Antibacterial Activity of Stilbenes Isolated From Reynoutria japonica Against Vancomycin-resistant Enterococci: An In Vitro and In Silico Approach
Abstract
Enterococcus species are Gram-positive cocci normally found in the gut but have emerged as opportunistic pathogens in healthcare settings, with rising multidrug resistance, particularly vancomycin-resistant Enterococci (VRE) posing major therapeutic challenges. This study evaluated the antibacterial and synergistic effects of plant-derived stilbenes from Reynoutria japonica against clinical VRE isolates. Ten clinical Enterococcus faecalis isolates were identified using standard morphological, biochemical, and Gram-staining methods, and antibiotic susceptibility testing. The antibacterial activity of polydatin, resveratroloside, and trans-resveratrol was assessed by agar well diffusion, and evaluation of minimum inhibitory concentrations (MICs) and evaluation of bactericidal activity concentrations (MBCs), with synergistic effects tested in combination with vancomycin. Molecular docking was performed with DapF, vanA, and gelE proteins. All isolates were resistant to vancomycin, ampicillin, imipenem, gentamicin, and ciprofloxacin but sensitive to linezolid. All three stilbenes displayed antibacterial activity. Notably, trans-resveratrol was the most active compound, with a MIC value of 0.156 mM and bactericidal activity (an MBC value equal to the MIC). Furthermore, all compounds exhibited synergistic effects when combined with vancomycin. Phytochemical analysis confirmed these stilbenes as major constituents in the root extract of R. japonica. Molecular docking studies revealed strong binding affinities of the compounds to key virulence and resistance-associated proteins (DapF, VanA, and GelE). These results indicate that trans-resveratrol and related stilbenes are promising candidates for development as alternative or adjunct therapeutic agents against VRE infections. Additionally, this study substantiates the traditional use of R. japonica as a source of bioactive antimicrobial compounds.
Keywords:
Enterococci, Reynoutria japonica, Stilbenes, HPLC, Antibacterial activity, Molecular dockingIntroduction
Enterococcus species are Gram-positive, facultative anaerobic cocci that are commensal inhabitants of the human gastrointestinal tract but can also colonize sites such as the respiratory and biliary tracts.1 Their ability to survive harsh conditions stems from metabolic flexibility and tolerance to variations in pH, salts, detergents, and antimicrobial agents, enhancing their competitiveness in diverse environments.2 Clinically, they are important opportunistic pathogens causing infections like endocarditis, surgical wound infections, bacteremia, and urinary tract infections, and are associated with high hospital prevalence and significant mortality rates.3
Their pathogenicity is linked to several virulence factors, including the endocarditis antigen (efaA), gelatinase (gelE), aggregation substance (agg), cytolysin, and enterococcal surface proteins (esp), which promote immune evasion, tissue invasion, biofilm formation, and inflammation.4 Since the 1980s, multidrug-resistant Enterococci have become major causes of hospital-acquired infections due to intrinsic resistance and the acquisition of mobile genetic elements conveying resistance to aminoglycosides, β-lactams, glycopeptides, and other classes.5 Vancomycin-resistant Enterococcus (VRE) has become widespread globally, limiting treatment options and creating significant therapeutic challenges.6
