol (Fig. 4) is one of the substances formed through the action of intestinal bacteria on lignan precursors. It can be excreted in the form of glucuronide in the urine and bile of both humans and mammals. Enterodiol has garnered interest in research for its potential in tumor prevention and various physiological effects.21 Saurulignan A-E (Fig. 4) have been reported from Saururus chinensis and Schisandra chinensis and Saurulignan E has been associated with platelet aggregation inhibitory activity.22
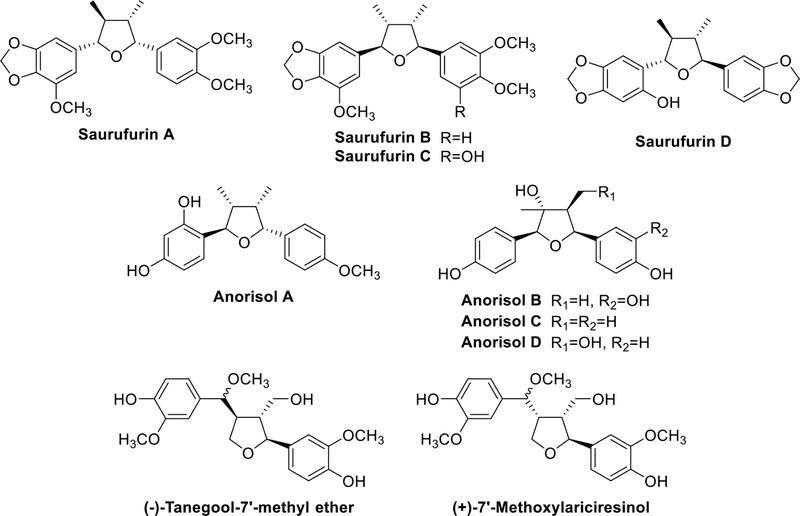
Tetrahydrofuran lignans – Tetrahydrofuran lignans have a furan ring formed by 7-O-7′, 9-O-9′, or 7-O-9′. They have been isolated from plants of various families such as Saururaceae, Asteraceae, Lauraceae, Oleaceae, and others. The benzene rings usually have one or more OMe and OH groups in 7-O-7′ type tetrahydrofuran lignans. Saurufurins A–D (Fig. 5) were obtained from the aerial parts of Saururus chinensis (Saururaceae).22 Anorisols A−D (Fig. 5) were isolated from the roots, stems, leaves and twigs of Anogeissus rivularis (Combretaceae).23 Most compounds belong to the 7-O-9′ type. Tetrahydrofuran lignans isolated from the fruit of Arctium lappa L. (Asteraceae) demonstrated a significant hepatoprotective effect against D-galactosamine-induced cytotoxicity in HL-7702 hepatic cells.24 (-)-tanegool-7′-methyl ether and (+)-7′-methoxylariciresinol (Fig. 5), isolated from the roots and rhizomes of Sinopodophyllum emodi, exhibited excellent cytotoxicity against HeLa and KB cell lines.25
Arylnaphthalene and aryltetralin lignans – Arylnaphthalene lignans are similar to diarylbutane lignans, but they differ in the six-membered ring connecting C-6 and C-7′. Common substituents on the benzene ring typically include functional groups like OH and OMe. Notable examples of arylnaphthalene lignans, including justicidin A, B, taiwanin C, and diphyllin (Fig. 6), have been isolated from Justicia procumbens (Acanthaceae). These lignans exhibit potent cytotoxic effects against various cancer cells in in vitro studies.26,27 Many arylnaphthalene lignans serve as components of traditional herbal remedies and have garnered increased attention over recent decades due to their demonstrated cytotoxic, antiviral, fungicidal, antiprotozoal, and antiplatelet activities in cell-based assays.28–31 The diverse biological activities of these lignans underscore their potential significance in the development of therapeutic agents.
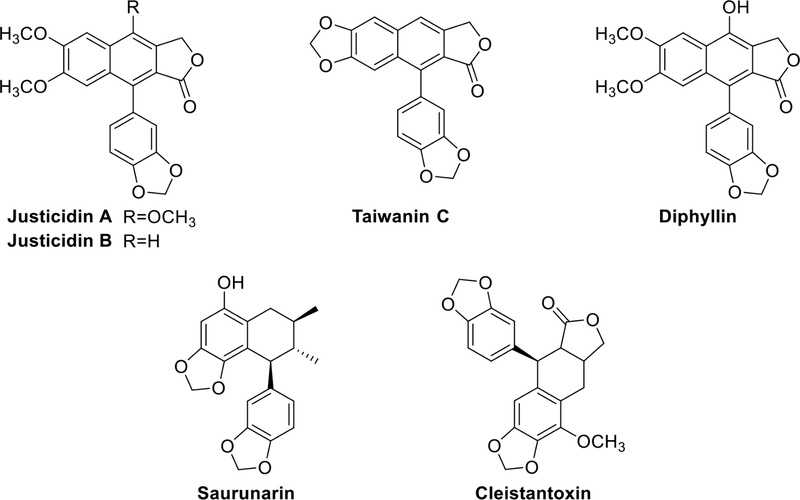
Examples of arylnaphthalene (Justicidin A, B, Taiwanin C, and Diphyllin) and aryltetralin (Saurunarin and Cleistantoxin) lignans.
On the other hand, aryltetralin lignans have a tetrahydronaphthalene (tetralin) core. Saurunarin (Fig. 6) was also isolated from the aerial parts of Saururus chinensis (Saururaceae).22 Cleistantoxin (Fig. 6), isolated from the fruits of Cleistanthus indochinensis (Euphorbiaceae), exhibits strong cytotoxic activity against various cancer cell lines. Its structure is reminiscent of podophyllotoxin, well-known for its antiviral and antitumor properties.32 Its structure is similar to that of podophyllotoxin, which is well-known for its antiviral and antitumor properties.
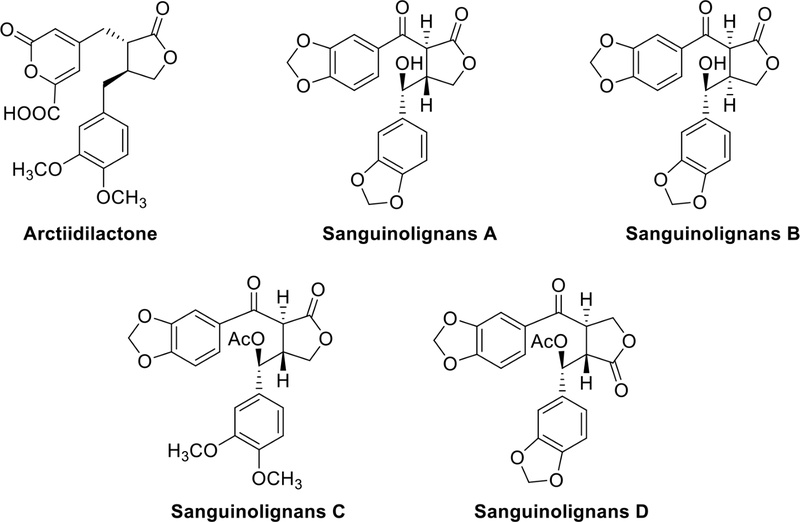
Dibenzylbutyrolalctone lignans – Dibenzylbutyrolalctone lignans which are derived from the dibenzylbutane skeleton, have a lactone ring formed by an ester bond at C-9′. Arctiidilactone (Fig. 7), isolated from the fruits of Arctium lappa L. (Asteraceae), is a novel rare butyrolactone lignan with a 6-carboxyl-2-pyrone moiety. Moreover, several butyrolactone lignans were isolated from this plant demonstrated enhanced anti-inflammatory effects.33 Sanguinolignans A–D (Fig. 7) were isolated from the leaves of Piper sanguineispicum Trel. (Piperaceae). Notably, sanguinolignan A (Fig. 7) exhibited moderate antileishmanial activity, but interestingly it had a very low cytotoxicity on macrophage and cancer or even normal cell lines.34
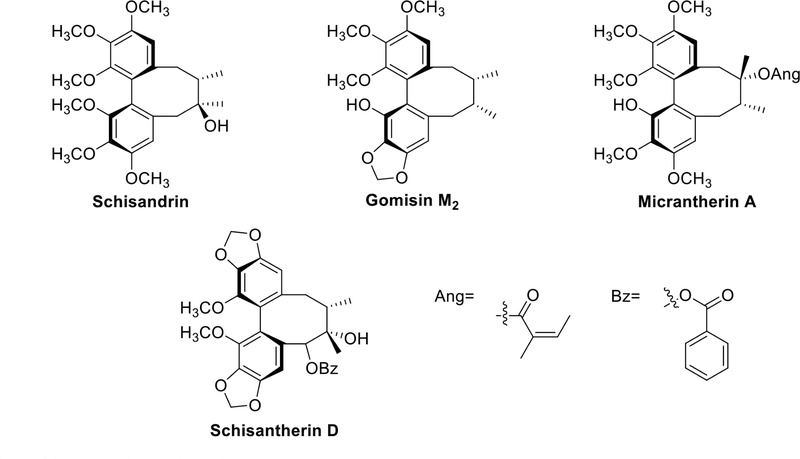
Dibenzocyclooctadiene lignans are unique class of lignans characterized by the C-2/C-2′ linkage and the typical C-8/C-8′ junction of the phenylpropanoid precursor units. Structurally, these lignans feature a cyclooctadiene ring, preventing free rotation of the two benzene rings. This structural feature leads to a distinctive configuration between the two benzene rings and the cyclooctadiene ring structures. These lignans are primarily isolated from and are characteristic compounds of the Schisandraceae family. Dibenzocyclooctadiene lignans play an outstanding role in liver protection.34 Schisandrin, gomisin M2, micrantherin A and schisantherin D (Fig. 8), have been isolated from Schisandra chinensis and have demonstrated hepatoprotective effects.35
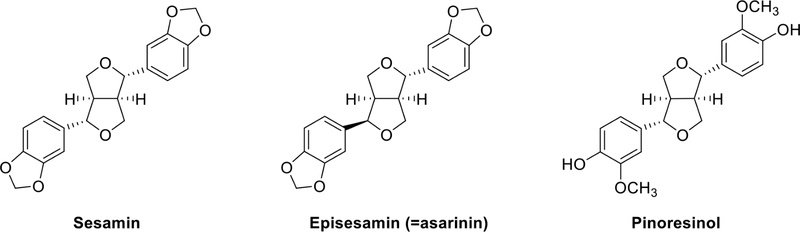
Furofuran lignans – Furofuran lignans are defined by a dual-ring oxygen structure comprising two connected furan rings, allowing for various optical isomers. Notably, these structures readily form glycosides by easily associating with sugars, and the alteration of substituents on the glycosylation group is a prominent feature. Furfuran-type lignans, such as sesamin, episesamin, or pinoresinol (Fig. 9), are widely distributed in edible plants.36 Most dietary lignans of this type are metabolized by gut microflora to enterolactone and enterodiol, collectively known as enterolignans and traditionally classified as phytoestrogens.37 Rich sources of lignans include flaxseed, sesame seeds, cereal products, and Brassica vegetables.36,38
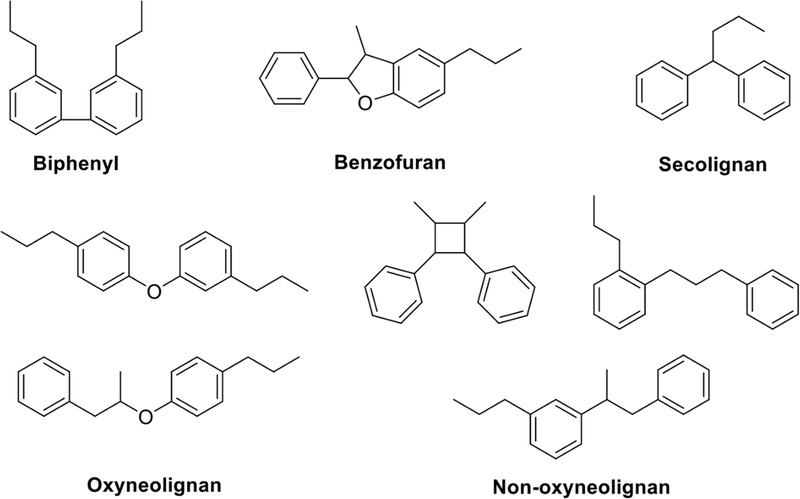
Neolignans – “Neolignans” are compounds in which, unlike typical lignans, two phenylpropanoid units are not linked by C-8 atoms. This category encompasses biphenyls, benzofurans, secoliganans, and various other types (Fig. 10).
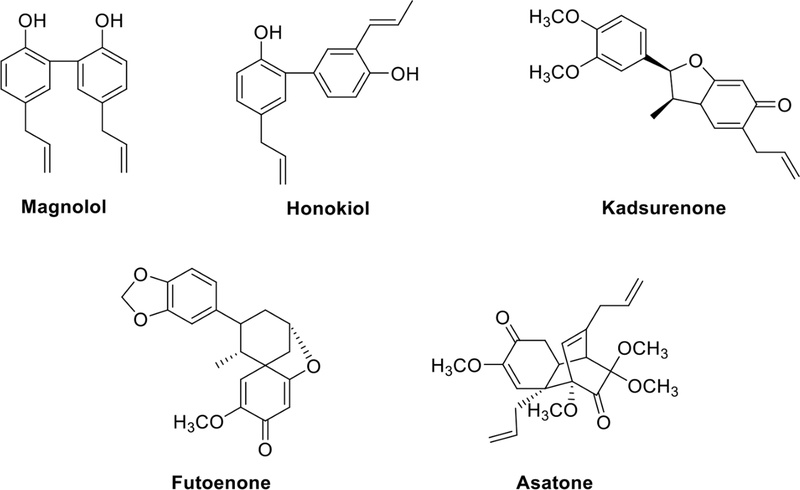
Biphenyl lignans, such as magnolol and honokiol (Fig. 11), are directly linked to each other through the phenyl parent nucleus. These compounds are isolated from the root and stem bark of the magnolia tree and have demonstrated properties associated with spontaneous movement inhibition, central muscle relaxation, anti-stress effects on gastric ulcers, and the inhibition of gastric acid secretion. In the aerial parts or stems of several Piper species, kadsurenone (Fig. 11), identified as a benzofuran-type neolignan, has been shown to inhibit PAF-induced production of prostacyclin (PGI2) and significantly reduce the release of inflammatory mediators.39 Futoenone (Fig. 11), a novel spiro-cyclohexadienone derivative isolated from the stems of Piper kadsura, has been found to inhibit PAF-induced platelet aggregation40 and demonstrate activity against matrix metalloproteinases, including stromelysin, collagenase, and gelatinase.41 Asatone (Fig. 11) is an active component extracted from the Chinese herb Radix et Rhizoma Asari. It exhibits anti-inflammatory effects by activating NF-κB and downregulating p-MAPK (ERK, JNK, and p38) pathways.42
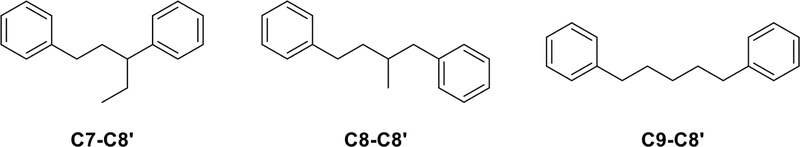
Norlignans – Norlignans are characterized by the absence of the central carbon-carbon linkage between the two phenylpropanoid units, which sets them apart from classical lignans. These compounds exhibit variations in their chemical structures based on the linkage position between the two units. There are three main groups of norlignans classified according to the linkage type: C7-C8′ linkage type, C8-C8′ linkage type, and C9-C8′ linkage type (Fig. 12).
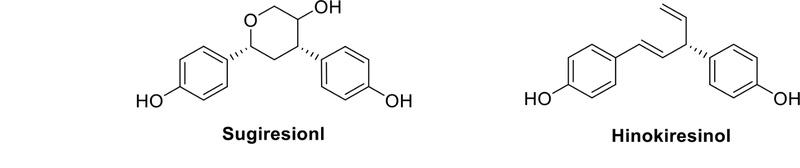
Norlignans are primarily found in gymnosperms, particularly in the Taxodiaceae and Cupressaceae within the gymnospermous plants. Sugiresinol and hinokiresinol (Fig. 13) are found in Cryptmeria japonica and Chamaecyparis obtuse, respectively.43 The discovery of norlignans in these plant types is essential for understanding their biological activities and chemical characteristics. The absence of the central carbon-carbon linkage in norlignans compared to classical lignans may have implications for their properties and biological activities. Like other lignans, norlignans are subjects of study for potential health benefits and biological activities, particularly in the context of antioxidant properties and plant defense mechanisms.44
Lignan oligomer (= polymeric lignans) – Oligolignans, consisting of sequilignans (trimers) and dilignans (tetramers), are condensed forms of lignans resulting from the condensation of three or four lignan units, respectively. The structures of these oligomers can vary based on the specific connections between lignan units. The linkages between units often involve various types of bonds, such as ether linkages or other covalent bonds. This diversity in bonding patterns contributes to the unique chemical and biological properties exhibited by lignan oligomers. The study of oligolignans is crucial for understanding the complexity of lignan chemistry and exploring potential applications, especially in the context of their distinctive properties and functions.
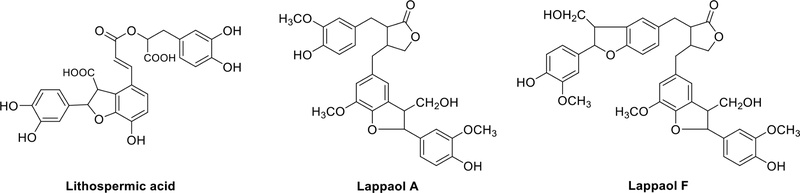
Lithospermic acid (Fig. 14), a prominent oligolignan found in the roots of Lithospermum erythrorhizon, exhibits noteworthy anti-oxidative, anti-inflammatory, and hepatoprotective activities.45 In the seeds of Arctium lappa, lignans like lappaol A and lappaol F (Fig. 14) have been identified. Notably, lappaol F stands out for its ability to inhibit nitric oxide (NO) production, suggesting that it may possess anti-inflammatory properties.46
Lignoid: flavolignan, coumarolignan, xantholignan – “Lignoid” typically refers to compounds or substances that share structural similarities with lignans. This may have lignan-like structure or share certain features with lignans, but they are not necessarily classified as lignans themselves. The term is often used in the context of phytochemistry to describe compounds that are related to lignans but may have variations in their chemical structure.
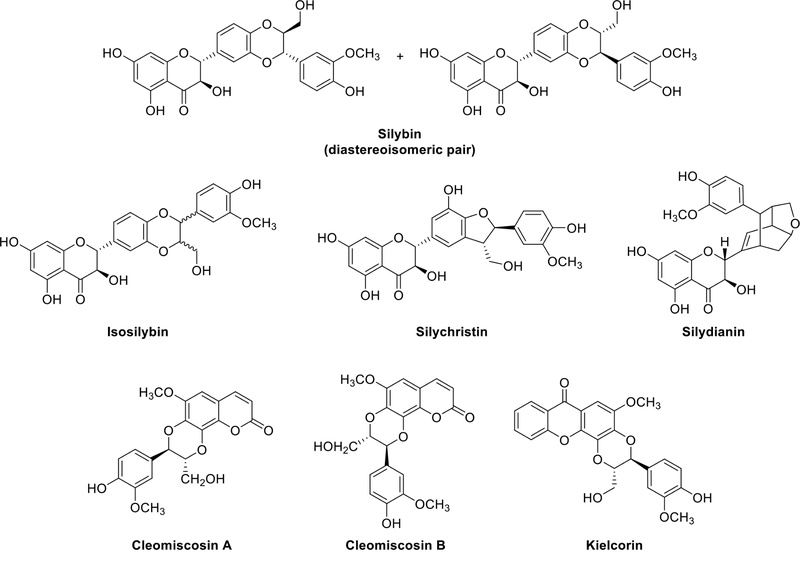
