PTP1B Inhibitory Constituents from Dystaenia takeshimana: Mechanistic Insights via Kinetic and Molecular Docking Analyses
Abstract
Dystaenia takeshimana (Nakai) Kitagawa belongs to the family Apiaceae (Umbelliferae) and is a perennial herb naturally endemic to Ulleung Island, Republic of Korea. Following its successful introduction and cultivation in mainland Korea, a phytochemical investigation of this plant was carried out. As a result, thirteen compounds were isolated and structurally characterized, including one flavonoid glycoside (1), two polyacetylenes (2 and 3), one caffeoylquinic acid derivative (4), one aliphatic hydroxy acid (5), four phenolic acids and their derivatives (6, 8–10), one aromatic alcohol derivative (7), two carbohydrates (11 and 12), and one phytosterol (13). The structures of these compounds were elucidated based on spectroscopic analyses and comparison with previously reported literature data. To the best of our knowledge, this study represents the first report on the isolation of falcarinol (2), trans-4-hydroxy-2-nonenoic acid (5), and sucrose (11) from D. takeshimana and the evaluation of their protein tyrosine phosphatase 1B (PTP1B) inhibitory activity. Compounds 2, 3, and 5 exhibited potent PTP1B inhibitory effects with IC50 values of 10.74, 8.31, and 25.24 µM, respectively. Enzyme kinetic studies indicated noncompetitive inhibition by 2 and mixed-type inhibition by 5 against PTP1B. Furthermore, molecular docking analysis was performed to investigate the binding interactions between the active compounds and the enzyme active site.
Keywords:
Dystaenia takeshimana, Apiaceae, PTP1B, Polyacetylene, Molecular dockingIntroduction
Dystaenia takeshimana (Nakai) Kitagawa, a member of the family Apiaceae (Umbelliferae), is a perennial herb reaching 1.5–2 m in height and endemic to Ulleung Island, Korea.1,2 Its leaves and roots have traditionally been used as food, forage, and oriental medicine.3 The genus Dystaenia comprises two species: D. ibukiensis, distributed in Japan, and D. takeshimana. Molecular phylogenetic analyses have demonstrated that the genus forms a monophyletic lineage and suggested that D. takeshimana likely evolved from D. ibukiensis.2 Previous pharmacological studies have reported several biological activities of D. takeshimana. Its extracts have shown antioxidant and α-glucosidase inhibitory activities.4 In addition, the anti-inflammatory activity of its ethanol extract has been demonstrated through the suppression of inducible nitric oxide synthase (iNOS) and cyclooxygenase-2 (COX-2) expression in lipopolysaccharide-stimulated RAW 264.7 macrophages.5 Antimicrobial activity has also been reported; the ethanol extract and n-hexane fraction of this plant inhibited several Listeria monocytogenes strains, leading to the isolation of falcarindiol as an active antimicrobial compound.6 More recently, D. takeshimana extracts have also been reported to exhibit anti-thrombotic activity, including the inhibition of platelet aggregation.3 Phytochemical investigation of the roots has revealed the presence of coumarins, flavonoids, phenethyl alcohol derivatives, sterols, polyacetylenes, and D-mannitol.1,6 Coumarins and flavonoids isolated from this plant have been reported to exhibit dual inhibitory activity against COX-2 and 5-lipoxygenase (5-LOX), while falcarindiol has been identified as an antimicrobial constituent.1 However, phytochemical investigations of D. takeshimana remain limited. In the present study, we investigated the chemical constituents of D. takeshimana cultivated and harvested in mainland Korea. As a result, thirteen compounds were isolated, and their protein tyrosine phosphatase 1B (PTP1B) inhibitory activities were evaluated using both in vitro and in silico approaches.