The medicinal plant Reynoutria japonica Houtt. (Polygonum cuspidatum Siebold & Zucc.) is commonly known as Japanese knotweed, is a perennial herb belonging to the Polygonaceae family. Native to East Asia, this species can grow 1–2 meters tall and is now widely distributed across South Korea, Japan, China, North America, and Europe.7,8 R. japonica is a rich source of bioactive metabolites. The young shoots of R. japonica are commonly consumed as food, whereas the roots have been historically used as a natural dye and as a medicinal ingredient. In traditional practice, the root is prepared as granules, powders, or decoctions to treat a wide range of conditions, including inflammation, infections, jaundice, skin burns, chronic bronchitis, and hyperlipidemia.9,10 Phytochemical investigations have identified a diverse array of secondary metabolites in this plant, notably quinones, stilbenes, flavonoids, and coumarins.10,11 Among them, stilbenes exhibit diverse biological activities especially resveratrol along with its derivatives, are associated with a broad spectrum of pharmacological activities, including anti-inflammatory, anticancer, antiviral, and antibacterial activities.12,13 This study evaluated the efficacy of stilbene compounds derived from Reynoutria japonica against clinical isolates of vancomycin-resistant Enterococci (VRE). The investigation assessed both their intrinsic antibacterial activity and their potential synergistic effects when combined with vancomycin. Furthermore, to elucidate potential mechanisms of action, in silico molecular docking analyses were conducted to examine the interactions of these stilbenes with key bacterial targets: the D-alanyl-D-alanine dipeptidase (DapF), the VanA ligase, and the gelatinase (GelE) enzyme.
Experimental
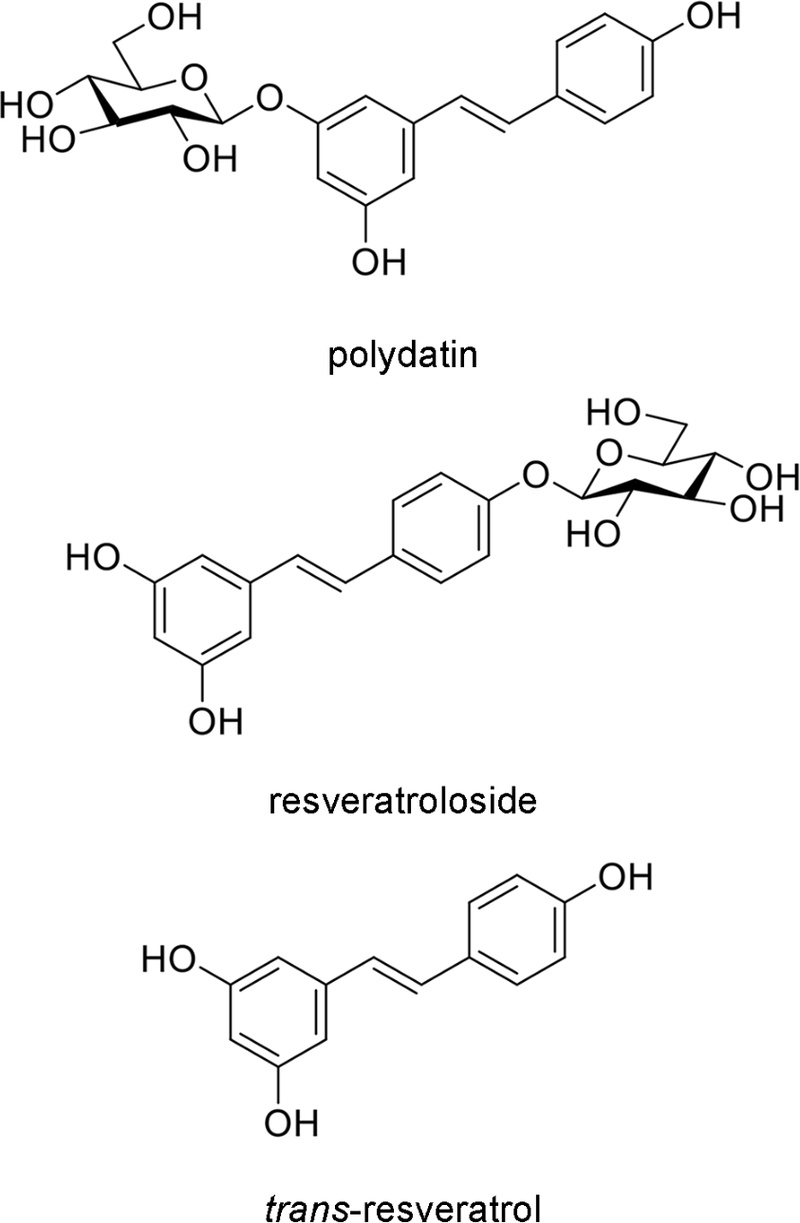
Isolation of stilbenes from R. japonica – The three stilbene derivatives polydatin, resveratroloside, and trans-resveratrol were isolated from an ethanolic extract of Reynoutria japonica Houtt. root. The detailed isolation procedures and comprehensive spectroscopic characterization data for these compounds have been previously reported.14,15
Enterococcal isolates – A total of ten enterococcal isolates were utilized in this study which were previously collected from the microbiology laboratories of three tertiary care hospitals in Peshawar, Pakistan (Rehman Medical Institute, Northwest Hospital, and Khyber Teaching Hospital), from September 2020 to May 2021.16 Following collection, samples were streaked onto Bile Esculin agar plates and incubated aerobically at 37°C for 18–24 hours. After incubation, colonies exhibiting black pigmentation and a pointed morphology were selected for further analysis. Putative enterococcal isolates were confirmed based on Gram-positive cocci morphology, positive pyrrolidonyl arylamidase (PYR) test results, growth in broth containing 6.5% NaCl, and the ability to hydrolyze esculin in the presence of bile.16
Antibiotic susceptibility testing – Antibiotic susceptibility testing was performed using the Kirby-Bauer disk diffusion method on Mueller-Hinton Agar (MHA) to identify multidrug-resistant isolates of Enterococcus spp. The assay was conducted in accordance with the guidelines published by the Clinical and Laboratory Standards Institute (CLSI, 2020).16 The following antibiotic disks were used: vancomycin (30 µg), ampicillin (30 µg), imipenem (30 µg), gentamicin (10 µg), linezolid (30 µg), and ciprofloxacin (5 µg).
Evaluation of minimum inhibitory concentrations (MICs) – The minimum inhibitory concentrations (MICs) of vancomycin and the stilbene compounds were determined using a broth microdilution method as described in a previous study with minor modifications.17 Bacterial suspension was prepared in Luria Bertani (LB) broth, and its turbidity was standardized to a 0.5 McFarland’s standard, equivalent to approximately 1–2 × 10⁸ CFU/mL. 100 µL of each serially diluted compound and 100 µL of the bacterial inoculum were added to the wells of a 96-well microtiter plate. Vancomycin, a glycopeptide antibiotic with known activity against enterococci, was included as a positive control.