Silymarin (Fig. 15) is a flavolignan compound found in the fruits of Silybum marianum, commonly known as milk thistle and belonging to the Compositae family. It constitutes approximately 1.5–3% of the fruit and is renowned for its applications in treating liver-related disorders.47 The liver-protective effects of silymarin are often attributed to mechanisms such as the inhibition of toxin absorption, antioxidant activity, and scavenging of free radicals. Cleomiscosins A and B (Fig. 15) are coumarinolignans isolated from Cleome viscosa (Cleomaceae), a plant commonly known as “Asian spider flower” or ”tickweed” and belonging to the Cleomaceae family. These compounds are recognized for their potential therapeutic applications, particularly in liver health.48 Kielcorin (Fig. 15), a specific xantholignoid found in Kielmeyera coriacea (Kiclmeyeroideae), exhibites antibacterial, anti-inflammatory, and hepatoprotective activity.49
Extraction of Lignans
The extraction process is a crucial step, and several parameters related to it, such as the choice of solvent, method, time, temperature, solvent-to-sample ratio, and the number of repeat extractions of a sample, should be considered for the optimum recovery of lignans from the plant matrix (Fig. 16). Traditional methods like Soxhlet, maceration, and digestion have been commonly applied for lignan extraction. While these conventional extraction methods are favored for their simplicity and lack of complex apparatus, they come with drawbacks such as extended extraction times and the use of large solvent volumes.
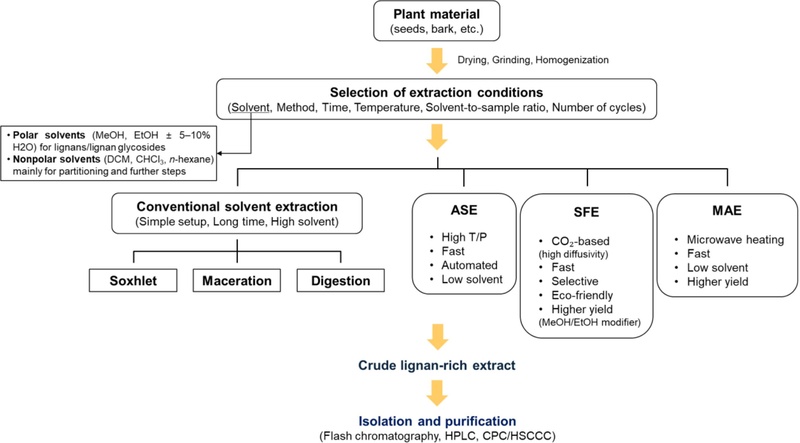
Overview of lignan extraction workflows. Plant material is dried, and ground, followed by extraction using conventional or alternative technologies, including solvent extraction, ASE, SFE, and MAE. After concentration, a crude lignan-rich extract is obtained and subjected to chromatographic isolation.
In addition to conventional methods, alternative extraction techniques are gaining popularity for lignan extraction. These methods include accelerated solvent extraction (ASE), supercritical fluid extraction (SFE), and microwave-assisted extraction (MAE) and offer potential advantages in terms of efficiency and reduced extraction time. More recently, advances in sample pretreatment have introduced various microextraction based approaches, such as dispersive liquid microextraction, miniaturized solid phase extraction, and novel sorbent assisted dispersive solid phase microextraction, which offer rapid, efficient, and environmentally benign alternatives for lignan extraction.50
Solvent extraction – The choice of extraction solvent for lignans should consider the physicochemical properties of the target compounds, such as polarity, solubility, and lipophilicity. Commonly, polar solvents like ethanol and methanol are used for extracting lignans, as they are relatively lipophilic. However, when dealing with a mixture of lignans and lignan glycosides, it is advisable to add around 5–10% water to the solvent to enhance extraction. Although some lignans can be extracted using nonpolar solvents like dichloromethane, chloroform, or n-hexane, these are typically employed in subsequent stages such as solvent partitioning, separation, and isolation.51
A recent advancement in lignan extraction involves the use of Accelerated Solvent Extraction (ASE), a highly effective method (Fig. 16). This automated process occurs at elevated temperature and pressure under an inert nitrogen atmosphere, allowing for fast and sequential extraction with relatively small solvent volumes. ASE has demonstrated success in extracting lignans from various tree species, particularly yielding lignan-rich extracts from Norway spruce knots.52–56
Solid-phase extraction (SPE) – Solid-phase extraction (SPE) is a versatile and crucial technique for recovering and purifying phenolic compounds from diverse sample types. It offers rapid and sensitive sample preparation, replacing cumbersome conventional isolation and extraction methods. The choice of sorbent, such as C18 reversed-phase, amino phases, or anion-exchange materials, depends on the specific analysis requirements.51 Although the utilization of solid-phase extraction (SPE) for lignans is not as widespread as various solvent extraction techniques, it has found application in the analysis of lignans in flaxseed or biological fluids.57 In these cases, SPE serves as a purification step after alkaline or enzymatic hydrolysis of a raw extract or is employed for the fractionation of oligomers from a non-hydrolyzed raw extract.58,59 Despite being less common, SPE has been successfully used for the extraction of simple phenols and polyphenols from plants and food, although lignans appear to have been somewhat neglected in these applications.60
Supercritical fluid extraction (SFE) – Supercritical fluid extraction (SFE) is considered an environmentally friendly extraction technique. This has several advantages, including reducing the use of extraction solvents, shortening extraction time, increasing yield and convenient method for sample preparation before analyzing compounds in plant matrices (Fig. 16). Supercritical solvents have unique properties, including high diffusivities, lower viscosities, and almost zero surface tensions. These properties provide advantages such as shorter extraction times, enhanced selectivity, and the absence of residual organic solvents in the final extract. The most used supercritical solvent is pure or modified carbon dioxide due to its low critical parameters, allowing for mild extraction conditions, including temperature and pressure. Carbon dioxide is recognized for its non-toxic, non-flammable, and environmentally friendly characteristics.61 SFE has been predominantly utilized for lignan extraction from various parts of Schisandra chinensis. Numerous studies have been conducted to optimize the SFE methods for the extraction of lignans from this plant. Recent publications have contributed to the understanding of key factors affecting lignan extractability using SFE.62 Notably, the addition of methanol or ethanol proven to be a significant factor in substantially increasing the lignan yield in SFE process. The successful application of SFE is not limited to Schisandra chinensis, as it has been effectively used for lignan extraction from diverse plants sources, including Forsythia koreana and Magnolia cortex.61
Microwave-assisted extraction (MAE) – Microwave-assisted extraction (MAE) has emerged as a significant method for lignan extraction, utilizing microwave energy as an electromagnetic radiation source. This energy induces molecular motion through the migration of ions and rotation of dipoles, aligning molecules in both the solvent and the matrix. MAE provides rapid energy delivery to the solvent, leading to swift heating due to collisions with surrounding molecules. The technique offers advantages such as prompt initiation, reduced analysis time, and lower solvent consumption compared to other extraction methods, making it a valuable tool for extracting natural products from plant materials (Fig. 16).63
MAE’s effectiveness was initially demonstrated in lignan extraction from flaxseeds. Specifically, the primary lignan, secoisolariciresinol diglucoside (SDG), exhibited a 1.3-fold increase in extraction yield with MAE compared to traditional methods, irrespective of sodium hydroxide concentration. Besides enhanced yields, MAE demonstrated benefits of shorter extraction times and a simplified procedure.64
These findings underscore MAE’s potential as a more efficient and time-effective method for extracting phenolic compounds from flaxseed, thereby enhancing its value in the extraction of natural products from plant materials. Importantly, MAE has been successfully optimized for preparing lignan-rich extracts from various plant sources, further establishing its versatility in phytochemical extraction processes.63
Isolation and Purification Methods
Flash chromatography – Flash chromatography on silica gel or Sephadex LH-20 columns is commonly employed for the initial fractionation, separation, and preparative isolation of natural compounds, including lignans. This approach offers distinct advantages for the preparative isolation of selected lignans from raw material due to its capability to handle larger sample loads compared to high-performance liquid chromatography (HPLC).
In the process of lignan isolation, the choice of suitable solvent mixtures is determined through systematic experiments involving both column chromatography and thin layer chromatography (TLC). For the preparative separation of lignans, the selection of the column or TLC material typically involves normal-phase or reversed-phase (RP) C18. Interestingly, RP C8 has been found in some cases to yield better separation than C18. Various solvent mixtures have been identified as suitable, including methanol/water, acetonitrile/water, acetonitrile/methanol/water, ethanol/water, n-hexane/chloroform/methanol, or n-hexane/ethyl acetate in different ratios. The choice between isocratic and gradient elution depends on the complexity of the extract.51,63
Semi-preparative HPLC of lignans – The use of a semi-preparative HPLC system equipped with a UV-Vis detector is highlighted. UV absorption is monitored at specific wavelengths, such as 254 or 280 nm. Commonly employed are reverse-phase C18 columns. Isocratic elution is utilized, maintaining a constant mobile phase composition. Mobile phases include acetonitrile-water mixtures or methanol-water mixtures, with varying proportions depending on the study. The addition of 0.1% formic acid to the mobile phase is noted to enhance peak resolution for structures containing hydroxyl groups. Gradient elution, involving a change in mobile phase composition during the analysis, is described as an effective method for isolating specific lignans in gram quantities.
Countercurrent separation of lignans – Centrifugal Partition Chromatography (CPC) and high-speed countercurrent chromatography (HSCCC) are highlighted as all-liquid chromatographic techniques, differing from conventional methods that use solid stationary phases. This distinction is emphasized due to the avoidance of adsorption losses and the formation of artifacts, common in conventional chromatography with active surfaces. One of the notable strengths of CPC and HSCCC lies in their versatility in selecting solvent systems. These techniques offer a wide range of solvent options, providing flexibility in the development of chromatographic methods.