Experimental
General experimental procedures − An Autobio PHOmo reader was used for spectrophotometric measurement in the PTP1B inhibitory assay. High-resolution electrospray ionization mass spectrometry (HRESIMS) was measured using a Micromass QTOF2-MS spectrometer. Nuclear magnetic resonance (NMR) spectra were acquired on a Varian Unity Inova 400 MHz (Varian, Inc., California, USA) and Bruker Avance 500 MHz (Bruker, Karlsruhe, Germany) spectrometers using tetramethylsilane (TMS) as an internal standard, with chemical shifts reported in δ (ppm). Open column chromatography was implemented with Sephadex LH-20 (Pharmacia Company), silica gel 60 (Merck, 230–400 mesh), and reversed-phase (RP)-C18 silica gel (Merck, 75 mesh). Thin-layer chromatography (TLC) was carried out using Merck precoated silica gel F254 plates and RP-18 F254s plates, and compounds were visualized by spraying with aqueous 10% H2SO4 and heating for 3–5 min.
Chemicals and Reagents − The organic solvents were purchased from Duksan Pharmaceutical Co. (Seoul, Korea). Protein tyrosine phosphatase 1B (PTP1B, human recombinant) and dithiothreitol (DTT) were purchased from NKMAX Co. Ltd. (Seongnam, Korea) and Bio-Rad Laboratories (Hercules, CA, USA), respectively. Reagents utilized in the enzyme inhibitory assay were procured from Sigma-Aldrich Co. (St. Louis, MO, U.S.A.), including p-nitrophenyl phosphate (p-NPP), ethylene diamine tetraacetic acid (EDTA), sodium citrate, sodium hydroxide, ursolic acid, and dimethyl sulfoxide (DMSO).
Plant material − The roots of Dystaenia takesimana were collected in June 2022 from the medicinal plant garden at the College of Pharmacy, Daegu Catholic University, Republic of Korea (3554′43.3″ N, 12848′16.4″ E). The plant material was authenticated by Professor Byung Sun Min, one of the authors. A voucher specimen (CUD-2343) has been deposited in the Laboratory of Pharmacognosy, College of Pharmacy, Daegu Catholic University, Republic of Korea.
Extraction and isolation − The dried roots of D. takesimana (9.8 kg) were extracted with MeOH (15 L × 4) under reflux and filtered. During concentration of the combined filtrates under reduced pressure, a poorly soluble white precipitate formed. The precipitate was collected by filtration and washed with MeOH to yield a mixture (1.0 kg) of compounds 11 and 12. The remaining filtrate was further evaporated under reduced pressure to afford a crude MeOH extract (1.2 kg). The MeOH extract was suspended in distilled H2O and successively partitioned with methylene chloride (CH2Cl2) and ethyl acetate (EtOAc) to yield the CH2Cl2 fraction (668 g), EtOAc fraction (7 g), and the remaining aqueous layer. The CH2Cl2 (668 g) was chromatographed by silica gel CC eluted with CH2Cl2:MeOH solvent system (stepwise, 100:1 → 100% MeOH, v/v) to afford 15 distinct fractions DTC1-DTC15. Fraction DTC5 (10 g) was divided into 13 sub-fractions DTC5.1–DTC5.13 using silica gel CC eluted with n-hexane: acetone (10:1, v/v). Sub-fraction DTC5.4 (1.8 g) was separated by silica gel CC (n-hexane: EtOAc, 10:1, v/v) to obtain compounds 2 (10.0 mg) and 13 (12 mg). Compound 3 (8 mg) was obtained from fraction DTC5.12 (121 mg) by silica gel CC (n-hexane: EtOAc, 5:1, v/v). The EtOAc fraction (7 g) was chromatographed by silica gel CC eluted with CH2Cl2:MeOH solvent system (30:1 → 100% MeOH, stepwise) to afford 10 fractions (DTE1-DTE10). Fraction DTE3 (250 mg) was separated by RP-18 CC (MeOH-H2O, 1:1, v/v) to yield compounds 9 (6 mg), 8 (10 mg), and 5 (3 mg). Compound 4 (5 mg) was obtained from fraction DTE6 (110 mg) by RP-18 CC (MeOH-H2O, 1.5:1, v/v). Fraction DTE4 (245 mg) was purified on a Sephadex LH-20 column (MeOH-H2O, 1:1, v/v) to yield compounds 6 (16 mg) and 7 (6 mg). Fraction DTE7 (245 mg) was separated on a Sephadex LH-20 column (MeOH-H2O, 1.5:1, v/v) to yield six sub-fractions DTE7.1–DTE7.6. Fraction DTE7.5 (420 mg) was subjected to a RP-18 column (MeOH-H2O, 2:1, v/v) to give 10 (10 mg) and 1 (5 mg).