Evaluation of minimum bactericidal concentrations (MBCs) – The minimum bactericidal concentration (MBC) was determined according to the protocols reported in a study.18 Bacteria were subculture from wells showing no visible growth in the MIC assay. Blood agar plates were prepared and allowed to solidify. The surface of each plate was divided into labelled sections to accommodate multiple samples. Using a sterile loop, bacterial samples were collected from the appropriate wells of the microtiter plate and streaked onto their designated sections on the blood agar. Both positive (bacterial growth control) and negative (sterility) controls were included. The inoculated plates were incubated at 37°C for 20–24 hours. Following incubation, the plates were examined for the presence or absence of bacterial growth to determine the MBC, defined as the lowest concentration that resulted in ≥99.9% killing of the initial inoculum.
Synergistic activities of stilbenes with vancomycin – The synergistic effects of polydatin, resveratroloside, and trans-resveratrol with vancomycin were evaluated against vancomycin-resistant Enterococci (VRE) isolates exhibiting high minimum inhibitory concentration (MIC) values. Checkerboard broth microdilution assay was employed to assess the interactions.19 A total of 200 µL of highest concentration of vancomycin solution was added to each well of the first row of microtiter plate, except for the first well. Subsequently, 100 µL of sterile MHB was added to the remaining wells of the plate. Vancomycin was diluted by double dilution method from the first row to the last row. Similarly, 200 µL of polydatin solution was added to each well of the 12th column followed by two-fold serial dilution from the 12th column to the 2nd column. After preparing the dilution mixture, 100 µL of bacterial suspension which was prepared in normal saline and adjusted to a 0.5 McFarland standard, was added to the wells containing different concentrations of vancomycin-polydatin combinations. The plate was incubated at 37°C for 24 hours. The same procedure was performed for resveratroloside and trans-resveratrol.
HPLC analysis – Chromatographic analysis was performed using an Agilent 1260 series HPLC system (Agilent Technologies, Palo Alto, CA, USA), equipped with a manual sampler, a quaternary pump, a degasser, and a column oven. Separation was achieved on a Capcell Pak C18 reversed-phase column (4.6 × 250 mm; Shiseido Co. Ltd., Kyoto, Japan) maintained at 30°C. Samples (10 μL) were injected onto the column and eluted using a gradient of acetonitrile and water at a flow rate of 0.8 mL/min. The gradient program was as follows: 10–30% acetonitrile (0–5 min), 30–60% (5–15 min), 60–65% (15–20 min), and 65–70% (20–30 min), followed by a 5-min conditioning phase to re-equilibrate the column to initial conditions. Elution was monitored at a wavelength of 254 nm. System control and data acquisition were managed using ChemStation software (Agilent Technologies).
Retrieval of protein structures – The three-dimensional structures of the target proteins, DapF (UniProt ID: Q838I3), VanA (UniProt ID: G1FK41), and GelE (UniProt ID: Q833V7) were retrieved from the UniProt via their corresponding UniProt accession numbers.20 The selected crystal structures were prepared for docking by removing water molecules, heteroatoms, and adding polar hydrogen atoms using BIOVIA Discovery Studio Visualizer.
