Quantitative HPLC-DAD Analysis and α-Glucosidase Inhibitory Activity of Major Compounds from the Stems and Pericarpium of Punica granatum L.
Abstract
Punica granatum L. has been widely used in traditional medicine and is known to contain bioactive ellagitannins with potential antidiabetic properties. The primary objective of this study was to develop and validate a reliable HPLC-DAD method for the quantitative analysis of major compounds isolated from stems and pericarpium of P. granatum, namely pedunculagin (1), flavogallonic acid (2), casuarinin (3), isocorilagin (4), methyl brevifolincarboxylate (5), eschweilenol C (6), ellagic acid-4-O-β-D-xylopyranoside (7), ellagic acid (8), and quercitrin (9), and to futher evaluate their α-glucosidase inhibitory activity. The analytical method demonstrated linearity with correlation coefficients (R²) exceeding 0.9991. The limits of detection (LOD) and limits of quantification (LOQ) ranged from 2.75 to 7.44 μg/mL and 8.33 to 22.54 μg/mL, respectively. Intra-day and inter-day precision showed relative standard deviation (RSD) values below 2.85% and 2.72%, while recovery values ranged from 96.87% to 107.72%, confirming the accuracy and reliability of the method. Quantitative results revealed that ellagitannins were the predominant constituents in both stems and pericarpium of P. granatum. Biological evaluation demonstrated that several isolated compounds exhibited strong α-glucosidase inhibitory activity, with compounds 1–3 showing significantly stronger activity than the reference inhibitor acarbose. Enzyme kinetic studies and molecular docking analyses were conducted to clarify their inhibition mechanisms, revealing mixed, uncompetitive and competitive inhibition modes associated with binding to catalytic or allosteric sites of the enzyme. This study integrates quantitative analysis with mechanistic enzyme inhibition evaluation, providing a practical analytical tool for quality control of P. granatum and scientific evidence supporting its potential application in antidiabetic therapy.
Keywords:
Punica granatum L., HPLC-DAD, Simultaneous determination, α-Glucosidase, Molecular dockingIntroduction
Pomegranate (Punica granatum L.), a member of the Punicaceae family, is widely cultivated worldwide. Beyond its culinary uses, various parts of the pomegranate plant, including the juice, peels, seeds, flowers, leaves, and bark, have long been utilized in traditional medicine.1 Recent studies have focused on examining the chemical composition of these different plant parts, while also exploring the pharmacological activities associated with their extracts.2–5 Phytochemical investigations have revealed that pomegranate is rich in polyphenolic compounds such as ellagitannins, gallotannins, phenolic acids, and flavonoids.6–8 Pomegranate fruit pieces (peels, seeds, and juice) extract has been documented to possess a variety of pharmacological benefits, including antidiabetic, anti-inflammatory, antioxidant, and hepatoprotective effects.9–12 Additionally, extracts from pomegranate leaves and bark have been shown to offer health-promoting properties, including antinociceptive, antitumor, antibacterial, anti-inflammatory, and antidepressant effects, as observed in both animal studies and human diseases.13–15
The quality of herbal plants can vary significantly due to factors such as origin, distribution location, cultivation methods, and processing techniques. This variability highlights the importance of developing a systematic approach to the analysis and evaluation of medicinal herbs. Recent advancements in analytical methods, particularly high-performance liquid chromatography (HPLC), have enabled more precise analyses of individual components in these natural products.16–20 Although pomegranate is rich in polyphenolic compounds and has shown antidiabetic potential, the α-glucosidase inhibitory activities and mechanisms of its individual constituents remain insufficiently understood.21–23
Therefore, this study aims to develop a reliable HPLC-DAD method for the simultaneous quantification of nine major compounds isolated from the stems and pericarpium of P. granatum, to compare their contents, and to evaluate the α-glucosidase inhibitory activity and mechanisms of the principal constituents.