Apigenin-7-O-glucoside (1) – Yellow amorphous powder. 1H-NMR (500 MHz, DMSO-d6): δ 12.96 (1H, s, OH-5), 7.96 (2H, d, J = 8.5 Hz, H-2′, H-6′), 6.94 (2H, d, J = 8.5 Hz, H-3′, H-5′), 6.87 (1H, s, H-2), 6.83 (1H, d, J = 2.0 Hz, H-8), 6.44 (1H, d, J = 2.0 Hz, H-6), 5.07 (1H, d, J = 7.5 Hz, H-1′′), 3.70 (1H, m, H-6′′a), 3.48 (1H, m, H-6′′b), 3.44 (1H, m, H-5′′), 3.30 (1H, m, H-3′′), 3.27 (1H, m, H-2′′), 3.17 (1H, m, H-41′′); 13C-NMR (125 MHz, DMSO-d6): δ 182.0 (C-4), 164.3 (C-2), 162.9 (C-7), 161.4 (C-4'), 161.1 (C-5), 156.9 (C-8a), 128.6 (C-2', C-6'), 116.0 (C-3', 5'), 105.3 (C-4a), 103.1 (C-3), 121.0 (C-1'), 99.9 (C-11′′), 99.5 (C-6), 94.8 (C-8), 77.2 (C-51′′), 76.5 (C-31′′), 73.1 (C-21′′), 69.4 (C-41′′), 60.6 (C-61′′).
Falcarinol (2) – Yellow amorphous powder. 1H-NMR (400 MHz, CDCl3): δ 5.91 (1H, ddd, J = 16.8, 10.4, 5.6 Hz, H-2), 5.50 (1H, m, H-10), 5.47 (1H, d, J = 16.8 Hz, H-1a), 5.35 (1H, m, H-9), 5.22 (1H, d, J = 10.4 Hz, H-1b), 4.89 (1H, brd, J = 5.6 Hz, H-3), 3.02 (2H, brd, J = 6.8 Hz, H-8), 2.01 (2H, q, J = 7.2 Hz, H2-11), 1.23–1.35 (10H, m, H2-12, 13, 14, 15, 16), 0.86 (3H, t, J = 7.2 Hz, H-17); 13C-NMR (100 MHz, CDCl3): δ 136.2 (C-2), 133.1 (C-10), 122.0 (C-9), 117.0 (C-1), 80.2 (C-7), 74.4 (C-4), 71.3 (C-5), 64.2 (C-6), 63.5 (C-3), 31.9 (C-15), 27.2 (C-11), 29.2 (C-12), 29.3 (C-13), 29.3 (C-14), 22.8 (C-16), 17.7 (C-8), 14.1 (C-17); HR-ESI-MS m/z 276.1726 [M + Na]+ (calcd for C17H24ONa, 276.1725).
Falcarindiol (3) –Yellow amorphous powder. 1H-NMR (400 MHz, CDCl3): δ 5.87 (1H, ddd, J = 16.4, 10.0, 5.2 Hz, H-2), 5.53 (1H, m, H-10), 5.45 (1H, m, H-9), 5.42 (1H, d, J = 16.4 Hz, H-1a), 5.19 (1H, d, J = 10.0 Hz, H-1b), 5.15 (1H, d, J = 8.0 Hz, H-8), 4.88 (1H, m, H-3), 2.04 (2H, q, J = 6.8 Hz, H2-11), 1.19–1.345 (10H, m, H2-12, 13, 14, 15, 16), 0.83 (3H, t, J = 6.4 Hz, H-17); 13C NMR (100 MHz, CDCl3): δ 135.8 (C-2), 134.0 (C-10), 127.7 (C-9), 117.1 (C-1), 79.7 (C-7), 78.2 (C-4), 70.2 (C-5), 68.7 (C-6), 63.1 (C-3), 58.2 (C-8), 31.7 (C-15), 27.6 (C-11), 29.1 (C-12), 29.1 (C-13), 29.2 (C-14), 22.6 (C-16), 14.1 (C-17); HR-ESI-MS m/z 283.1675 [M + Na]+ (calcd for C17H24O2Na, 283.1674).