Energy minimization – To alleviate steric clashes and optimize the system to a local energy minimum, all protein structures underwent energy minimization. The procedure was implemented using the AMBER force field within a Google Colab environment, utilizing a custom Python notebook (structure_relax_amber.ipynb), with parameters set to a maximum of 2000 iterations, an energy tolerance of 0.0001 kcal/mol, and a restraint weight of 10. Subsequently, the stereochemical quality of the energy-minimized structures was assessed by analyzing Ramachandran plots produced by the PdbSUM server.21
Active site prediction – The putative active sites of all target proteins were predicted using the AlphaFold2Bind algorithm, implemented via a custom Python script within a Google Colab environment (Binding_Site_Detection_with_AlphaFold2_Part_10.ipynb). Subsequently, the identified binding site residues were visualized and highlighted on the protein structures using PyMOL (Open version).22
Ligand retrieval and preparation – Three-dimensional structures of the ligands, polydatin (PubChem CID: 5281718) and resveratrol (PubChem CID: 445154), were retrieved in SDF format from the PubChem database.23 Both compounds were then visualized and converted into the Protein Data Bank (PDB) file format using BIOVIA Discovery Studio Visualizer for subsequent molecular docking analyses.
Preparation of ligands and target proteins for molecular docking – Molecular docking was performed using AutoDock tools within the MGLTools software package. All target protein structures and ligands were prepared for docking by conversion to the PDBQT format, which incorporates atomic partial charges and atom types. Protein preparation involved the removal of all water molecules and non-standard residues, followed by the addition of polar hydrogen atoms. Gasteiger partial charges were assigned to both the protein and ligand structures. The docking simulations were conducted using a grid-based approach, with the search space defined around the predicted active site of each target protein. The dimensions and center coordinates of the grid box for each protein were as follows:
DapF: size (x, y, z) = 50, 50, 50 Å; center (x, y, z) = −0.355, −0.067, −0.178.
VanA: size (x, y, z) = 50, 50, 50 Å; center (x, y, z) = −1.165, 2.111, 1.692.
GelE: size (x, y, z) = 54, 54, 54 Å; center (x, y, z) = 2.469, −5.840, 9.621.
Molecular docking – Molecular docking was performed to predict the binding affinities and poses of the selected compounds (polydatin and resveratrol) against the target proteins using AutoDock Vina.24 The docking simulations were executed using predefined configuration files that specified the search parameters for each protein. Following docking, the resulting ligand-protein complexes were visualized, and the poses with the most favorable binding affinities were extracted and converted to the Protein Data Bank (PDB) format using BIOVIA Discovery Studio Visualizer for subsequent analysis.
Results and Discussion
Antibiotics are considered as one of the most remarkable discoveries that saved millions of people from bacterial infections but due to its excessive use and misuse, bacteria developed antibiotic resistance which is a significant and emerging global health problem contributing to 1.2 million deaths worldwide in 2019.25 The escalating global crisis of antibiotic resistance necessitates the exploration of novel therapeutic strategies, including the use of phytochemicals to enhance the efficacy of existing antibiotics.26, 27 Plant-derived secondary metabolites, known for their antimicrobial defence roles, offer promise as synergists to potentiate standard treatments.28,29 In this context, we investigated the antibacterial and synergistic potential of stilbene compounds from R. japonica against vancomycin-resistant Enterococci (VRE), integrating in vitro assays with in silico molecular docking to elucidate their mechanisms of action.