The efficacy of CPC and HPCCC in the purification and isolation of lipid-soluble lignans is emphasized, with specific examples such as sesamin and sesamolin from sesame seeds.65 Additionally, HPCCC is underscored as an efficient method for isolating secoisolariciresinol diglucoside (SDG) from flaxseed, employing a gradient method for improved separation.66,67 Seven lignans including (−)-maglifloenone, futoenone, magnoline, cylohexadienone, fargesone C, fargesone A and fargesone B were isolated and purified from Magnolia sprengeri Pamp. using HSCCC with two-step separation. In the first step, a stepwise elution mode with the two-phase solvent system composed of petroleum ether-ethyl acetate-methanol-water (1:0.8:0.6:1.2, 1:0.8:0.8:1, v/v/v/v). In the second step, the residues were successfully separated by HSCCC with the solvent system composed of petroleum ether-ethyl acetate-methanol-water (1:0.8:1.2:0.6, v/v/v/v).68 Then arylnaphthalene lignans were purified from Justicia procumbens L. in 8 h using HSCCC with the two-phase solvent system composed of petroleum-ethyl acetate-methanol-water (1:0.7:1:0.7, v/v/v/v). The enriched mixture) was further separated using the solvent system consisting of petroleum-ethyl acetate-methanol-water (3:3.8:3:3.8, v/v/v/v).69 A combination of HSCCC and preparative HPLC has led to the purification of six lignans from Schisandra chinensis. HSCCC was performed with a two phase solvent system of n-hexane-ethyl acetate-methanol-water (1:1:1:1, v/v/v/v).70,71 Arctigenin and matairesiol, dibenzylbutyrolactone lignans, were obtained CPC operation from an enriched lignan extract of Forsythia koreana with a two-phase solvent system composed of n-hexane-ethyl acetate-methanol-water (5:5:5:5, v/v/v/v).72 Consecutive sample injection method to purify sesamin and sesamolin from Defatted sesame meal extract. CPC was carried out with a two-phase solvent system consisting of n-hexane-ethyl acetate-methanol-water (8:2:8:2, v/v/v/v) in descending mode.73 Podophyllotoxin, 4′-demethyl podophyllotoxin, and, deoxypodophyllotoxin were purified from Sinopodophyllum emodi by HSCCC using a solvent system composed of n-hexane-ethyl acetate-methanol-water (1.75:1.5:1:0.75, v/v/v/v).74
Qualitative and Quantitative Analysis
Thin layer chromatography (TLC) – Thin-Layer Chromatography (TLC) is emphasized as a simple and cost-effective technique for separating lignans. It is routinely utilized in lignan research, particularly for the initial qualitative assessment of plant extracts. In lignan TLC analysis, silica gel is predominantly employed as the stationary phase. The choice of eluent, or mobile phase, is determined by the physicochemical properties of the lignans present in the sample. Visualization of TLC spots or bands is commonly carried out using a UV lamp on commercially available plates that are impregnated with a fluorophore. Lignans exhibit the property of absorbing UV light, allowing plates without chemical treatment to be observed under a 254 nm wavelength UV light. Other alternative detection method involves spraying plates with various reagents, such as 5% sulfuric acid in ethanol.51,61,63
TLC was historically utilized not only for qualitative analysis but also for quantitative analysis (densitometry) and in procedures for isolation and purification (preparative TLC).61 However, more advanced methods have largely replaced the use of TLC in recent times for quantitative analysis and isolation. High Performance Thin Layer Chromatography (HPTLC) is highlighted as a method used for quantifying lignans in the seed oil of sesame species without saponification, affirming the validity of this method in lignan analysis.66 A more recent TLC method, thin layer chromatography-direct bioautography (TLC-DB), allows on-plate identification of active components, particularly in terms of antioxidative, antibacterial, and enzyme inhibition properties. Despite advancements and the replacement of TLC in some areas, it is emphasized that TLC remains valuable for specific applications. These include the initial examination of plant materials and the monitoring of purification stages in lignans analysis (Table 1).
High-performance liquid chromatography (HPLC) – HPLC usually used for analyzing lignans in biological samples due to its convenience, no need for time-consuming sample preparation. This technique commonly employs reversed phase (RP) columns, with occasional utilization of silica (normal phase) columns for lignan analyses. RP-18 (octadecylsilica) columns are widely utilized, whereas RP-8 columns are better suited for separating hydrophilic lignans, such as 7-hydroxymatairesinol isomers.75,76 HPLC is the most widely used method for the analysis of lignans. Traditionally, UV detection (at a single wavelength) may offer sufficient sensitivity and selectivity for determination of lignans. Traditionally, UV detection at wavelengths of 254 or 280 nm is widely used in plant extracts for lignan determination due to its satisfactory sensitivity. Nine lignans in Schisandra chinensis extract were quantified by HPLC-DAD at 254 nm.77 The total lignans from Arctii Fructus were quantitatively analyzed by HPLC to determine the quality of Arctii Fructus. Total 11 marker components, lappaol H, lappaol C, arctiin, arctignan D, arctignan E, matairesinol, arctignan G, isolappaol A, lappaol A, arctigenin, and lappaol, were successfully quantified at 280 nm with isocratic elution of 47.6% methanol.78
In recent years, a combined HPLC-UV and HPLC-MS method was developed to identification and quantative analysis of lignans. MS has emerged as the most sensitive lignan analysis method, providing quantitative analysis capabilities along with analyte structure determination, including high selectivity, resolution, speed, sensitivity, repeatability, molecular weight and purity. The UV and MS data led to the identification of group characteristics for the major subclasses that can be used to establish the structure of unknown lignan peaks in the chromatograms of crude extract of Linum usitatissimum and L. bienne Mill.79,80 In addition, HPLC-ESI-MS (coupled or in combination with a PDA detector) was used for the identification of 12 lignans (schisandrol A, schisandrol B, gomisin G, schisantherin A, schisantherin D, schisanhenol, (+)-anwulignan, deoxyschisandrin, schisandrin B, schisandrin C, 6-O-benzoylgomisin O, and interiotherin A) in extracts of Schisandra sphenanthera fruits.81
HPLC-MS method was applied to determine lignans in biological fluids. The major part of lignans in biological fluids are present in conjugated forms such as glucuronides and sulfates, so these samples are typically hydrolysed enzymatically using β-glucuronidase/sulfatase preparations prior to sample extraction and chromatographic analysis. Urine and blood (serum or plasma) are the most frequently analyzed biological fluids. The pharmacokinetic profile of arctiin, the major active lignan in fruits of Arctium lappa L., was carried out by an LC-UV-MS. Arctigenin, main metabolite of arctiin, was identified by HPLC-MS and quantified by HPLC-UV at 280 nm in plasma and organs of Sprague-Dawley rats.82 Another study of arctiin and its metabolites in rat urine and feces were performed by HPLC coupled Q-TOF mass spectrometer. Arctiin and its three metabolites, (−)-enterolactone, (−)-arctigenin and (2R,3R)-2-(3′-hydroxybenzyl)-3-(3″,4″-dimethoxybenzyl)-butyrolactone were identified and quantitative analysed by UV at 280 nm.83 The absorption profiles of Schisandra chinensis in the human Caco-2 cell monolayer and rat everted gut sac models, as well as in rat plasma. By analyzing MSn characteristics of peaks acquired by HPLC-DAD-APCI-MSn determination, absorbable lignans in the Schisandra extract and related metabolites were identified.84 Picropodophyllin and its isomer podophyllotoxin in human serum samples with electrospray ionization of hexylamine adducts were determined by HPLC-MS/MS. The mobile phase was consisted of 2.5 mM hexylamine and 5 mM formic acid in water and methanol. The hexylamine adducts rather than proton adducts were more sensitive. The limit of quantification (LOQ) was 0.01 μmol/L for picropodophyllin and podophyllotoxin.85
After oral administration of magnolol and honokiol emulsion, pharmacokinetic study applied. A sensitive and specific ultra-performance liquid chromatography/tandem mass spectrometry (UPLC-MS/MS) method was developed for the investigation of the pharmacokinetics of magnolol and honokiol in rats. The plasma samples were deproteinized with acetonitrile, the post-treatment samples were analyzed on an Agela C18 column interfaced with a triple quadrupole tandem mass spectrometer in negative electrospray ionization mode. Acetonitrile and 5 mmol/L ammonium acetate buffer solution (65:35, v/v) was used as the mobile phase.86 The pharmacokinetics and safety of multiple oral doses study of sesame lignans (sesamin and episesamin) was conducted. Sesame lignans and metabolites were analyzed by an Acquity UPLC system coupled to a Quattro Micro MS System. Metabolites were extracted from plasm after hydrolysis with β-glucuronidase/sulfatase, and applied to an Oasis HLB solid-phase extraction column preconditioned.87
Chiral HPLC separation for lignans – Lignans, chiral plant metabolites that are generally found in enantiomerically pure (+)- or (−)-form or in pure enantiomeric excess. The enantiomeric composition could be determined by HPLC coupled to UV/laser polarimetric detection.88 Typically, enantiomeric configuration was determined through isolation followed by optical rotation measurement. The separation of enantiomers in a mixture can also be separated on a chiral HPLC column. This requires a reference to a known pure enantiomer. Enantiomeric lignans and neolignans were isolated from Phyllanthus glaucus. The racemic or partial racemic mixtures were successfully separated by chiral HPLC using different types of chiral columns (Daicel chiral-pak IA column or Daicel chiral-pak ASH column) with various mobile phases (composition of n-hexane–ethanol–formic acid). The absolute configurations of enantiomers were determined by computational analysis of their electronic circular dichroism (ECD) spectrum by comparing their experimental ECD spectra and optical rotation values. Chiral resolution of phyllanglaucin A was performed on Daicel chiral-pak IA column (eluted with n-hexane-ethanol-formic acid, 70:30:0.1, v/v/v).89 Sesamin, with a remarkable rotation and significant Cotton effects, was isolated from Juglans mandshurica Maxim. (±)-Sesamin was analyzed by a Daicel Chiralpak IC (eluted with n-hexane-2-propanol, v/v, 2:1) to obtain (+)-sesamin and (-)-sesamin. The absolute configurations were determined by computational analysis of their ECD spectrum. In Aβ1–42 aggregation assay, (−)-sesamin (80.6±1.53% inhibiton) was more potent inhibitory activity than (+)-sesamin (67.7±2.04%) at the concentration of 20μM.90