trans-4-Hydroxy-2-nonenoic acid (5) – Yellow amorphous powder. 1H-NMR (400 MHz, methanol-d4): δ 6.91 (1H, dd, J = 15.6, 5.2 Hz, H-3), 5.98 (1H, dd, J = 15.6, 1.6 Hz, H-2), 4.22 (1H, m, H-4), 1.55 (2H, m, H2-5), 1.28-1.37 (6H, m, H2-6, 7, 8), 0.91 (3H, t, J = 6.8 Hz, H-9); 13C-NMR (100 MHz, methanol-d4): δ 170.2 (C-1), 152.5 (C-3), 121.1 (C-2), 71.6 (C-4), 37.5 (C-5), 32.9 (C-7), 26.1 (C-6), 23.6 (C-8), 14.3 (C-9).
PTP1B inhibitory assay – The inhibition assay against recombinant human PTP1B enzyme was conducted using p-nitrophenyl phosphate (p-NPP) as the substrate, according to our previously published protocols.7,8 Ursolic acid was included as a positive control in this experiment as it is a well-known PTP1B inhibitor widely used as a reference compound in PTP1B inhibitory assays. The inhibitory activity of PTP1B was evaluated via an in vitro assay conducted on a 96-well plate. Samples were prepared at varying concentrations (1–100 μM) by diluting them in a reaction buffer containing 50 mM citrate buffer (pH 6.0, 0.1 M NaCl, 1 mM EDTA, and 1 mM DTT). Next, 10 μL of the test samples were added to each well, followed by 30–40 μL of reaction buffer and 10 μL of PTP1B enzyme (5 μg/mL). The mixture was incubated at 37°C for 10 minutes before adding 50 μL of 2.0 mM p-NPP substrate. Then, the reaction was mixed gently and incubated again at 37°C for 20 minutes. After 20 minutes, 10 µL of 10 M NaOH was added to stop the reaction. Under alkaline conditions, the dephosphorylation of p-NPP produced the yellow p-nitrophenolate anion, allowing spectrophotometric determination of PTP1B activity, which could be quantified at 405 nm. The percent inhibition (%) was obtained by the following equation:
where Ac is the absorbance of the control, and As is the absorbance in the presence of the test sample.
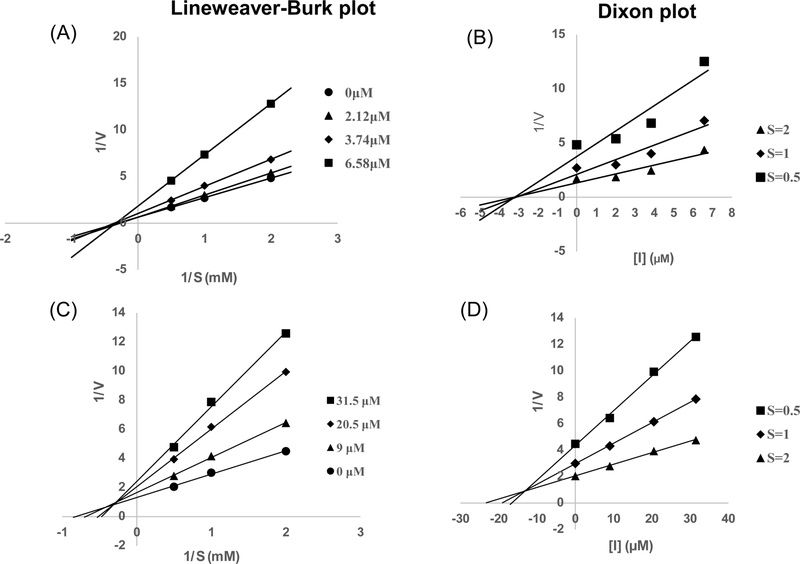
Enzyme kinetic analysis with PTP1B – The kinetic mechanisms of the active compounds (2 and 5) in PTP1B inhibition were investigated using complementary Lineweaver-Burk and Dixon plots.7,8 The PTP1B inhibition of compounds 2 and 5 was monitored at different substrate concentrations (2, 1, and 0.5 mM of p-NPP for 2 and 5) on Dixon plots (single reciprocal plots). In addition, Lineweaver-Burk plots for the inhibition of PTP1B of compounds 2 and 5 were obtained in the presence of its various concentrations (0, 2.12, 3.74, and 6.58 μM for 2 and 0, 9.0, 20.5, and 31.5 μM for 5). The kinetic results were analyzed using the SigmaPlot 12.0 software (SPCC Inc., Chicago, IL, U.S.A). The kinetic data were graphically presented by the Lineweaver-Burk and Dixon plots, from which the type of enzyme inhibition was determined. Kinetic constant values (Ki) were assessed using Dixon plots.