Experimental
General experimental procedures – 1H (500 MHz) and 13C (125 MHz) nuclear magnetic resonance (NMR) and 2D-NMR were conducted using a Bruker Avance Digital 500 MHz spectrometer (Karlsruhe, Germany) (in ppm rel. to tetramethylsilane (TMS) as internal standard) (NFEC-2025-02-303505) at the Medibio Core Facility of Kyungpook National University. HPLC was conducted using a Waters Alliance system, comprising a Waters 2998 Photodiode Array (PDA) Detector, a Waters 1525 Binary HPLC pump, and an Agilent Eclipse XD8-C18 column (20 × 250 mm, 4 μm). Column chromatography (CC) was carried out on Merck silica gel (60–200 μm), Merck Lichroprep RP-18 (40–63 µm), MCI gel (75–150 μm), and Sephadex LH-20 (25–100 μm). Thin-layer chromatography (TLC) was performed using glass plates coated with silica gel 60 F254 and RP-18 F254S (Merck , Germany). Compounds were detected under UV light and then visualized by spraying the plates with 10% sulfuric acid reagent followed by heating at 110°C for 1–2 minutes. The solvents for extraction and isolation were supplied by Daejung Chemicals (Gyeonggi-do, Korea), while HPLC-grade acetonitrile was purchased from Burdick & Jackson (Muskegon, MI, USA). The water was purified through the Milli-Q water purification system from Merck Millipore.
Plant material – The stems of Punica granatum were collected from the Kyungpook National University Herb Garden in January 2022, while the pericarpium was purchased from a traditional market at Jeonju-si, Korea in December 2022. Plant identification was verified by Professor Byung Sun Min at Daegu Catholic University. Voucher specimens of the stems and pericarpium (24A–PG and 24B–PGP) were deposited in the Laboratory of Pharmacognosy, College of Pharmacy, Kyungpook National University, Korea.
Extraction and isolation – The dried stems of P. granatum (PG) (18.4 kg) were extracted with methanol (MeOH, 10 L × 3 times) under reflux for 4 hours and evaporated under reduced pressure to give a MeOH residue (1.40 kg). The MeOH residue was suspended in water and successively partitioned with n-hexane, methylene chloride (CH2Cl2), ethyl acetate (EtOAc), butanol (BuOH) to yield a n-hexane extract (99.7 g) CH2Cl2 extract (63.1 g), EtOAc extract (409.9 g), BuOH extract (167.2 g) and water layer after removal of the solvents.
The EtOAc extract of PG (409.9 g) was subjected to vacuum liquid chromatography on silica gel using a gradient of CH2Cl2-MeOH (9:1 → 1:1, v/v) to yield six fractions, 1A–1F. Fraction 1B (13.2 g) was separated by silica gel CC and eluted with CH2Cl2-MeOH (30:1 → 1:1, v/v), to obtain subfractions, 2A–2E. Fraction 2B (2.1 g) was chromatographed by reverse phase C18 (RP-18) CC using MeOH-H2O (1:1 → 2:1, v/v) to obtain compounds 9 (75.1 mg). Fraction 2D (130.6 g) was further separated on silica gel CC using a mixture of solvents EtOAc-MeOH-H2O (6:1:0.1, v/v/v), to give six fractions 3A–3F. Subfraction 3B (11.9 g) was fractioned by silica gel CC, eluting with EtOAc-MeOH-H2O (9:1:0.1, v/v/v), to give six subfraction 3B1–3B6. Subfraction 3B2 (5.2 g) was separated by MCI CC using MeOH–H2O (1:2, v/v) to give 4A–4H. Fraction 4A (1.6 g) was further separated by RP-18 CC using acetone–H2O (1:4, v/v), followed by Sephadex LH-20 CC with MeOH–H2O (1:3, v/v) as eluent to obtain compound 3 (138.6 mg). Compounds 4 (76.9 mg) was isolated from subfraction 4C (529.7 mg) by RP-18 CC using acetone–H2O (1:4, v/v). From fraction 3B4 (589.2 mg), compounds 8 (83.8 mg) and 7 (36.3 mg) were isolated by RP-18 CC using acetone-H2O (1:2, v/v).