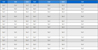
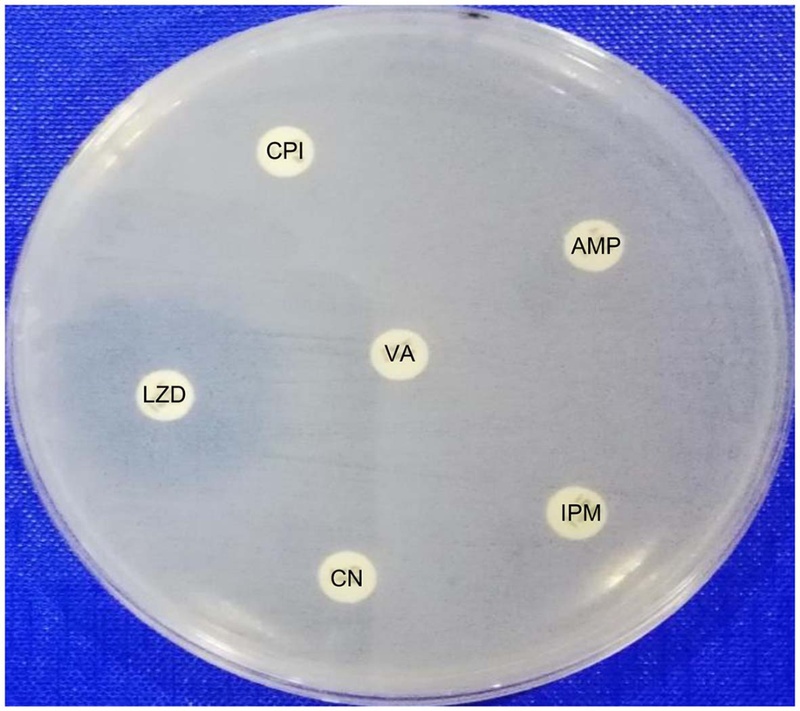
All ten clinical VRE isolates used in this study exhibited a multidrug-resistant phenotype, demonstrating susceptibility only to linezolid and resistance to vancomycin, ampicillin, imipenem, gentamicin, and ciprofloxacin (Table 1, and Fig. 1). This resistance profile is consistent with global reports, such as studies by Sienko et al. and Coombs et al., which documented high-level resistance to imipenem/ampicillin and ampicillin, respectively, underscoring the therapeutic challenge posed by VRE.30,31
The intrinsic antibacterial activity of the stilbenes polydatin, resveratroloside and trans-resveratrol were subsequently evaluated. The minimum inhibitory concentration (MIC) determinations revealed that trans-resveratrol was the most potent compound, with an MIC of 0.156 ± 0.002 mM, while polydatin and resveratroloside showed MICs of 2.5 ± 0.003 mM and 5 ± 0.003 mM, respectively (Table 2). The potent activity of trans-resveratrol aligns with previous findings by Ma et al., who demonstrated its efficacy against a range of pathogens.32 Furthermore, the minimum bactericidal concentration (MBC) was found to be equal to the MIC for all three compounds, confirming their bactericidal, rather than bacteriostatic, mode of action against the tested VRE strains.
Given the potential of phytochemicals to augment conventional antibiotics, we assessed the combination of each stilbene with vancomycin.29 The co-administration significantly enhanced vancomycin’s activity against the VRE strains (Table 3), indicating a clear synergistic interaction. This finding is particularly significant as it suggests a strategy for potentially overcoming vancomycin resistance, allowing the drug to regain activity against resistant strains.
To contextualize the relevance of our tested compounds, we analyzed the crude ethanolic extract of R. japonica roots using RP-HPLC. The chromatogram, monitored at 254 nm, displayed three major peaks corresponding to the retention times of the authentic standards for polydatin, resveratroloside, and trans-resveratrol (Fig. 2, 3A and 3B). This confirms the presence of these bioactive stilbenes as major constituents in the native plant material, validating the use of the isolated compounds for our assays and supporting the findings of Shan et al. and Song et al. on the antibacterial activity of R. japonica extracts.33,34
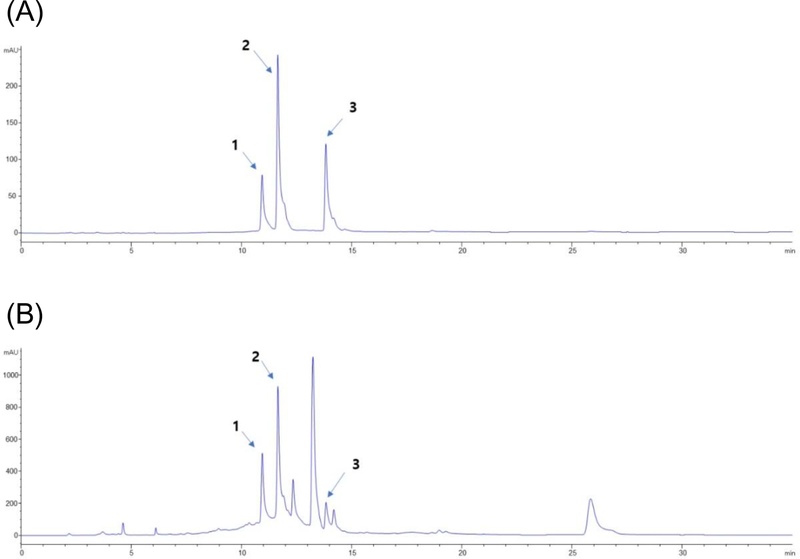
HPLC chromatogram of R. japonica roots crude extract (254 nm). (A) HPLC chromatogram of standards (B) HPLC chromatogram of ethanolic crude extract. 1. resveratroloside, polydatin and trans-resveratrol.
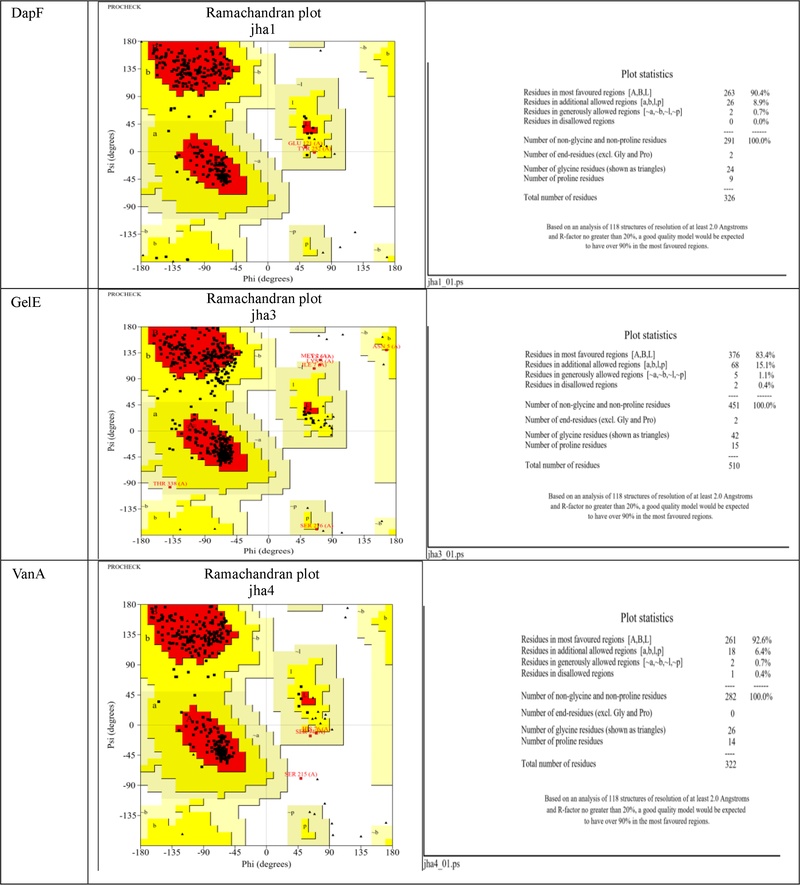
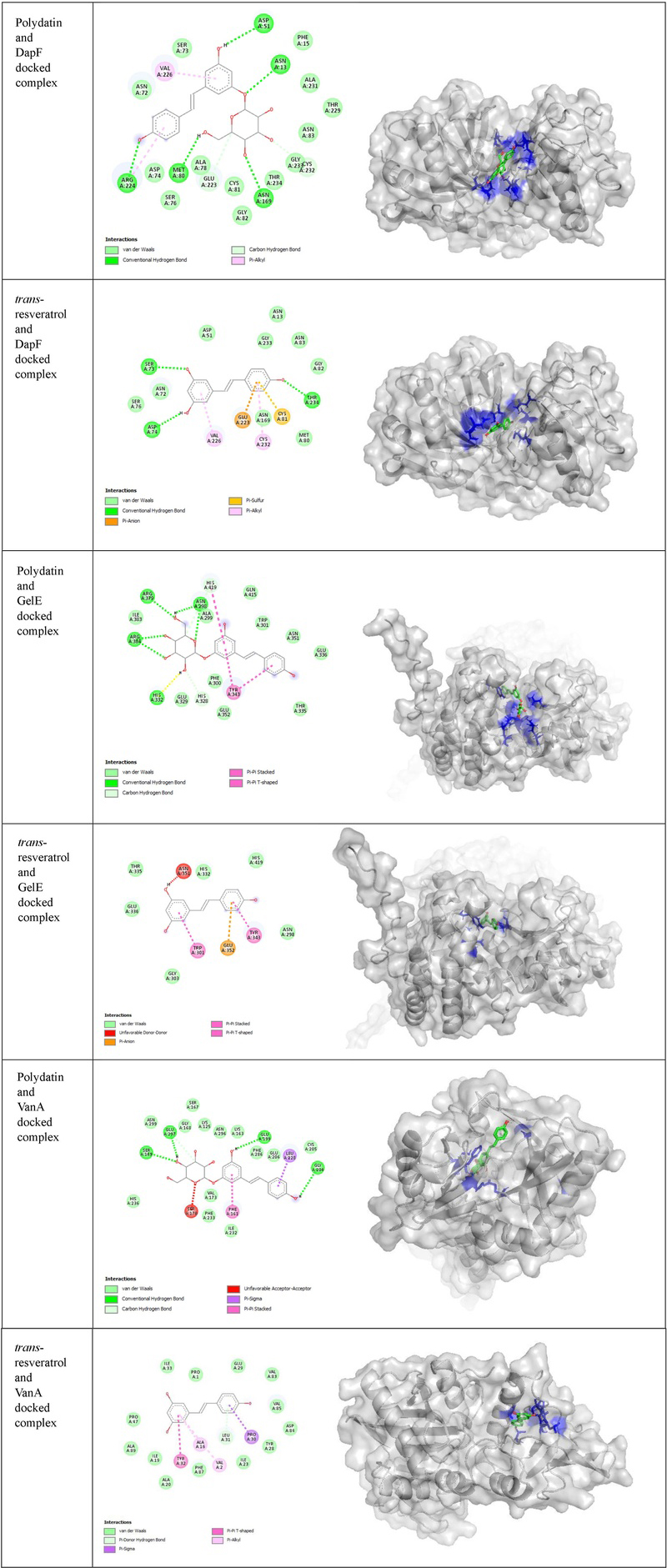
To elucidate the potential molecular mechanisms underlying the observed antibacterial effects, we performed molecular docking simulations with three key proteins in Enterococcus faecalis: D-alanine-D-alanine dipeptidase (DapF, involved in peptidoglycan biosynthesis), VanA ligase (central to vancomycin resistance), and gelatinase (GelE, a virulence factor). The target proteins were prepared and validated, with Ramachandran plots confirming that over 90% of residues were in allowed regions for each protein (Fig. 4), ensuring the reliability of the docking simulations.