(±)-Licarin A, a neolignan obtained by the oxidative coupling reaction of isoeugenol, was directly performed by chiral high-performance liquid chromatography (HPLC-PDA) protocol (Chiralpack® AD column; n-hexane-2-propanol, 9:1, v/v). The schistosomicidal and trypanocidal structure-activity relationships for (±)-licarin A and its (+)- and (−)-enantiomers were also displayed.91 The synthetic racemates (±)-larreatricins and (±)-8′-epi-larreatricins were separated using a Chirobiotic V column eluted with ethanol-hexanes (12:88, v/v) and then, chiral HPLC analysis of the four isolated diastereomers from L. tridentata revealed that larreatricin and 8′-epi-larreatricin were present primarily as the (−)-form.92
Structural Elucidation
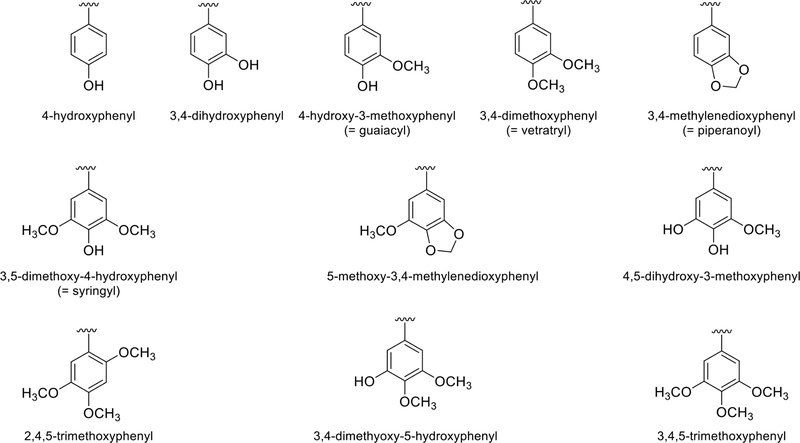
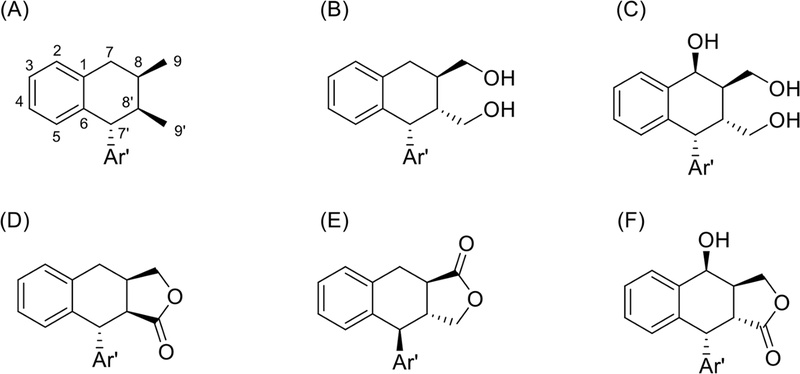
Structural identification of lignans using spectroscopic methods – Most chemical structures of lignans can be divided into aryl groups (aromatic rings) and side chains. The aromatic rings are usually oxy-substituted, particularly at the para position to side-chain substitution. Generally, the aromatic ring is oxygenated at the 3-, and 4-positions and is represented in Fig. 17.
In addition, the side chain moiety is mostly 8-8′ (or β-β′) connected, which can be used to confirm the structures. The two aryl groups (aromatics), which occur frequently in lignans, are depicted in Fig. 1, so identifying the side chain structures is important for lignan structure analysis. 1H and 13C NMR were usually used to determine the chemical structures of lignans. Lignans were classified into several derivatives according to their side chain structures. The methods of structural analysis using NMR are described below for each derivative.
For most lignans, twelve carbon peaks corresponding to the aromatic ring can be observed in 13C-NMR, and then the remaining six side chain carbons were interpreted by 1D and 2D NMR. The linkage of the side chains can be seen through the coupling of the protons corresponding to C7-C8-C9 (C7′-C8′-C9′), and the C8-C8′ linkage of each side chain can also be determined. HMBC also confirms the connection between the H-7 proton of the side chain and the C-1 carbon of the aromatic ring to confirm the basic structure. The structural features of each lignan derivative and the structural analysis method using NMR are described. In particular, coupling constants, 13C-NMR values, and methods for determining relative configurations using NOE (or NOESY) are included. The absolute configuration is usually determined by optical rotation, circular dichroism (CD), or electronic circular dichroism (ECD).
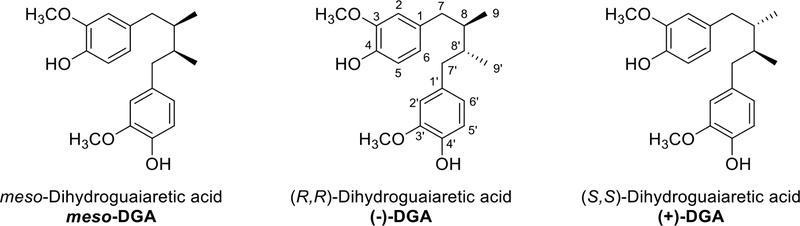
Dibenzylbutane and dibenzylbutanediol derivatives – Dihydorguaiaretic acid (DGA) was isolated from Myristica argentea, Pycnanthus angolensis, Saururus cernuus, meso-dihydroguaiaretic acid was also isolated Saururus chinensis.93–96 DGA and meso-DGA showed nine carbon signals, six aromatic ring signals, three aliphatic signals and an additional one methoxy signal in 13C-NMR. Sometimes lignans are symmetrical, meaning that although though the backbone is made up of 18 carbons, only half that number, nine signals, are observed. In this case, the molecular weight should be checked and compared with that of simple phenylpropanoids. The Ar-CH2-CH-CH3 (or Ar-CH2-CH-CH2OH, dibenzylbutanediol) moiety was confirmed by two proton signals (benzylic methylenes) 2.3 and 2.7 ppm (dd, J = 13.5, 9.3 or 5.1 Hz) in 1H-NMR. The structures of the aromatic regions are confirmed by the splitting pattern of the proton in the aromatic region, and the structure of the side chain part is confirmed by proton splitting and coupling constant in 1H NMR. Most lignans have a C8-C8′ linkage, so the C7-C8-C9 linkage and C7′-C8′-C9′ linkage can be identified, and the 8-8′ linkage can also be used to identify the structure. 1H-, 13C-NMR and optical rotation data of dihydroguaiaretic acid (Fig. 18) were described in Table 2.93,97,98
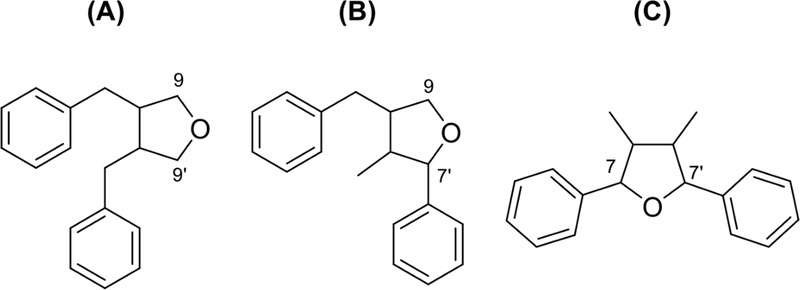
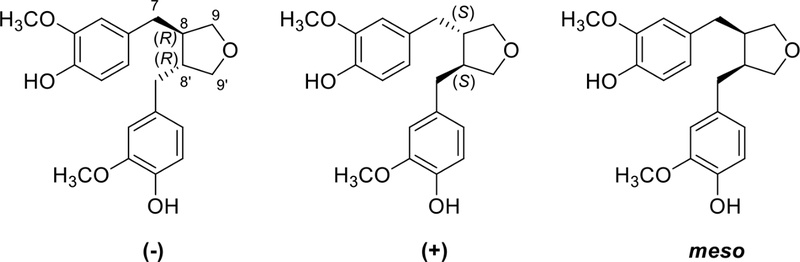
Tetrahydrofuran lignans – Tetrahydrofuran lignans have a furan ring formed by 9-O-9′, 7-O-9′, or 7-O-7′ linkage as shown in Fig. 19.
Tetrahydrofuran lignan type A (furan ring formed by 9-O-9′ linkage) showed two benzylic methylenes, two methines, and two oxymethylenes. Shonanin (4,4′-dihydroxy-3,3′-dimethoxy-9,9′-epoxylignan) was isolated from the leaves of Calocedrus formosana.99 The molecular weight was m/z 344, while 13C-NMR spectrum displayed only 10 signals, indicating that shonanin is a symmetric molecule. 1H-NMR data displayed at δ 2.15 (2H, m, H-8,8′), 2.53 (4H, m) 8.52 (2H, dd, J = 8.7, 5.7 Hz), 3.90 (2H, dd, J = 8.7, 6.6 Hz) indicated that benzylic methylene, methine, and oxymethylene (Ar-CH2-CH-CH2-O) in side chain. The 13C-NMR data of (±)-shonanin and meso-form (Fig. 20) were shown in Table 3.99–101
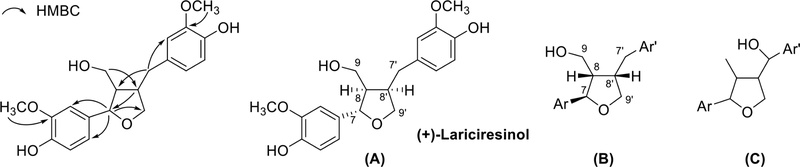
Tetrahydrofuran lignan type B (furan ring formed by 7-O-9′ linkage) showed one benzylic methylene, two methines, one oxymethylene, and one methyl. Terminal CH3 often changes to CH2OH, and hydroxy can substitute to benzylic methylene to form benzylic oxymethine. Consistent with these structural features, lariciresinol shows characteristic long-range HMBC correlations in which H-7 correlates with C-2, C-6, C-8, and C-9′, and H-8 and H-8′ correlate with C-7, C-7′, C-9, and C-9′. These correlations collectively confirm that the compound possesses a tetrahydrofuran skeleton formed through a 7-O-9′ linkage (Fig. 21, Table 4).102,103
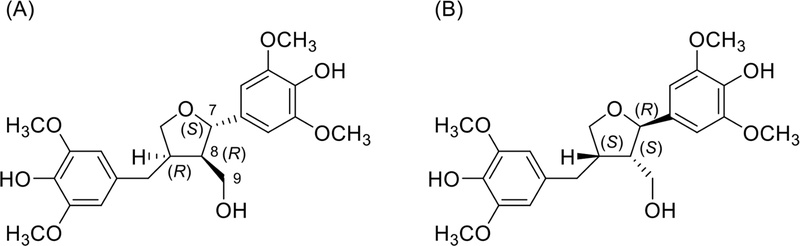
The relative configuration of B type 7-O-9′ furan lignans could be determined by the coupling constant between H-7 and H-8. The coupling constant between H-7 and H-8 (J7−8 = 6.0Hz) confirmed trans-configuration (J7−8 = 3.6 Hz in cis-configuration). The absolute configuration of aglycones of alangilinoside C and ligalbumosides A are determined by the NOESY and CD spectra (Fig. 22). The CD spectra of (A) and (B) was opposite from 180 to 260 nm, A type (positive) and B type (negative).104
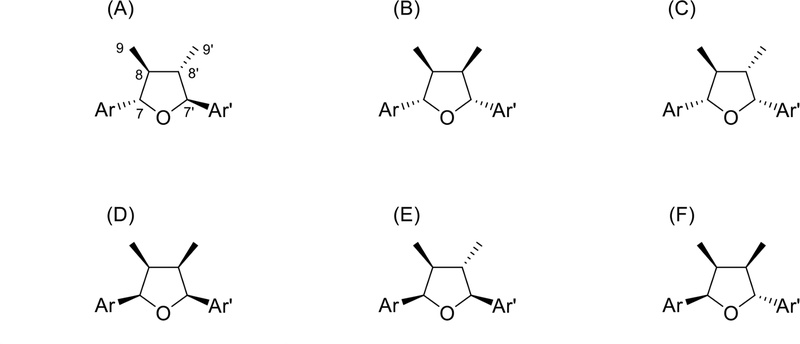
Tetrahydrofuran lignan type C (furan ring formed by 7-O-7′ linkage, Fig. 23) showed two benzylic oxymethines, two methines, and two methyls in 1H-NMR. The comparison of the specific rotation, coupling constants, and the chemical shifts of the aliphatic protons and carbons identified the relative configuration. The 13C-NMR data of relative configuration were described in Table 5.105 The absolute configurations of anorisols A–D (tetrahydrofuran lignans) were established by comparison of the experimental electronic circular dichroism (ECD) spectra with the calculated ECD spectra.23
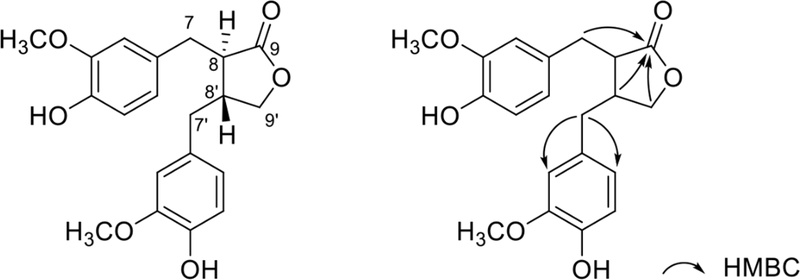
Dibenzylbutyrolactone derivatives – Dibenzylbutyrolactone lignans, such as matairesinol,106 are composed of the side chains Ar-CH2-CH-C=O and Ar-CH2-CH-CH2-O that are connected through 8-8′ and 9-O-9′ linkages (Fig. 24). These side changes were characterized by the signals at δC 34.5 (C-7, benzylic methylene, δH 2.88, 2.94), 46.4 (C8, methane, δH 2.56), 178.6 (C-9, carbonyl), 38.2 (C-7′, benzylic methylene, δH 2.53, 2.61), 40.9 (C-8′, methine, δH, 2.47), and 71.2 (C-9′, oxymethylene, δH 3.88, 4.15).107 The 2D NMR (HMBC) spectrum clearly showed correlations between H-7, H-8′, and H-9′ and C-9 confirming the characteristic dibenzylbutyrolactone skeleton. The aromatic ring system was assigned based on the HMBC correlations from H-7 to C-6 and C-2 (Fig. 24). The remaining aromatic signals were readily assigned by further HMBC correlations, and a similar approach was applied to resolve the signals of the second aromatic ring.108 The relative configuration of H-8 and H-8′ was determined by the chemical shifts. The trans-derivatives showed a poorly resolved spectrum with a four-proton multiplet (H-8, 8′, 7′a, 7′b) at δ 2.5–2.6, a two-proton multiplet (H-7a, 7b) at δ 2.9, with a very small nonequivalence of the protons of each of the two benzyl groups, and the distinct nonequivalence of the protons of the C-9′ methylene group (δ 3.9 and 4.2). In the cis-derivatives, the benzylic methylenes and H-7 and H-7′ were relatively well resolved within a broad range (δ 2.3–3.3), while the hydrogens in each of the benzyl groups were distinctly nonequivalent, although the hydrogens of the C-9′ methylene group were almost equivalent in the δ 4.0–4.1 range. The optical rotations of (8R,8′R) was levorotatory (negative) and the (8S, 8′S) isomer was dextrorotatory (positive) in the trans derivatives.109
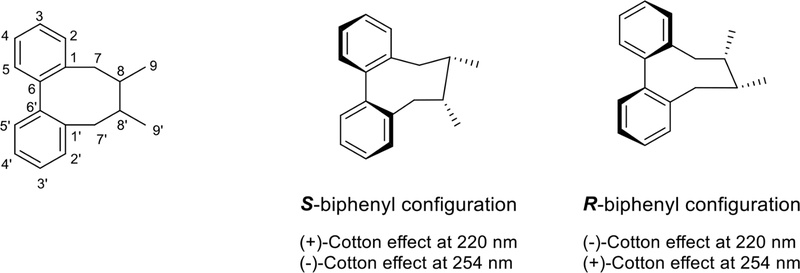
Dibenzocyclooctadiene derivatives – Dibenzocyclooctadiene lignans constitute a distinctive class characterized by the C-2/C-2′ biaryl linkage and the typical C-8/C-8′ junction of phenylpropanoid precursor units. The structural diversity of dibenzocyclooctadienes arises from various combinations of the following structural elements: (i) the substitution pattern of the biaryl unit, allowing for potential replacement of hydrogen(s) with hydroxyl, methoxyl, or methylenedioxy groups; (ii) the substitution pattern and configuration of stereocenters along the aliphatic bridge; (iii) the absolute configuration of the biaryl axis. These lignans exhibit multiple substitutions of methoxyl (OCH3) or methylenedioxy (-OCH2O-) in the aromatic region, leading to a lower count of aromatic protons (Fig. 25).110,111
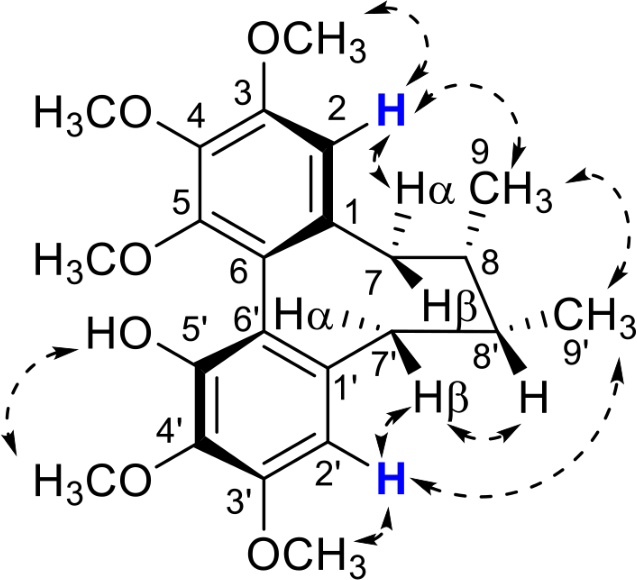
The two aromatic protons (H-2 and H-2′) with chemical shifts between 6.4 and 7.0 ppm in 1H-NMR, are useful for stereochemical and conformational information. NOE (or NOESY) is a powerful approach for identifying the aromatic substitution pattern and stereostructure of a lignan. NOESY correlations of H-2/CH3-9, H-9β; H-2′/H7′β, H-8′ and CH3-9′/CH3-9 indicated a cyclooctadiene lignan with a twisted boat/chair conformation, whose C-8 and C-8′ carbons were in S and R absolute configurations (Fig. 26).112