Molecular docking simulation in PTP1B inhibition – The interactions and binding of the active compounds (2 and 5) with PTP1B (Protein Data Bank [PDB] ID: 1T49) were investigated using Autodock Vina 1.2.7, following the previously established procedure.9 An allosteric inhibitor, 3-(3,5-dibromo-4-hydroxy-benzoyl)-2-ethyl-benzofuran-6-sulfonic acid (4-sulfamoyl-phenyl)-amide (compound A, PDB ID: 1T49) and a catalytic inhibitor, (3-((5-[(N-acetyl-3-(4-[(carboxycarbonyl)(2-carboxyphenyl)amino]-L-naphthyl)-L-alanyl)amino]pentyl)oxy)-2-naphthoic acid (compound C, PDB ID: 1NNY) were obtained from the RCSB Protein Data Bank website (https://www.rcsb.org/). The 3D structures of the tested compounds (2 and 5) were constructed using ChemDraw 3D Professional 19. AutoDock tool (Ver. 1.5.7, The Scripps Research Institute, La Jolla, CA, USA) was used for the addition of Kollman charges to the PTP1B enzyme, and then the receptor was saved to a ‘.pdbqt’ file. Ligand energy is minimized by computing Gasteiger charges using AutoDock Tools 1.5.7 before conversion to ‘.pdbqt’ files for docking simulations. The binding interactions between PTP1B residues and the corresponding binding affinity scores were considered the most relevant molecular interactions. The results were analyzed using Discovery Studio and PyMOL.
Statistical analysis − Statistical analyses were performed by one-way analysis of variance (ANOVA), followed by the Fisher least significant difference test. P < 0.05 was considered statistically significant. The results are presented as the mean±standard error of the mean (SEM).
Results and Discussion
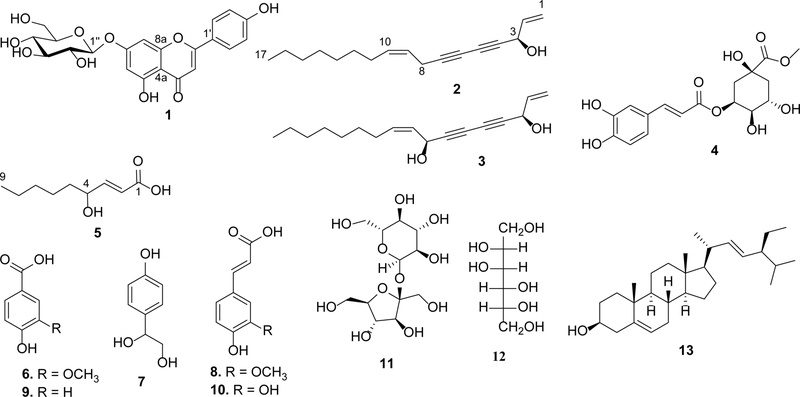
The known compounds were characterized through spectroscopic analysis and confirmed by comparison with published reference data. The chemical structures of the thirteen known compounds were determined as follows: apigenin-7-O-glucoside (1), falcarinol (2),10,11 falcarindiol (3),6,11 3-O-caffeoylquinic acid methyl ester (4),12trans-4-hydroxy-2-nonenoic acid (5),13 vanillic acid (4-hydroxy-3-methoxybenzoic acid, 6),14 2-hydroxy-2-(4'-hydroxyphenyl)ethanol (7),1 ferulic acid (4-hydroxy-3-methoxycinnamic acid, 8),15 4-hydroxybenzoic acid (9),12 caffeic acid (3,4-dihydroxycinnamic acid, 10),12 sucrose (11),16 D-mannitol (12),1 and stigmasterol (13)14 (Fig. 1).