The BuOH extract of PG (167.2 g) was separated on silica gel using a gradient of CH2Cl2-MeOH (9:1 → 0:1, v/v) to afford 5A–5F. Fraction 5C (36.7 g) was further separated by silica gel CC using a gradient of CH2Cl2-MeOH (9:1 → 0:1, v/v) to give 5C1–5C5. From fraction 5C3 (5.7 g), compounds 6 (110.9 mg) was isolated by RP-18 CC using acetone-H2O (1:4, v/v).
The dried pericarpium of P. granatum (PGP) (11.4 kg) was extracted three times with MeOH (10 L × 4 h) under reflux, then the solvent was removed under reduced pressure and gave a MeOH extract (4.1 kg). The MeOH extract was suspended in water and then successively partitioned with n-hexane, EtOAc, BuOH to afford a n-hexane extract (120.8 g), EtOAc extract (100.9 g), BuOH extract (455.2 g) and water layer.
The EtOAc extract of PGP (100.9 g) was subjected to silica gel CC using a gradient of CH2Cl2-MeOH (20:1 → 0:1, v/v) to yield seven fraction 6A–6G. Fraction 6E (16.1 g) was separated by MCI CC using gradient of MeOH-H2O (1:2 → 1:0, v/v) to yield subfraction 6E1–6E8. From fraction 6E4 (1.26g), compound 5 (20.8 mg) was isolated by using RP-18 CC eluting with aceton-H2O (1:3, v/v).
The BuOH extract of PGP (455.2 g) was separated on silica gel using a gradient of CH2Cl2-MeOH (15:1 → 0:1, v/v) to afford eight fraction 7A–7H. Fraction 7D (57.7 g) was separated by silica gel CC using CH2Cl2-MeOH (10:1 → 0:1, v/v) to give four fraction 7D1–7D4. Compounds 8 (202.1 mg) was isolated from fraction 7D1 (1.1 g) by RP-18 CC eluting with acetone-H2O (4:1 → 1:1, v/v). Fraction 7D3 (27.5 g) was chromatographed by MCI CC using a gradient of MeOH-H2O (1:2 → 1:0, v/v) to afford 8A–8E. From fraction 8B (2.5 g), compounds 1 (35.8 mg) were isolated by RP-18 CC using a gradient of MeOH-H2O (1:4 → 1:0, v/v) as eluent. From fraction 8E (2.1 g), compound 6 (58.6 mg) was isolated by crystallization in MeOH. Fraction 7F (56.3 g) was subjected to silica gel CC using a gradient of EtOAc-MeOH-H2O (8:1:0.1 → 2:1:0.1, v/v/v) to yield five subfraction 9A–9E. Compounds 2 (79.4 mg) and 4 (74.9 mg) were isolated from fraction 9B (1.7 g) by RP-18 CC with MeOH-H2O (1:4, v/v) as eluent.