13C-NMR can identify the position of a methoxy carbon substitution. Compared to other methoxy carbons, the chemical shift of a methoxy carbon adjacent to the biphenyl bond is about 5 ppm upfield. For example, the chemical shift of 3-OCH3 and 3′-OCH3 is at 55.0 ppm, while other methoxy carbons appear at approximately 60.0 ppm. In addition, the chemical shifts of C-7 and C-7′ are effected by hydroxyl or ester group substitution depending on the substituent′s configuration. For 7-β, this substitution shifts C-7 (or C-7′) downfield (δ ≥ 80 ppm); for 7-α, the substitution shifts C-7 upfield (δ ca. 73 ppm).111,112
The biphenyl chromophore in a dibenzocyclooctadiene lacks rotational freedom, resulting in R or S biphenyl configurations. Determination of the absolute configuration of this biphenyl chromophore is feasible through circular dichroism (CD) spectroscopy. In the CD spectrum of a lignan derivative, if both a (+)-Cotton effect at 220 nm and a (–)-Cotton effect at 254 nm are observed, the biphenyl unit typically possesses the S configuration. Conversely, a lignan exhibiting an R configuration will display a CD spectrum with a (–)-Cotton effect at 220 nm and a (+)-Cotton effect at 254 nm.110,113
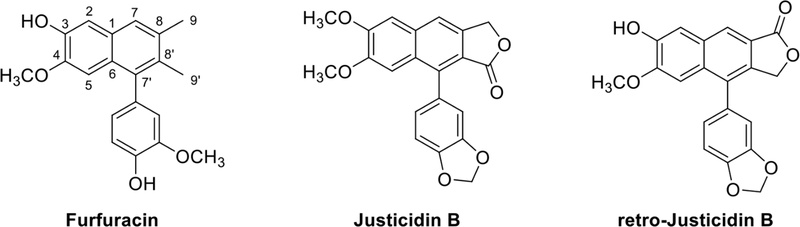
Arylnaphthalene and aryltetralin lignans – The arylnaphthalen lignans showed strong UV absorption maxima at 236 and 287 nm due to the naphthalene chromophore.114 The appearance of 16 aromatic carbons in 13C-NMR is characteristic of arylnaphthalene lignans. Other lignan derivatives typically exhibit 12 aromatic carbons, but arylnaphthalene is unique in that it exhibits 16 aromatic carbons. Two aromatic methyl signals (3H, s) showed ca. 2.07 ~ 2.40 ppm, and the terminal methyl (-CH3) groups are often found to be modified with -CH2OH or lactone ring. Arylnaphthalene lignan lactones are found in a variety of dietary and medicinal herbs including Phyllanthus, Justicia, Haplophyllum, and Cleistanthus.115 Arylnaphthalene lignan lactones are classified into two types based on the position of lactone. The representative arylnaphthalene lignan lactones, justicidin B and retro-justicidin B (Fig. 27), could be distinguished by 13C-NMR data, C-9 (ca. 69.5 ppm) in justicidin and C-9′ (ca. 66.5 ppm) in retro-justicidin B.
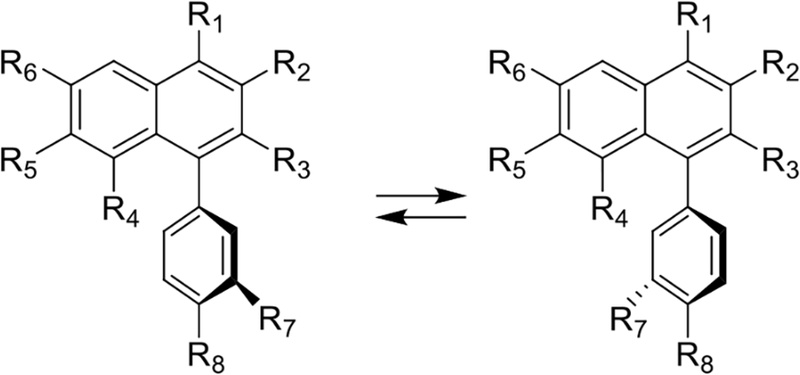
In some arylnaphthalene lignan lactones, the hindered rotation in the phenyl-naphthalene bond lead to atropisomers. The methylene groups (on both the methylenedioxy and lactone groups) give rise to AB patterns, rather than singlets, since the two protons are diastereotopic when rotation is slow.116
Atropisomers are stereoisomers arising because of hindered rotation about a single bond, where energy differences due to steric strain or other contributors create a barrier to rotation that is high enough to allow for isolation of individual conformers (Fig. 28).
Aryltetralin lignans are similar to diarylbutane lignans, but they differ in the six-membered ring connecting C-6 and C-7′, or have additional γ-butyrolactone ring. Therefore, side chains usually are -CH2-CH-CH3, -CH-CH-CH3. The structure of a representative aryltetralin lignan is shown below (Fig. 29). 13C-NMR data was described in Table 6.105
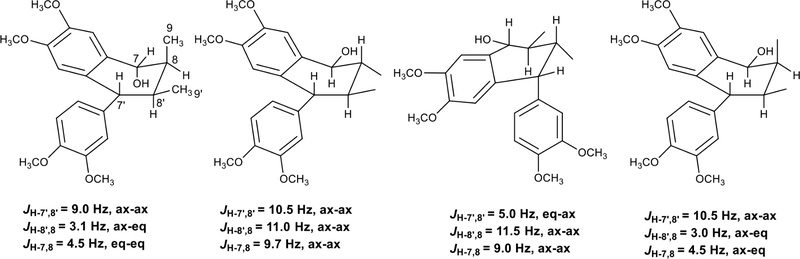
Holostylol A and its conformers were obtained from Holostylis reniformis.117 Their chemical structures of determined by NMR and CD spectroscopic methods. The 1H- and 13C- NMR spectra showed signals for two aromatic rings signals, four methine carbons, and two methyl groups (Fig. 30). The position of methine hydrogens were deduced by coupling constants between the methine hydrogens (JH-7′,8′ = 9.0Hz, JH-8′,8 = 3.1Hz, JH-7,8 = 4.5Hz), which suggested a trans di-axial orientation of H-7′, H-8′, a cis axial-equatorial orientation of H-8, H-8′, and a trans di-equatorial orientation of H-7, H-8 (Fig. 30). NOESY correlations also confirmed its spatial structures. H-7 correlated with H-8, CH3-9. H-8′ correlated with CH3-9′, H-8, H-6′, and H-2′.
The 7′R configuration. The positive Cotton effects at ̇λ≅289 nm characteristic of an α-aryl at C-7′ (7′R) of these derivatives also indicates 7′R configuration of holostylol A.118 The absolute configuration of holostylol A was determined to be 7R,7′R,8S,8′S.
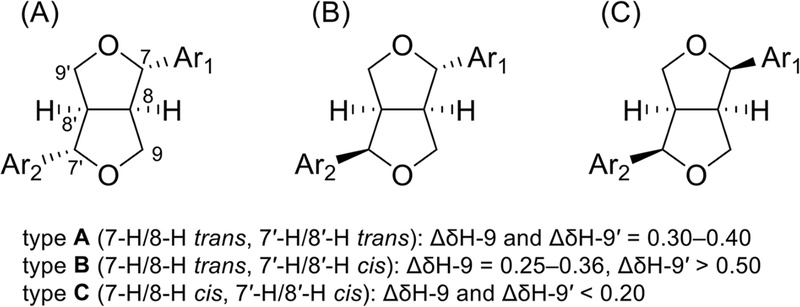
Furofuran derivatives – Furofuran lignans are an important group of molecules with a characteristic 2,6-diaryl-3,7-dioxabicyclo[3.3.0]octane skeleton, so they possess a 7,9′:7′,9-diepoxy moiety. Their side chains are composed of two set -OCH2-CH-CH2O-, which linked at 8-8′ to form furofuran derivatives. Therefore, side chains of furofuran lignans (H-8/H-8′ type) showed two oxygenated methine signals, and two oxygenated methylene signals, and two methine signals in 1H- and 13C-NMR. Furofuran lignans with 8-OH group exhibited two oxygenated methine signals, two oxygenated methylene signals, and one methine signal in 1H-NMR. Additional oxygenated quaternary carbon was shown in 13C-NMR.
The relative configuration of H-8/H-8′ type furofuran lignans (Fig. 31) were classified three types: (A) H-7/H-8 trans, H-7′/H-8′ trans (1–3); (B) H-7/H-8 trans, H-7′/H-8′ cis; and (C) H-7/H-8 cis, H-7′/H-8′ cis. Chemical shift differences (ΔδH-9 and ΔδH-9′) of two methylenes (H-9 and H-9′) can be used to determine the relative configuration of C-7/C-8 and C-7′/C-8′.119 13C-NMR data furofuran parts were described in the Table 7.105
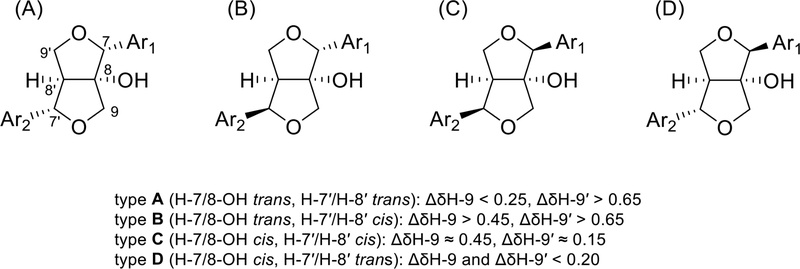
Furofuran lignans with 8-OH (Fig. 32) were classified into four groups according to the orientations of the two aromatic rings: (A) H-7/8-OH trans, H-7′/8′-H trans; (B) H-7/8-OH trans, H-7′/H-8′ cis; (C) H-7/8-OH cis, H-7′/H-8′ cis; and (D) H-7/8-OH cis, H-7′/H-8′ trans.119
The relative stereochemistry was also confirmed through NOESY experiment and the absolute stereochemistry was determined by comparing the specific optical rotation or ECD spectra.120
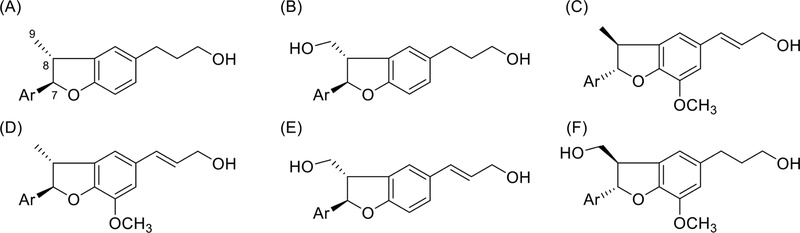
Benzofuran neolignans – Dihydrobenzofuran neolignans contain two C6C3 units displaying 7-O-4′ and 8-3′ (or 5′) linkages (Fig. 33). The characteristic UV absorbance was observed at 207–212 nm, 228–233 nm and 280–285 nm. Spectroscopic characteristic data of dihydrobenzofuran lignans were set of H-7/C7 (δH 5.07–5.13 1H, d, J = 8.8–9.6 Hz / δC 92.5–93.2), H-8/C-8 (δ 3.40–3.46 ,1H/ δC 45.2-45.8) and CH3-9 (δH 1.37–1.41, 3H, d, J = 6.8 Hz / δC 17.0–17.7). When 9-CH3 changed to 9-CH2OH, 13C-NMR data was observed at C-8 (δC 52.9-55.2) and C-9 (δC 62.9-64.5).121 13C-NMR chemical shift ranges for the resonances belonging to C-7 to C-9 positions dihyrobenzofuran lignans (Table 8).105
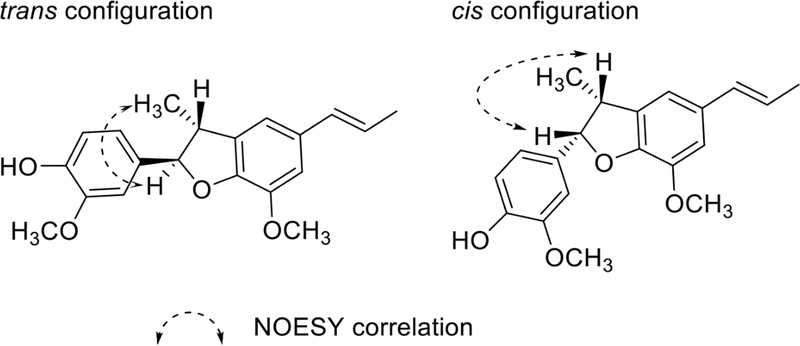
The coupling constant between H-7 and H-8 indicates the relative configuration (usually, J = 9.5 Hz, trans; J = 8.5 Hz, cis) of C-7 and C-8, but small J value difference, NOE (or NOESY) required to the determine relative configuration of C-7 and C-8. NOE correlation of H-7/H-8 and no correlation H-7/CH3-9 indicates cis relative configuration. In case of trans configuration, correlation H-7/CH3-9 was observed in NOESY (Fig. 34).121–123
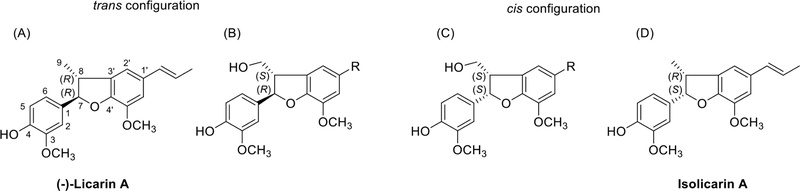
The absolute configuration at C-7 and C-8 were determined by the circular dichroism and optical rotation properties (Fig. 35). If methyl group at C-9 position changed into CH2OH, C-8 configuration was changed due to priority.123 In case of C-9 methyl, trans enantiomer, negative Cotton effect ~ 230 nm and positive Cotton effect ~ 280 nm (Table 9), the absolute configuration at C-7, C-8 were 7S,8S.121,124
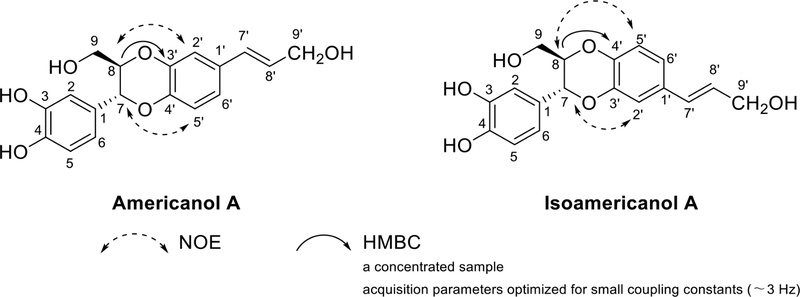
Benzodioxane neolignans – A benzodioxane-type lignan is formed by connecting the C-7 and C-8 of one C6C3 unit to the aryl ring carbon of another C6C3 unit via two oxygen atoms. Americanol A and isoamericanol A, structural isomers, were isolated from Phytolacca americana,125 therefore these two compounds showed similar NMR spectral data, but characteristically different shifts for H-2′ and H-5′ as measured in CD3OD, indicating a distinction between americinol and isoamericinol types of neolignans: in the spectrum of americanol A, a relatively large difference of 0.2 ppm between the resonances of H-2′ and H-5′, whereas the corresponding difference in the spectrum of isoamericanol A is only 0.07 ppm. NOE effects of H-2′/H-8 and H-5′/H-7 were shown in americanol, and NOEs were found between H-2′ and H-7, and between H-5′ and H-8 in isoamericanol (Fig. 36). In addition, HMBC spectra using a concentrated sample and acquisition parameters optimized for small coupling constants (~ 3 Hz), showed a correlation between H-8 and C-3′ for americanol and a correlation between H-8 and C-4′ for isoamericanol.126
The relative configuration of H-7 and H-8 can be determined by coupling constant: the trans isomer has a ~ 8 Hz coupling constant, while the coupling constant for the cis isomer is ~ 2 Hz. The assignment of absolute stereochemistry was determined by optical rotation and ECD (or) curve. When looking at the ECD spectra of the trans compounds, in particular the peak at ~ 250 nm; the 7S,8S enantiomer showed a positive curve and the 7R,8R enantiomer showed a negative curve in this region. In case of cis compounds, positive curve at ~ 250 nm is 7S,8R configuration and negative curve is 7R,8S configuration. Optical rotations also be used to determine the C-7 and C-8 configuration of benzodioxane-type lignans.127–129
In conclusion, recent advancements in analytical and preparative techniques have significantly enhanced the ability to isolate, purify, and structurally characterize lignans. Moving forward, greater standardization of extraction protocols, improved methods for chiral separation, and the integration of advanced spectroscopic tools will further strengthen lignan research. These methodological developments promise to deepen understanding of lignan bioactivity and facilitate their application as therapeutic, nutritional, and agrochemical agents. It is hoped that this review will guide researchers who are new to the field and stimulate continued innovation in the study and utilization of lignans.
Acknowledgments
This study was supported by Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (RS-2023-00245441 and NRF-2020R1A6A1A03042854).
Conflicts of Interest
The authors declare that they have no conflicts of interest.
References
-
Bagniewska-Zadworna, A.; Barakat, A.; Łakomy, P.; Smoliński, D. J.; Zadworny, M. Plant Sci. 2014, 229, 111–121.
[https://doi.org/10.1016/j.plantsci.2014.08.015]