The isolated compounds were evaluated for their PTP1B inhibitory activity, and the results are expressed as IC50 values. As shown in Table 1, compared with the positive control (ursolic acid, IC50 = 9.75 ± 0.39 μM), two polyacetylenes, falcarinol (2) and falcarindiol (3), exhibited potential PTP1B inhibitory activities, with IC50 values of 10.74 ± 0.11 μM and 8.31 ± 0.08 μM, respectively. These findings indicate the promising PTP1B inhibitory potential of polyacetylenes, which is further supported by previous reports describing the inhibitory activity of falcarindiol (3).12 A structure–activity relationship was also suggested, as the presence of an additional hydroxyl group at C-8 in falcarindiol (3) appears to enhance its PTP1B inhibitory activity compared with falcarinol (2). Meanwhile, trans-4-hydroxy-2-nonenoic acid (5) and apigenin-7-O-glucoside (1) exhibited moderate PTP1B inhibitory activities, with IC50 values of 25.24 ± 0.50 μM and 37.95 ± 0.96 μM, respectively. To the best of our knowledge, this is the first report describing the PTP1B inhibitory activities of compounds 2 and 5. In addition, cytotoxicity of the isolated compounds toward RAW264.7 and HeLa cells was evaluated using an MTT assay (MTT: 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide). None of the tested compounds showed significant cytotoxicity, even at concentrations up to 100 μM.
Compounds 2 and 5 were selected for both kinetic and molecular docking studies. Their PTP1B inhibition mechanisms were investigated using two kinetic methods, namely the Lineweaver-Burk and Dixon plots. The inhibition mechanisms at different concentrations of falcarinol (2) and trans-4-hydroxy-2-nonenoic acid (5) were evaluated by analyzing the effects of varying substrate concentrations using the Dixon plot, while the effects of different inhibitor concentrations were analyzed using the Lineweaver–Burk plot. In the Lineweaver–Burk plot, the plotted lines for falcarinol (2) intersected at the same point on the x-axis, indicating noncompetitive inhibition against PTP1B with an inhibition constant (Ki) of 3.20 µM. In contrast, the Lineweaver-Burk plot of trans-4-hydroxy-2-nonenoic acid (5), the lines of the inhibitors that intersected at the xy region indicated a mixed inhibition. The inhibition constants (Ki) were calculated from the Dixon plots, in which the intersection point on the x-axis corresponds to −Ki.