Pedunculagin (1) – Pale amorphous powder. Mixture of α and β-anomers (1a and 1b). 1a: 1H-NMR (500 MHz, methanol-d4): δ 6.62, 6.61, 6.50, 6.37 (each 1H, s, HHDP H-3), 5.48 (1H, t, J = 9.7 Hz, α-Glc-3), 5.37 (1H, d, J = 3.8 Hz, α-Glc-1), 5.34 (1H, m, α-Glc-6a), 5.10 (2H, m, α-Glc-2, and 4), 4.59 (1H, ddd, J = 1.5, 7.0, 10.0 Hz, α-Glc-5), 3.83 (1H, dd, J = 1.5, 13.0 Hz, α-Glc-6b). 13C-NMR (125 MHz, methanol-d4): δ 170.81, 170.38, 169.72, 169.07 (HHDP C-7), 145.89, 145.87, 145.85, 145.78, 145.01, 144.86, 144.72, 144.62 (HHDP C-4 and 6), 137.51, 137.51, 137.17, 137.11 (HHDP C-5), 126.69, 126.48, 126.28, 126.06 (HHDP C-2), 116.61, 116.54, 115.57, 114.98 (HHDP C-1), 108.63, 107.92, 107.88, 107.75 (HHDP C-3), 92.27 (α-Glc-1), 76.46 (α-Glc-3), 76.28 (α-Glc-2), 70.44 (α-Glc-4), 67.80 (α-Glc-5), 64.19 (α-Glc-6). 1b: 1H-NMR (500 MHz, methanol-d4): δ 6.62, 6.59, 6.54, 6.37 (each 1H, s, HHDP H-3), 5.26–5.32 (2H, m, β-Glc-3, and 6a), 5.13 (1H, m, β-Glc-4), 4.97 (1H, d, J = 8.2 Hz, β-Glc-1), 4.87 (1H, m, β-Glc-2), 4.16 (1H, ddd, J = 1.0, 6.5, 10.0 Hz, β-Glc-5), 3.89 (1H, dd, J = 1.0, 13.0 Hz, β-Glc-6b); 13C-NMR (125 MHz, methanol-d4): δ 170.85, 170.23, 169.59, 169.12 (HHDP C-7), 145.93, 145.87, 145.87, 145.84, 145.04, 144.88, 144.80, 144,71 (HHDP C-4 and 6), 137.54, 137.54, 137.47, 137.20 (HHDP C-5), 126.48, 126.48, 126.23, 125.98 (HHDP C-2), 116.61, 116.57, 115.64, 114.89 (HHDP C-1), 108.67, 108.20, 107.83, 107.66 (HHDP C-3), 95.80 (β-Glc-1), 78.80 (β-Glc-2), 78.22 (β-Glc-3), 73.11 (β-Glc-5), 70.01 (β-Glc-4), 64.13 (β-Glc-6).
Flavogallonic acid (2) – Yellow amorphous powder. 1H-NMR (500 MHz, methanol-d4): δ 7.40, 7.30 (each 1H, s, H-3ʹ and 3ʹʹ); 13C-NMR (125 MHz, methanol-d4): δ 170.85, 161.55, 160.14 (C-7, 7ʹ and 7ʹʹ), 148.84, 147.20, 145.35, 144.30, 140.27, 139.86, 139.11, 137.52, 136.86 (C-4, 4ʹ, 4ʹʹ, 5, 5ʹ, 5ʹʹ, 6, 6ʹ, and 6ʹʹ), 125.96, 121.23, 119.10 (C-2, 2ʹ, and 2ʹʹ), 114.40, 114.03, 111.76 (C-1, 1ʹ, and 1ʹʹ), 111.26, 109.22, 108.91 (C-3, 3ʹ, and 3ʹʹ).
Casuarinin (3) – Pale amorphous powder. Pale amorphous powder. 1H-NMR (500 MHz, methanol-d4): δ 7.09 (2H, s, Galloyl H-2 and 6), 6.83, 6.53, 6.41 (each 1H, s, HHDP H-3 and 3ʹ), 5.52 (1H, d, J = 5.0 Hz, H-1), 5.47 (1H, dd, J = 2.0, 9.0 Hz, H-4), 5.38 (1H, t, J = 2.0 Hz H-3), 5.33 (1H, dd, J = 3.0, 9.0 Hz, H-5), 4.93 (1H, dd, J = 3.0, 13.0 Hz, H-6a), 4.70 (1H, dd, J = 2.0, 5.0 Hz, H-2), 4.08 (1H, d, J = 13.0 Hz, H-6b); 13C-NMR (125 MHz, methanol-d4): δ 170.81, 170.27, 169.35, 167.02 (HHDP C-7 and 7ʹ), 166.93 (Galloyl C-7), 146.76, 146.41, 145.79, 145.75, 144.80, 144.71, 144.31, 144.16 (HHDP C-4, 4ʹ, 6 and 6ʹ) 146.28 (Galloyl C-3 and 5), 140.14 (Galloyl C-4), 139.80, 137.70, 136.86, 135.59 (HHDP C-5 and 5ʹ), 127.31, 126.81, 124.94, 119.96 (HHDP C-2 and 2ʹ), 120.57 (Galloyl C-1), 117.73, 116.62, 115.76, 116.46 (HHDP C-1 and 1ʹ), 110.30 (Galloyl C-2 and 6), 109.06, 107.54, 105.07 (HHDP C-3 and 3ʹ), 77.82 (H-2), 74.58 (H-4), 71.51 (H-5), 70.45 (H-3), 67.59 (H-1), 64.99 (H-6).