-
Landete, J. M. Food Res. Int. 2012, 46, 410–424.
[https://doi.org/10.1016/j.foodres.2011.12.023]

-
Holmbom, B.; Eckerman, C.; Eklund, P.; Hemming, J.; Nisula, L.; Reunanen, M.; Sjöholm, R.; Sundberg, A.; Sundberg, K.; Willför, S. Phytochem. Rev. 2003, 2, 331–340.
[https://doi.org/10.1023/B:PHYT.0000045493.95074.a8]

-
Cornwell, T.; Cohick, W.; Raskin, I. Phytochemistry 2004, 65, 995–1016.
[https://doi.org/10.1016/j.phytochem.2004.03.005]

-
Li, J.; Ma, X.; Luo, L.; Tang, D.; Zhang, L. J. Agric. Food Chem. 2023. 71, 16419–16434.
[https://doi.org/10.1021/acs.jafc.3c02680]

-
Cederroth, C. R.; Auger, J.; Zimmermann, C.; Eustache, F.; Nef, S. Int. J. Androl. 2010, 33, 304–316.
[https://doi.org/10.1111/j.1365-2605.2009.01011.x]

-
Adlercreutz, H. Crit. Rev. Clin. Lab. Sci. 2007, 44, 483–525.
[https://doi.org/10.1080/10408360701612942]

-
Peterson, J.; Dwyer, J.; Adlercreutz, H.; Scalbert, A.; Jacques, P.; McCullough, M. L. Nutr. Rev. 2010, 68, 571–603.
[https://doi.org/10.1111/j.1753-4887.2010.00319.x]

-
Wang, L.-X.; Wang, H.-L.; Huang, J.; Chu, T.-Z.; Peng, C.; Zhang, H.; Chen, H.-L.; Xiong, Y.-A.; Tan, Y.-Z. Phytochemistry 2022, 202, 113326.
[https://doi.org/10.1016/j.phytochem.2022.113326]

-
Moss, G. P. Pure Appl. Chem. 2000, 72, 1493–1523.
[https://doi.org/10.1351/pac200072081493]

-
Davin, L. B.; Lewis, N. G. Phytochem. Rev. 2003, 2, 257–288.
[https://doi.org/10.1023/B:PHYT.0000046175.83729.b5]

-
Satake, H.; Ono, E.; Murata, J. J. Agric. Food Chem. 2013, 61, 11721–11729.
[https://doi.org/10.1021/jf4007104]

-
Satake, H.; Koyama, T.; Bahabadi, S. E.; Matsumoto, E.; Ono, E.; Murata, J. Metabolites 2015. 5, 270–290.
[https://doi.org/10.3390/metabo5020270]

-
Morimoto, K.; Satake, H. Biol. Pharm. Bull. 2013, 36, 1519–1523.
[https://doi.org/10.1248/bpb.b13-00437]

-
Kim, H. J.; Ono, E.; Morimoto, K.; Yamagaki, T.; Okazawa, A.; Kobayashi, A.; Satake, H. Plant Cell Physiol. 2009, 50, 2200–2209.
[https://doi.org/10.1093/pcp/pcp156]

-
Umezawa, T.; Ragamustari, S. K.; Nakatsubo, T.; Wada, S.; Li, L.; Yamamura, M.; Sakakibara, N.; Hattori, T.; Suzuki, S.; Chiang, V. L. Plant Biotechnol. 2013, 30, 97–109.
[https://doi.org/10.5511/plantbiotechnology.12.1230a]

-
Ragamustari, S. K.; Yamamura, M.; Ono, E.; Hattori, T.; Suzuki, S.; Suzuki, H.; Shibata, D.; Umezawa, T. Plant Biotechnol. 2014, 31, 257–267.
[https://doi.org/10.5511/plantbiotechnology.14.0722a]

-
Umezawa, T. Phytochem. Rev. 2003, 2, 371–390.
[https://doi.org/10.1023/B:PHYT.0000045487.02836.32]

-
Ragamustari, S. K.; Nakatsubo, T.; Hattori, T.; Ono, E.; Kitamura, Y.; Suzuki, S.; Yamamura, M.; Umezawa, T. Plant Biotechnol. 2013, 30, 375–384.
[https://doi.org/10.5511/plantbiotechnology.13.0527b]

-
Noguchi, A.; Fukui, Y.; Iuchi‐Okada, A.; Kakutani, S.; Satake, H.; Iwashita, T.; Nakao, M.; Umezawa, T.; Ono, E. Plant J. 2008, 54, 415–427.
[https://doi.org/10.1111/j.1365-313X.2008.03428.x]

-
Jang, W. Y.; Kim, M.-Y.; Cho, J. Y. Int. J. Mol. Sci. 2022, 23, 15482.
[https://doi.org/10.3390/ijms232415482]

-
Tsai, W.-J.; Shen, C.-C.; Tsai, T.-H.; Lin, L.-C. J. Nat. Prod. 2014, 77, 125–131.
[https://doi.org/10.1021/np400772h]