The results demonstrated strong and specific binding of both polydatin and trans-resveratrol to the active sites of these targets (Table 4, and Fig. 5). Notably, polydatin formed a highly stable complex with VanA, exhibiting a binding affinity of −9.7 kcal/mol stabilized by hydrogen bonds with key residues Leu280, Glu297, and Asn299. This high-affinity interaction suggests a potential mechanism where polydatin could directly interfere with the VanA-mediated alteration of cell wall precursors, potentially re-sensitizing VRE to vancomycin, a finding that provides a molecular rationale for the observed synergy.
Similarly, polydatin showed strong binding to DapF (−8.4 kcal/mol) and GelE, interacting with residues critical for their catalytic activity. The superior in silico binding affinity of polydatin across all targets, compared to trans-resveratrol, can be attributed to its ability to form a greater number of stabilizing hydrogen bonds.
A compelling and critical observation emerged from comparing the computational and experimental data: despite its consistently lower predicted binding affinities (e.g., −7.2 kcal/mol with VanA), trans-resveratrol demonstrated significantly superior in vitro antibacterial potency. This discrepancy underscores a fundamental principle in drug discovery, that in silico binding predictions, while invaluable for understanding specific interactions, do not fully capture the complexity of a biological system.
We hypothesize that the potent efficacy of trans-resveratrol may be driven by factors beyond high-affinity binding to a single protein. Its smaller size and distinct physicochemical properties likely facilitate superior cellular uptake or enable membrane disruption. Furthermore, its activity may involve pleiotropic effects, such as simultaneously targeting multiple essential pathways or inducing oxidative stress, which are not captured by static docking against individual proteins. In contrast, polydatin's robust in silico performance highlights its potential as a highly specific inhibitor, particularly of VanA.
In conclusion, our integrated approach validates the antibacterial and synergistic properties of stilbenes against VRE. The docking study provides a molecular foundation for these activities, with polydatin emerging as a strong specific binder to resistance and virulence factors, and trans-resveratrol standing out as a highly potent antibacterial agent with a likely multi-faceted mechanism. These findings highlight the importance of combining in silico and experimental methods to fully evaluate phytochemical candidates for combating multidrug-resistant bacteria.
In conclusion, the rises of VRE has been influenced by the prevalence of Enterococci in the gut flora, increased use of medical equipment, prolonged hospital stays and most importantly, inappropriate and irrational antibiotic use. It has been observed from all over the world that the prevalence of VRE is seriously rising. VRE bacteremia and UTI cause increased mortality and morbidity worldwide. Phytochemicals or plants derived compounds i-e., polydatin, resveratrol and trans-resveratrol from R. japonica can be used in conjunction with vancomycin to boost their action or synergism and enhance their antibacterial activity even at lower doses. Our finding showed that among the tested compounds, trans-resveratrol was the most potent compound against Vancomycin Resistant Enterococci (VRE). Stilbenes in combination with vancomycin enhanced its activity against VRE. This leads to the conclusion that these natural substances can be a good substitute for existing antibiotics to treat VRE.
Conflicts of Interest
The authors declare that they have no conflicts of interest.
References
- Liu, D. Laboratory Models for Foodborne Infections; CRC Press: Boca Raton, United States of America; pp 175–183.
-
García-Solache, M.; Rice, L. B. Clin. Microbiol. Rev. 2019, 32, e00058-18
[https://doi.org/10.1128/CMR.00058-18]