To further investigate the interaction and binding modes of the active compounds 2 (falcarinol) and 5 (trans-4-hydroxy-2-nonenoic acid) with PTP1B, molecular docking simulations were performed. These results were compared with those of known inhibitors, including compound A (an allosteric inhibitor) and compound C (a catalytic inhibitor), using AutoDock Vina 1.2.7 software. The binding energies and interacting residues, including hydrogen bonds, Van der Waals, and hydrophobic interactions, of compounds 2 and 5 with PTP1B are listed in Table 2. According to the simulation results shown in Fig. 3, both compound A (an allosteric inhibitor) and 2 (falcarinol) were stably positioned in the same pocket (allosteric site) of PTP1B with negative binding energies of −9.71 and −6.27 kcal/mol, respectively. One hydrogen bonding interaction was observed between the hydroxyl group of 2 (falcarinol) and the key allosteric residue of PTP1B, Glu276. In addition, four residues involved in Van der Waals interactions with 2 were Asn193, Ser187, Lys279, and Gly277. Falcarinol (2) and compound A shared several similar binding residues, including the allosteric residue Glu276 via hydrogen-bond interaction; Gly277 and Lys279 via Van der Waals interactions; and Phe196, Ala189, Phe280, and Leu192 via hydrophobic interactions. According to the simulation results shown in Fig. 3a, trans-4-hydroxy-2-nonenoic acid (5) demonstrated a mixed-type inhibition via binding to both the catalytic and allosteric sites of PTP1B. Similar to 2 (falcarinol), the PTP1B- trans-4-hydroxy-2-nonenoic acid (5) complex at the allosteric site showed a −5.68 kcal/mol binding energy with one hydrogen bond and the key allosteric residue of PTP1B, Glu276. Van der Waals interactions were observed between trans-4-hydroxy-2-nonenoic acid (5) and PTP1B residues Asn193, Ala189, Pro188, and Gly277, which may help this compound be further stabilized in the allosteric site of PTP1B. Catalytic inhibition by trans-4-hydroxy-2-nonenoic acid (5) against PTP1B (−5.49 kcal/mol) displayed two hydrogen-bond interactions with key PTP1B catalytic residues, Cys215 and Arg221. In addition, trans-4-hydroxy-2-nonenoic acid (5) and compound C shared several similar binding residues, including the PTP1B catalytic residues Cys215 and Arg221 via hydrogen-bond interactions; Asp48, Ser216, and Gly220 via Van der Waals interactions; and Tyr46 via hydrophobic interactions. The molecular docking results of compounds 2 and 5 were consistent with the kinetic data, providing structural insight into their inhibition mechanism.
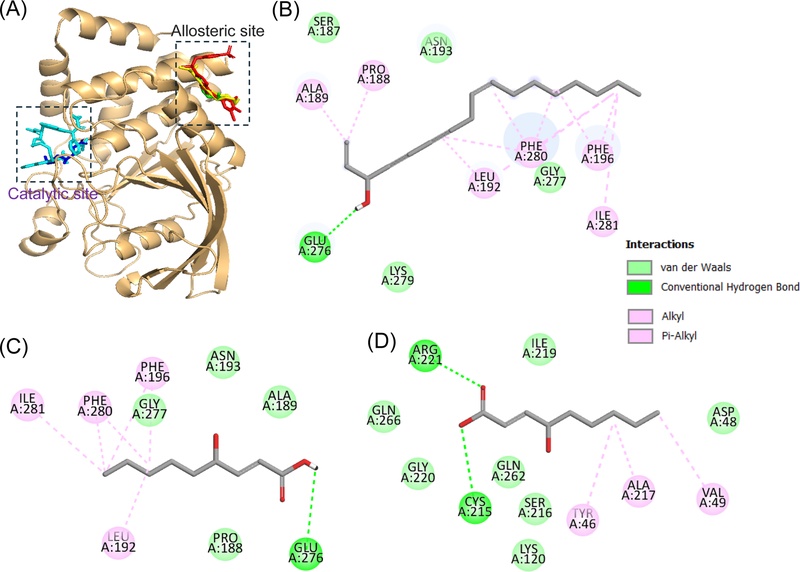
Molecular docking related to PTP1B inhibition by compound A (red stick), C (cyan stick), 2 (yellow stick), and 5 (green and blue stick) (A). 2D diagram of PTP1B inhibition by 2 (B) and 5 (C) at the PTP1B allosteric site and 5 (D) at the PTP1B catalytic site. The figure was generated using PyMOL and Discovery Studio Visualizer.
In summary, a phytochemical investigation of D. takeshimana, a plant endemic to Ulleung Island and recently introduced for cultivation in mainland Korea, resulted in the isolation of thirteen compounds. Among them, falcarinol (2), trans-4-hydroxy-2-nonenoic acid (5), and sucrose (11) were reported from this plant for the first time. Several isolates, particularly two polyacetylenes (2–3) and one aliphatic hydroxy acid (5), exhibited potential PTP1B inhibitory activity. Enzyme kinetic and molecular docking analyses further supported their inhibitory mechanisms, highlighting the potential of this plant as a natural source of PTP1B inhibitors.