-
Sukbangnop, W.; Hosen, A.; Hongthong, S.; Kuhakarn, C.; Tuchinda, P.; Chaturonrutsamee, S.; Thanasansurapong, S.; Akkarawongsapat, R.; Limthongkul, J.; Napaswad, C. Fitoterapia 2021, 151, 104885.
[https://doi.org/10.1016/j.fitote.2021.104885]

-
Yang, Y.-N.; Huang, X.-Y.; Feng, Z.-M.; Jiang, J.-S.; Zhang, P.-C. J. Agric. Food Chem. 2014, 62, 9095–9102.
[https://doi.org/10.1021/jf501859x]

-
Sun, Y.-J.; Li, Z.-L.; Chen, H.; Liu, X.-Q.; Zhou, W.; Hua, H.-M. Planta Med. 2012, 78, 480–484.
[https://doi.org/10.1055/s-0031-1298201]

-
Jin, H.; Yin, H.-L.; Liu, S.-J.; Chen, L.; Tian, Y.; Li, B.; Wang, Q.; Dong, J.-X. Fitoterapia 2014, 94, 70–76.
[https://doi.org/10.1016/j.fitote.2014.01.025]

-
Fukamiya, N.; Lee, K.-H. J. Nat. Prod. 1986, 49, 348–350.
[https://doi.org/10.1021/np50044a030]

-
Asano, J.; Chiba, K.; Tada, M.; Yoshii, T. Phytochemistry 1996, 42, 713–717.
[https://doi.org/10.1016/0031-9422(96)00024-6]

-
Chen, C.-C.; Hsin, W.-C.; Ko, F.-N.; Huang, Y.-L.; Ou, J.-C.; Teng, C.-M. J. Nat. Prod. 1996, 59, 1149–1150.
[https://doi.org/10.1021/np960443+]

-
Liu, B.; Zhang, T.; Xie, Z.-T.; Hong, Z.-C.; Lu, Y.; Long, Y.-M.; Ji, C.-Z.; Liu, Y.-T.; Yang, Y.-F.; Wu, H.-Z. J. Ethnopharmacol. 2022, 294, 115392.
[https://doi.org/10.1016/j.jep.2022.115392]

-
Hemmati, S.; Seradj, H. Molecules 2016, 21, 820.
[https://doi.org/10.3390/molecules21070820]

-
Trinh Thi Thanh, V.; Cuong Pham, V.; Doan Thi Mai, H.; Litaudon, M.; Guéritte, F.; Retailleau, P.; Nguyen, V. H.; Chau, V. M. J. Nat. Prod. 2012, 75, 1578–1583.
[https://doi.org/10.1021/np3003832]

-
Yang, Y.-N.; Huang, X.-Y.; Feng, Z.-M.; Jiang, J.-S.; Zhang, P.-C. J. Agric. Food Chem. 2015, 63, 7958–7966.
[https://doi.org/10.1021/acs.jafc.5b02838]

-
Liu, S.-Q.; Yang, Y.-P.; Hussain, N.; Jian, Y.-Q.; Li, B.; Qiu, Y.-X.; Yu, H.-H.; Wang, H.-Z.; Wang, W. Pharmacol. Res. 2023, 195, 106872.
[https://doi.org/10.1016/j.phrs.2023.106872]

-
Yang, S.; Yuan, C. Arab. J. Chem. 2021, 14, 103310.
[https://doi.org/10.1016/j.arabjc.2021.103310]

-
Sok, D.-E.; Cui, H. S.; Kim, M. R. Recent Pat. Food Nutr. Agric. 2009, 1, 87–95.
[https://doi.org/10.2174/2212798410901010087]

-
Heinonen, S.; Nurmi, T.; Liukkonen, K.; Poutanen, K.; Wähälä, K.; Deyama, T.; Nishibe, S.; Adlercreutz, H. J. Agric. Food Chem. 2001, 49, 3178–3186.
[https://doi.org/10.1021/jf010038a]

-
Kim, Y.; Kim, H.-W.; Sung, J.; Kim, Y. Front. Nutr. 2024, 11, 1409309.
[https://doi.org/10.3389/fnut.2024.1409309]

-
Ladeira Macedo, A.; Carvalho Costa dos Santos, T.; Leda Valverde, A.; de Lima Moreira, D.; Rocha Alves Vasconcelos, T. Mini Rev. Med. Chem. 2017, 17, 693–720.
[https://doi.org/10.2174/1389557516666161130094826]

-
Shen, T.; Hussaini, I. M. In methods in enzymology: Kadsurenone and other related lignans as antagonists of platelet-activating factor receptor; Colowick, S. P.; Kaplan, N. O. Ed; Elsevier; United States of America, 1990, pp 446–454.
[https://doi.org/10.1016/0076-6879(90)87051-4]

-
Yeh, L.-A.; Chen, J.; Baculi, F.; Gingrich, D. E.; Shen, T. Y. Bioorg. Med. Chem. Lett. 1995, 5, 1637–1642.
[https://doi.org/10.1016/0960-894X(95)00282-X]

-
Chang, H.-Y.; Chen, Y.-C.; Lin, J.-G.; Lin, I.-H.; Huang, H.-F.; Yeh, C.-C.; Chen, J.-J.; Huang, G.-J. Am. J. Chin. Med. 2018, 46, 651–671.
[https://doi.org/10.1142/S0192415X18500349]

-
Muraoka, O.; Zheng, B.-Z.; Fujiwara, N.; Tanabe, G. J. Chem. Soc. Perkin 1 1996, 405–411.
[https://doi.org/10.1039/P19960000405]

-
Suzuki, S.; Umezawa, T. J. Wood Sci. 2007, 53, 273–284.
[https://doi.org/10.1007/s10086-007-0892-x]

-
Jin, C. J.; Yu, S. H.; Wang, X.-M.; Woo, S. J.; Park, H. J.; Lee, H. C.; Choi, S. H.; Kim, K. M.; Kim, J. H.; Park, K. S. PLoS One 2014, 9, e98232.
[https://doi.org/10.1371/journal.pone.0098232]

-
Chan, Y.-S.; Cheng, L.-N.; Wu, J.-H.; Chan, E.; Kwan, Y.-W.; Lee, S. M.-Y.; Leung, G. P.-H.; Yu, P. H.-F.; Chan, S.-W. Inflammopharmacology 2011, 19, 245–254.
[https://doi.org/10.1007/s10787-010-0062-4]

-
Vargas-Mendoza, N.; Morales-González, Á.; Morales-Martínez, M.; Soriano-Ursúa, M. A.; Delgado-Olivares, L.; Sandoval-Gallegos, E. M.; Madrigal-Bujaidar, E.; Álvarez-González, I.; Madrigal-Santillán, E.; Morales-Gonzalez, J. A. Biomedicines 2020, 8, 122.
[https://doi.org/10.3390/biomedicines8050122]

-
Ray, A. B.; Chattopadhyay, S. K.; Kumar, S.; Konno, C.; Kiso, Y.; Hikino, H. Tetrahedron 1985, 41, 209–214.
[https://doi.org/10.1016/S0040-4020(01)83488-8]

-
Fernandas, E. R.; Carvalho, F. D.; Remião, F. G.; Bastos, M. L.; Pinto, M. M.; Gottlieb, O. R. Pharm. Res. 1995, 12, 1756–1760.
[https://doi.org/10.1023/A:1016230125496]

-
Yang, Y.-T.; Zhang, Y.; Bian, Y.; Zhu, J.; Feng, X.-S. Food Chem. X 2025, 26, 102249.
[https://doi.org/10.1016/j.fochx.2025.102249]

-
Willför, S.; Smeds, A. I; Holmbom, B. R. J. Chromatogr. A 2006, 1112, 64–77.
[https://doi.org/10.1016/j.chroma.2005.11.054]

-
Willför, S.; Hemming, J.; Reunanen, M.; Eckerman, C.; Holmbom, B. Holzforschung 2003, 57, 27–36.
[https://doi.org/10.1515/HF.2003.005]

-
Willför, S.; Hemming, J.; Reunanen, M.; Holmbom, B. Holzforschung 2003, 57, 359–372.
[https://doi.org/10.1515/HF.2003.054]

-
Willför, S. M.; Ahotupa, M. O.; Hemming, J. E.; Reunanen, M. H. T.; Eklund, P. C.; Sjöholm, R. E.; Eckerman, C. S. E.; Pohjamo, S. P.; Holmbom, B. R. J. Agric. Food Chem. 2003. 51, 7600–7606.
[https://doi.org/10.1021/jf030445h]

-
Willför, S.; Nisula, L.; Hemming, J.; Reunanen, M.; Holmbom, B. Holzforschung 2004, 58, 650–659.
[https://doi.org/10.1515/HF.2004.119]

-
Willför, S.; Nisula, L.; Hemming, J.; Reunanen, M.; Holmbom, B. Holzforschung 2004, 58, 335–344.
[https://doi.org/10.1515/HF.2004.052]

-
Hu, Y.; Tse, T. J.; Shim, Y. Y.; Purdy, S. K.; Kim, Y. J.; Meda, V.; Reaney, M. J. Crit. Rev. Food Sci. Nutr. 2024, 64, 5057–5072.
[https://doi.org/10.1080/10408398.2022.2148627]

-
Kamal-Eldin, A.; Peerlkamp, N.; Johnsson, P.; Andersson, R.; Andersson, R. E.; Lundgren, L. N.; Åman, P. Phytochemistry 2001, 58, 587–590.
[https://doi.org/10.1016/S0031-9422(01)00279-5]

-
Johnsson, P.; Kamal-Eldin, A.; Lundgren, L. N.; Åman, P. J. Agric. Food Chem. 2000, 48, 5216–5219.
[https://doi.org/10.1021/jf0005871]

-
Tura, D.; Robards, K. J. Chromatogr. A 2002, 975, 71–93.
[https://doi.org/10.1016/S0021-9673(02)00879-8]

-
Slanina, J.; Glatz, Z. J. Chromatogr. B Analyt. Technol. Biomed. Life Sci. 2004, 812, 215–229.
[https://doi.org/10.1016/S1570-0232(04)00548-3]

-
Dai, Z.; Xin, H.; Fu, Q.; Hao, H.; Li, Q.; Liu, Q.; Jin, Y. J. Sep. Sci. 2019, 42, 2444–2454.
[https://doi.org/10.1002/jssc.201900222]

-
Patyra, A.; Kołtun-Jasion, M.; Jakubiak, O.; Kiss, A. K. Plants 2022, 11, 2323.
[https://doi.org/10.3390/plants11172323]

-
Beejmohun, V.; Fliniaux, O.; Grand, É.; Lamblin, F.; Bensaddek, L.; Christen, P.; Kovensky, J.; Fliniaux, M.-A.; Mesnard, F. Phytochem. Anal. 2007, 18, 275–282.
[https://doi.org/10.1002/pca.973]

-
Wang, X.; Lin, Y.; Geng, Y.; Li, F.; Wang, D. Cereal Chem. 2009, 86, 23–25.
[https://doi.org/10.1094/CCHEM-86-1-0023]

-
Dar, A. A.; Arumugam, N. Bioorg. Chem. 2013, 50, 1–10.
[https://doi.org/10.1016/j.bioorg.2013.06.009]

-
Reshma, M. V.; Balachandran, C.; Arumughan, C.; Sunderasan, A.; Sukumaran, D.; Thomas, S.; Saritha, S. S. Food Chem. 2010, 120, 1041–1046.
[https://doi.org/10.1016/j.foodchem.2009.11.047]

-
Sun, Y.; Yu, Z.; Duan, W.; Fang, L.; Xu, S.; Wang, X. Chromatogr. B Analyt. Technol. Biomed. Life Sci. 2011, 879, 3775–3779.
[https://doi.org/10.1016/j.jchromb.2011.10.023]

-
Jiang, J.; Dong, H.; Wang, T.; Zhao, R.; Mu, Y.; Geng, Y.; Zheng, Z.; Wang, X. Molecules 2017, 22, 2024.
[https://doi.org/10.3390/molecules22122024]

-
Zhu, L.; Li, B.; Liu, X.; Huang, G.; Meng, X. Food Chem. 2015, 186, 146–152.
[https://doi.org/10.1016/j.foodchem.2014.09.008]

-
Chu, C.; Zhang, S.; Tong, S.; Li, X.; Yan, J. J. Sep. Sci. 2013. 36, 3958–3964.
[https://doi.org/10.1002/jssc.201300896]

-
Kim, C. Y.; Ahn, M.-J.; Kim, J. J. Sep. Sci. 2006, 29, 656–659.
[https://doi.org/10.1002/jssc.200500316]

-
Jeon, J.-S.; Park, C. L.; Syed, A. S.; Kim, Y.-M.; Cho, I. J.; Kim, C.Y. Chromatogr. B Analyt. Technol. Biomed. Life Sci. 2016, 1011, 108–113.
[https://doi.org/10.1016/j.jchromb.2015.12.062]

-
Wang, P.; Liu, Y.; Chen, T.; Xu, W.; You, J.; Liu, Y.; Li, Y. Phytochem. Anal. 2013, 24, 603–607.
[https://doi.org/10.1002/pca.2438]

-
Eklund, P. C.; Sundell, F. J.; Smeds, A. I.; Sjöholm, R. E. Org. Biomol. Chem. 2004, 2, 2229–2235.
[https://doi.org/10.1039/B402849J]

-
Eklund, P. C.; Willför, S. M.; Smeds, A. I.; Sundell, F. J.; Sjöholm, R. E.; Holmbom, B. R. J. Nat. Prod. 2004, 67, 927–931.
[https://doi.org/10.1021/np0340706]