-
Wang, A.; Gaca, J. G.; Chu, V. H. JAMA 2018, 320, 72–83.
[https://doi.org/10.1001/jama.2018.7596]

-
Madsen, K. T.; Skov, M. N.; Gill, S.; Kemp, M. Open. Microbiol. J. 2017, 11, 1–11.
[https://doi.org/10.2174/1874285801711010001]

- Fahim, N. A. I.; Masud, R. I.; Salam, S.; Hasan, M. A. E.; Rahman, A.-M. M.; Punom, S. A.; Rahman, M. T. Ger. J. Vet. Res. 2025, 5, 95–112.
-
Miller, W. R.; Murray, B. E.; Rice, L. B.; Arias, C. A. Infect. Dis. Clin. 2016, 30, 415–439.
[https://doi.org/10.1016/j.idc.2016.02.006]

-
Patocka, J.; Navratilova, Z.; Ovando, M. Mil. Med. Sci. Lett. 2017, 86, 17–31.
[https://doi.org/10.31482/mmsl.2017.004]

-
Shan, B.; Cai, Y.-Z.; Brooks, J. D.; Corke, H. Food Chem. 2008, 109, 530–537.
[https://doi.org/10.1016/j.foodchem.2007.12.064]

-
Kirino, A.; Takasuka, Y.; Nishi, A.; Kawabe, S.; Yamashita, H.; Kimoto, M.; Ito, H.; Tsuji, H. J. Nutr. Sci. Vitaminol. 2012, 58, 278–286.
[https://doi.org/10.3177/jnsv.58.278]

-
Peng, W.; Qin, R.; Li, X.; Zhou, H. J. Ethnopharmacol. 2013, 148, 729–745.
[https://doi.org/10.1016/j.jep.2013.05.007]

-
Khalil, A. A. K.; Park, W. S.; Lee, J.; Kim, H.-J.; Akter, K.-M.; Goo, Y.-M.; Bae, J.-Y.; Chun, M.-S.; Kim, J.-H.; Ahn, M.-J. Arch. Pharm. Res. 2019, 42, 505–511.
[https://doi.org/10.1007/s12272-019-01160-x]

-
Akinwumi, B. C.; Bordun, K.-A. M.; Anderson, H. D. Int. J. Mol. Sci. 2018, 19, 792.
[https://doi.org/10.3390/ijms19030792]

-
De Filippis, B.; Ammazzalorso, A.; Amoroso, R.; Giampietro, L. Drug Dev. Res. 2019, 80, 285–293.
[https://doi.org/10.1002/ddr.21525]

-
Khalil, A. A. K.; Akter, K.-M.; Kim, H.-J.; Park, W. S.; Kang, D.-M.; Koo, K. A.; Ahn, M.-J. Plants 2020, 9, 222.
[https://doi.org/10.3390/plants9020222]

-
Khalil, A. A. K.; Park, W. S.; Kim, H.-J.; Akter, K. M.; Ahn, M.-J. Nat. Prod. Sci. 2016, 22, 220–224.
[https://doi.org/10.20307/nps.2016.22.3.220]