Acknowledgments
This research was supported by the National Research Foundation of Korea (NRF) (NRF-2021R1A2C2011940) and Gyeongsangbuk-do RISE (Regional Innovation System & Education; 2025-RISE-15-107).
Conflicts of Interest
The authors have declared that there are no conflicts of interest.
References
-
Kim, J. S.; Kim, J. C.; Shim, S. H.; Lee, E. J.; Jin, W. Y.; Bae, K.; Son, K. H.; Kim, H. P.; Kang, S. S.; Chang, H. W. Arch. Pharm. Res. 2006, 29, 617–623.
[https://doi.org/10.1007/BF02968244]

-
Pfosser, M.; Jakubowsky, G.; Schlüter, P.; Fer, T.; Kato, H.; Stuessy, T.; Sun, B.-Y. Plant Syst. Evol. 2005, 256, 159–170.
[https://doi.org/10.1007/s00606-005-0374-9]

- Kang, D.-G.; Lee, Y.-J.; Kim, J. S.; Sohn, H.-Y. J. Life Sci. 2022, 32, 303–309.
-
Kim, J. H.; Jeong, G. H.; Jeong, Y.-H.; Kim, T. H. Food Sci. Preserv. 2019, 26, 246–252.
[https://doi.org/10.11002/kjfp.2019.26.2.246]

-
Lee, J.-Y.; Yoo, D.-H.; Joo, D.-H.; Chae, J.-W. Microbiol. Biotechnol. Lett. 2016, 44, 571–576.
[https://doi.org/10.4014/mbl.1610.10006]

- Oh, J.-A.; Shin, D.-H.; Baek, N.-I. Korean J. Food Sci. Technol. 1999, 31, 984–993.
-
Pham, T. L.; Ha, M. T.; Min, B. S.; Kim, J. A. Nat. Prod. Sci. 2025, 31, 62–73.
[https://doi.org/10.20307/nps.2025.31.1.62]

-
Ha, M. T.; Le, T. T.; Tran, T. T.; Tran, T. H.; Kim, J. A.; Min, B. S. Nat. Prod. Sci. 2025, 31, 129–137.
[https://doi.org/10.20307/nps.2025.31.2.129]

-
Dinh, T. N. A.; Le, T. T.; Ha, M. T.; Pham, T. L.; Shin, J.; Byun, J. K.; Min, B. S.; Kim, J. A. J. Agric. Food. Chem. 2025, 73, 29566–29580.
[https://doi.org/10.1021/acs.jafc.5c05819]

- Corell, M.; Sheehy, E.; Evans, P.; Brunton, N.; Valverde, J. Nat. Prod. Commun. 2013, 8, 1123–1126.
-
El-Houri, R. B.; Kotowska, D.; Christensen, K. B.; Bhattacharya, S.; Oksbjerg, N.; Wolber, G.; Kristiansen, K.; Christensen, L. P. Food Funct. 2015, 6, 2135–2144.
[https://doi.org/10.1039/C5FO00223K]

-
Ha, M. T.; Lee, T. H.; Kim, C. S.; Prajapati, R.; Kim, J. A.; Choi, J. S.; Min, B. S. Phytochemistry 2022, 197, 113100.
[https://doi.org/10.1016/j.phytochem.2022.113100]

-
Mu, L.-H.; Qiang-Feng, J.; Liu, P.; Hu, Y.; Zhao, H.-X. Chem. Nat. Compd. 2013, 48, 1004–1007.
[https://doi.org/10.1007/s10600-013-0450-2]

- Akbar, A.; Ail, M. S.; Zikr-Ur-Rehman, S.; Lateef, M.; Saeed, Z. Int. J. Biol. Biotech. 2020, 17, 9–15.
-
Kim, C.-K.; Yu, J.; Lee, M. Foods 2023, 12, 1146.
[https://doi.org/10.3390/foods12061146]

-
Pedersen, N. R.; Wimmer, R.; Emmersen, J.; Degn, P.; Pedersen, L. H. Carbohydr. Res. 2002, 337, 1179–1184.
[https://doi.org/10.1016/S0008-6215(02)00112-X]