-
Lee, H. J.; Kim, C. Y. Food Chem. 2010, 120, 1224–1228.
[https://doi.org/10.1016/j.foodchem.2009.11.068]

-
Zhang, J.-Y.; Mei, J.-W.; Wang, H.-Y.; Xu, Z.-H. Phytochem. Anal. 2022, 33, 1214–1224.
[https://doi.org/10.1002/pca.3172]

-
Bose, S.; Munsch, T.; Lanoue, A.; Garros, L.; Tungmunnithum, D.; Messaili, S.; Destandau, E.; Billet, K.; St-Pierre, B.; Clastre, M. Front. Plant Sci. 2020, 11, 508658.
[https://doi.org/10.3389/fpls.2020.508658]

-
Schmidt, T. J.; Hemmati, S.; Fuss, E.; Alfermann, A.W. Phytochem. Anal. 2006, 17, 299–311.
[https://doi.org/10.1002/pca.918]

-
Liu, H.; Zhang, J.; Li, X.; Qi, Y.; Peng, Y.; Zhang, B.; Xiao, P. Phytomedicine 2012, 19, 1234–1241.
[https://doi.org/10.1016/j.phymed.2012.07.017]

- He, F.; Dou, D.-Q.; Sun, Y.; Zhu, L.; Xiao, H.-B.; Kang, T.-G. Planta Med. 2012, 78, 800–806.
-
Wang, W.; Pan, Q.; Han, X.-Y.; Wang, J.; Tan, R.-Q.; He, F.; Dou, D.-Q.; Kang, T.-G. Fitoterapia 2013, 86, 6–12.
[https://doi.org/10.1016/j.fitote.2013.01.016]

-
Yang, J.-M.; IP, S.-P. P.; Yeung, H.-K. J.; Che, C.-T. Acta Pharm. Sin. B 2011, 1, 46–55.
[https://doi.org/10.1016/j.apsb.2011.04.007]

-
Rönquist-Nii, Y.; Eksborg, S.; Axelson, M.; Harmenberg, J.; Ekman, S.; Bergqvist, M.; Beck, O. Chromatogr. B Analyt. Technol. Biomed. Life Sci. 2011. 879, 326–334.
[https://doi.org/10.1016/j.jchromb.2010.12.017]

-
Sheng, Y.-L.; Xu, J.-H.; Shi, C.-H.; Li, W.; Xu, H.-Y.; Li, N.; Zhao, Y.-Q.; Zhang, X.-R. J. Ethnopharmacol. 2014, 155, 1568–1574.
[https://doi.org/10.1016/j.jep.2014.07.052]

-
Tomimori, N.; Tanaka, Y.; Kitagawa, Y.; Fujii, W.; Sakakibara, Y.; Shibata, H. Biopharm. Drug Dispos. 2013, 34, 462–473.
[https://doi.org/10.1002/bdd.1862]

-
Halls, S. C.; Lewis, N. G. Tetrahedron 2003, 14, 649–658.
[https://doi.org/10.1016/S0957-4166(03)00099-5]

-
Wu, Z.; Lai, Y.; Zhou, L.; Wu, Y.; Zhu, H.; Hu, Z.; Yang, J.; Zhang, J.; Wang, J.; Luo, Z. Xue, Y.; Zhang, Y. Sci. Rep. 2016, 6, 24809.
[https://doi.org/10.1038/srep24809]

-
Wang, J.; Zhou, L.; Cheng, Z.-Y.; Wang, Y.-X.; Yan, Z.-Y.; Huang, X.-X.; Song, S.-J. Nat. Prod. Res. 2020, 34, 2225–2228.
[https://doi.org/10.1080/14786419.2019.1577839]

-
Pereira, A. C.; Magalhães, L. G.; Gonçalves, U. O.; Luz, P. P.; Moraes, A. C. G.; Rodrigues, V.; da Matta Guedes, P. M.; da Silva Filho, A. A.; Cunha, W. R.; Bastos, J. K. Nanayakkara, N. P. D.; e Silva, M. L. A. Phytochemistry 2011, 72, 1424–1430.
[https://doi.org/10.1016/j.phytochem.2011.04.007]

-
Moinuddin, S. G. A.; Hishiyama, S.; Cho, M.-H.; Davin, L. B.; Lewis, N. G. Org. Biomol. Chem. 2003, 1, 2307–2313.
[https://doi.org/10.1039/B302632A]

-
Nakatani, N.; Ikeda, K.; Kikuzaki, H.; Kido, M.; Yamaguchi, Y. Phytochemistry 1988, 27, 3127–3129.
[https://doi.org/10.1016/0031-9422(88)80013-X]

-
Kubanek, J.; Fenical, W.; Hay, M. E.; Brown, P. J.; Lindquist, N. Phytochemistry 2000, 54, 281–287.
[https://doi.org/10.1016/S0031-9422(00)00076-5]

-
Njoku, C. J.; Hopp, D. C.; Alali, F.; Asuzu, I. U.; McLaughlin, J. L. Planta Med. 1997, 63, 580–581.
[https://doi.org/10.1055/s-2006-957779]

-
Moon, T. C.; Seo, C. S.; Haa, K.; Kim, J. C.; Hwang, N. K.; Hong, T. G.; Kim, J. H.; Kim, D. H.; Son, J. K.; Chang, H. W. Arch. Pharm. Res. 2008, 31, 606–610.
[https://doi.org/10.1007/s12272-001-1200-y]

-
Wang, Q.; Yang, Y.; Li, Y.; Yu, W.; Hou, Z. J. Tetrahedron 2006, 62, 6107–6112.
[https://doi.org/10.1016/j.tet.2006.03.111]

-
Kawaguchi, Y.; Yamauchi, S.; Masuda, K.; Nishiwaki, H.; Akiyama, K.; Maruyama, M.; Sugahara, T.; Kishida, T.; Koba, Y. Biosci. Biotechnol. Biochem. 2009, 73, 1806–1810.
[https://doi.org/10.1271/bbb.90167]

-
Fang, J.-M.; Hsu, K.-C.; Cheng, Y.-S. Phytochemistry 1989, 28, 3553–3555.
[https://doi.org/10.1016/0031-9422(89)80393-0]

-
Wukirsari, T.; Nishiwaki, H.; Hasebe, A.; Shuto, Y.; Yamauchi, S. J. Agric. Food Chem. 2013, 61, 4318–4325.
[https://doi.org/10.1021/jf400300n]

-
Tomioka, K.; Ishiguro, T.; Koga, K. Chem. Pharm. Bull. 1985, 33, 4333–4337.
[https://doi.org/10.1248/cpb.33.4333]

-
Nam, A.-M.; Paoli, M.; Castola, V.; Casanova, J.; Bighelli, A. Nat. Prod. Commun. 2011, 6, 379–385.
[https://doi.org/10.1177/1934578X1100600316]

-
Consonni, R.; Ottolina, G. Molecules 2022, 27, 2340.
[https://doi.org/10.3390/molecules27072340]

-
Nhiem, N. X.; Lee, H. Y.; Kim, N. Y.; Park, S. J.; Kim, E. S.; Han, J. E.; Yang, H.; Kim, S. H. Magn. Reson. Chem. 2012, 50, 772–777.
[https://doi.org/10.1002/mrc.3875]

-
Agrawal, P. K.; Pathak, A. K. Magn. Reson. Chem. 1994, 32, 753–773.
[https://doi.org/10.1002/mrc.1260321210]

-
Solyomváry, A.; Beni, S.; Boldizsar, I. Mini Rev. Med. Chem. 2017, 17, 1053–1074.
[https://doi.org/10.2174/1389557516666160614005828]

-
Umehara, K.; Sugawa, A.; Kuroyanagi, M.; Ueno, A.; Taki, T. Chem. Pharm. Bull. 1993, 41, 1774–1779.
[https://doi.org/10.1248/cpb.41.1774]

-
Ekholm, F. S.; Eklund, P.; Leino, R. Carbohydr. Res. 2010, 345, 1963–1967.
[https://doi.org/10.1016/j.carres.2010.06.008]

-
Li, N.; Wu, J.-L.; Sakai, J.-I.; Ando, M. J. Nat. Prod. 2003, 66, 1421–1426.
[https://doi.org/10.1021/np030247k]

-
Chang, J.; Reiner, J.; Xie, J. Chem. Rev. 2005, 105, 4581–4609.
[https://doi.org/10.1021/cr050531b]

-
Zhang, L.; Jia, Y.-Z.; Li, B.; Peng, C.-Y.; Yang, Y.-P.; Wang, W.; Liu, C.-X. Chin. Herb. Med. 2021, 13, 157–166.
[https://doi.org/10.1016/j.chmed.2021.01.005]

-
Ikeya, Y.; Taguchi, H.; Sasaki, H.; Nakajima, K.; Yosioka, I. Chem. Pharm. Bull. 1980, 28, 2414–2421.
[https://doi.org/10.1248/cpb.28.2414]

-
Hu, D.; Yang, Z.; Yao, X.; Wang, H.; Han, N.; Liu, Z.; Wang, Y.; Yang, J.; Yin, J. Phytochemistry 2014, 104, 72–78.
[https://doi.org/10.1016/j.phytochem.2014.04.014]

-
Rangkaew, N.; Suttisri, R.; Moriyasu, M.; Kawanishi, K. Fitoterapia 2009, 80, 377–379.
[https://doi.org/10.1016/j.fitote.2009.05.005]

-
Park, S.; Kim, S.; Shin, D. Phytochem. Rev. 2021, 20, 1033–1054.
[https://doi.org/10.1007/s11101-020-09735-z]

-
Charlton, J. L.; Oleschuk, C. J.; Chee, G.-L. J. Org. Chem. 1996, 61, 3452–3457.
[https://doi.org/10.1021/jo952048e]

-
Da Silva, T.; Lopes, L. M. Phytochemistry 2006, 67, 929–937.
[https://doi.org/10.1016/j.phytochem.2006.02.007]

-
Lopes, L. M. X.; Yoshida, M.; Gottlieb, O. R. Phytochemistry 1982, 21, 751–755.
[https://doi.org/10.1016/0031-9422(82)83181-6]

-
Shao, S.-Y.; Yang, Y.-N.; Feng, Z.-M.; Jiang, J.-S.; Zhang, P.-C. J. Nat. Prod. 2018. 81, 1023–1028.
[https://doi.org/10.1021/acs.jnatprod.8b00044]

-
Liu, W.-J.; Chen, Y.-J.; Chen, D.-N.; Wu, Y.-P.; Gao, Y.-J.; Li, J.; Zhong, W.-J.; Jiang, L. Nat. Prod. Res. 2018, 32, 933–938.
[https://doi.org/10.1080/14786419.2017.1371163]

-
Sawasdee, K.; Chaowasku, T.; Lipipun, V.; Dufat, T.-H.; Michel, S.; Likhitwitayawuid, K. Fitoterapia 2013, 85, 49–56.
[https://doi.org/10.1016/j.fitote.2012.12.028]

-
Li, F.; Yang, X.-W. Helv. Chim. Acta 2007, 90, 1491–1496.
[https://doi.org/10.1002/hlca.200790155]

-
Zhang, D.-D.; Zhao, P.; Huang, S.-W.; Song, S.-J.; Huang, X.-X. Nat. Prod. Res. 2023, 37, 1349–1355.
[https://doi.org/10.1080/14786419.2021.2007094]

-
Chen, H.; Zhu, Y.; Zhang, Y.-L.; Zeng, M.-N.; Cao, Y.-G.; Sun, P.-T.; Cao, B.; Du, K.; Zhao, X.; Wang, X.-W.; Zheng, Z.-K.; Feng, W.-S. Phytochemistry 2022, 203, 113336.
[https://doi.org/10.1016/j.phytochem.2022.113336]

-
Takahasi, H.; Yanagi, K.; Ueda, M.; Nakade, K.; Fukuyama, Y. Chem. Pharm. Bull. 2003, 51, 1377–1381.
[https://doi.org/10.1248/cpb.51.1377]

-
Waibel, R.; Benirschke, G.; Benirschke, M.; Achenbach, H. Phytochemistry 2003, 62, 805–811.
[https://doi.org/10.1016/S0031-9422(02)00357-6]

-
Pilkington, L. I.; Barker, D. J. Org. Chem. 2012, 77, 8156–8166.
[https://doi.org/10.1021/jo3015006]

-
Pilkington, L. I.; Wagoner, J.; Polyak, S. J.; Barker, D. Org. Lett. 2015, 17, 1046–1049.
[https://doi.org/10.1021/acs.orglett.5b00189]

-
Yang, F.; Zhu, W.; Sun, S.; Ai, Q.; Edirisuriya, P.; Zhou, K. J. Nat. Prod. 2020, 83, 825–833.
[https://doi.org/10.1021/acs.jnatprod.9b00279]

-
Lu, Y.; Chen, D.-F. J. Chromatogr. A 2009, 1216, 1980–1990.
[https://doi.org/10.1016/j.chroma.2008.09.070]

-
Wagner, H.; Bauer, R.; Melchart, D.; Xiao, P.-G.; Staudinger, A. Chromatographic fingerprint analysis of herbal medicines; Springer: United States of America, 2011; pp 37–44.
[https://doi.org/10.1007/978-3-7091-0763-8_4]

-
Kamal‐Eldin, A.; Appelqvist, L. Å.; Yousif, G. J. Am. Oil Chem. Soc. 1994, 71, 141–147.
[https://doi.org/10.1007/BF02541548]