-
Niaz, F.; Faheem, M.; Khattak, M.; Khawaja, I. A.; Ahn, M.-J.; Sarker, U.; Jamal, S. B.; Ullah, R.; Khalil, A. A. K. Biomed. Res. Int. 2022, 2022, 6197375.
[https://doi.org/10.1155/2022/6197375]

-
Kadeřábková, N.; Mahmood, A. J. S.; Mavridou, D. A. I. NPJ Antimicrob. Resist. 2024, 2, 37.
[https://doi.org/10.1038/s44259-024-00051-6]

- AL-Saadi, Z. N. Am. J. Biomed. Life Sci. 2016, 4, 75–80.
-
Bellio, P.; Fagnani, L.; Nazzicone, L.; Celenza, G. MethodsX 2021, 8, 101543.
[https://doi.org/10.1016/j.mex.2021.101543]

- Bateman, A.; Martin, M.-J.; Orchard, S.; Magrane, M.; Adesina, A.; Ahmad, S.; Bowler-Barnett, E. H.; Bye-A-Jee, H.; Carpentier, D.; Denny, P. Nucleic Acids Res. 2025, 53, D609–D617.
-
Laskowski, R. A. Nucleic Acids Res. 2009, 37, D355–D359.
[https://doi.org/10.1093/nar/gkn860]

- Gazizov, A.; Lian, A.; Goverde, C.; Ovchinnikov, S.; Polizzi, N. F. bioRxiv 2023, 2023-10.
-
Kim, S.; Chen, J.; Cheng, T.; Gindulyte, A.; He, J.; He, S.; Li, Q.; Shoemaker, B. A.; Thiessen, P. A.; Yu, B.; Zaslavsky, L.; Zhang, J.; Bolton, E. E. Nucleic Acids Res. 2023, 51, D1373–D1380.
[https://doi.org/10.1093/nar/gkac956]

-
Trott, O.; Olson, A. J. J. Comput. Chem. 2010, 31, 455–461.
[https://doi.org/10.1002/jcc.21334]

- Murray, C. J.; Ikuta, K. S.; Sharara, F.; Swetschinski, L.; Aguilar, G. R.; Gray, A.; Han, C.; Bisignano, C.; Rao, P.; Wool, E. The Lancet 2022, 399, 629–655.
-
Murray, B. E. J. Infect. Dis. 1991, 163, 1185–1194.
[https://doi.org/10.1093/infdis/163.6.1185]

-
Murugaiyan, J.; Kumar, P. A.; Rao, G. S.; Iskandar, K.; Hawser, S.; Hays, J. P.; Mohsen, Y.; Adukkadukkam, S.; Awuah, W. A.; Ruiz Alvarez, M. J.; Nanono, S.; Nansubuga, E. P.; Tilocca, B.; Roncada, P.; Roson-Calero, N.; Moreno-Morales, J.; Amin, R.; Kumar, B. K.; Kumar, A.; Toufik, A.-R.; Zaw, T. N.; Akinwotu, O. O.; Satyaseela, M. P.; van Dongen, M. B. M. Antibiotics 2022, 11, 200.
[https://doi.org/10.3390/antibiotics11020200]

-
Wallace, R. J. Proc. Nutr. Soc. 2004, 63, 621–629.
[https://doi.org/10.1079/PNS2004393]

-
Cheesman, M. J.; Ilanko, A.; Blonk, B.; Cock, I. E. Pharmacogn. Rev. 2017, 11, 57–72.
[https://doi.org/10.4103/phrev.phrev_21_17]

-
Sieńko, A.; Czaban, S.; Ojdana, D.; Majewski, P.; Wieczorek, A.; Sacha, P.; Tryniszewska, E. A.; Wieczorek, P. J. Med. Sci. 2018, 87, 195–203.
[https://doi.org/10.20883/jms.288]

-
Coombs, G. W.; Pearson, J. C.; Daley, D. A.; Le, T.; Robinson, O. J.; Gottlieb, T.; Howden, B. P.; Johnson, P. D.; Bennett, C. M.; Stinear, T. P. J. Clin. Microbiol. 2014, 52, 897–905.
[https://doi.org/10.1128/JCM.03286-13]

-
Ma, D. S.; Tan, L. T.-H.; Chan, K.-G.; Yap, W. H.; Pusparajah, P.; Chuah, L.-H.; Ming, L. C.; Khan, T. M.; Lee, L.-H.; Goh, B.-H. Front. Pharmacol. 2018, 9, 102.
[https://doi.org/10.3389/fphar.2018.00102]

-
Shan, B.; Cai, Y.-Z.; Brooks, J. D.; Corke, H. J. Agric. Food. Chem. 2007, 55, 5484–5490.
[https://doi.org/10.1021/jf070424d]

-
Song, J.-H.; Kim, S.-K.; Chang, K.-W.; Han, S.-K.; Yi, H.-K.; Jeon, J.-G. Arch. Oral. Biol. 2006, 51, 1131–1140.
[https://doi.org/10.1016/j.archoralbio.2006.06.011]