Isocorilagin (4) – Pale amorphous powder. 1H-NMR (500 MHz, methanol-d4): δ 7.06 (2H, s, Galloyl H-2 and 6), 6.69, 6.66 (each 1H, s, HHDP H-3 and 3ʹ), 6.37 (1H, d, J = 2.0 Hz, Glc-1), 4.96 (1H, t, J = 11.0 Hz, Glc-5), 4.81 (1H, m, Glc-3), 4.52 (1H, dd, J = 8.0, 10.5 Hz, Glc-6a), 4.47 (1H, d, J = 3.0 Hz, Glc-4), 4.16 (1H, dd, J = 8.0, 10.5 Hz, Glc-6b), 3.99 (1H, d, J = 2.0 Hz, Glc-2); 13C-NMR (125 MHz, methanol-d4): δ 170.11, 168.51 (HHDP C-7 and 7ʹ), 166.67 (Galloyl C-7), 146.37 (Galloyl C-3 and 5), 146.03, 145.61 (HHDP C-4 and 4ʹ), 145.30, 145.20 (HHDP C-6 and 6ʹ), 140.39 (Galloyl C-4), 138.16, 137.66 (HHDP C-5 and 5ʹ), 125.47, 125.43 (HHDP C-2 and 2ʹ), 120.59 (Galloyl C-1), 117.18, 116.68 (HHDP C-1 and 1ʹ), 110.92 (Galloyl C-2 and 6), 110.13, 108.29 (HHDP C-3 and 3ʹ), 95.00 (Glc-1), 76.18 (Glc-5), 71.64 (Glc-3), 69.45 (Glc-2), 65.01 (Glc-6), 62.44 (Glc-4).
Methyl brevifolincarboxylate (5) – Yellow amorphous powder. 1H-NMR (500 MHz, DMSO-d6): δ 7.30 (1H, s,H-3ʹ), 4.41 (1H, dd, J = 1.8, 7.7 Hz, H-4), 3.62 (3H, s, OCH3), 2.97 (1H, dd, J = 7.7, 18.7 Hz, H-5b), 2.46 (1H, d, J = 1.8 Hz, H-5a); 13C-NMR (125 MHz, DMSO-d6): δ 193.04 (C-1), 172.54 (C-6), 160.14 (C-7ʹ), 149.69 (C-2), 145.87 (C-4ʹ), 143.60 (C-6ʹ), 140.33 (C-3), 138.42 (C-5ʹ), 115.01 (C-2ʹ), 113.05 (C-1ʹ), 108.17 (C-3ʹ), 52.07 (OCH3), 40.63 (C-5), 36.99 (C-4).
Eschweilenol C (6) – White amorphous powder. 1H-NMR (500 MHz, methanol-d4): δ 7.92, 7.55 (each 1H, s, H-3 and 3ʹ), 5.60 (1H, d, J = 1.5 Hz, Rha-1), 4.23 (1H, m, Rha-2), 4.03 (1H, dd, J = 3.3, 9.5 Hz, Rha-3), 3.77 (1H, m, Rha-5), 3.55 (1H, t, J = 9.5 Hz, Rha-4), 1.31 (1H, d, J = 6.3 Hz, Rha-6); 13C-NMR (125 MHz, methanol-d4): δ 161.82, 161.74 (C-7 and 7ʹ), 150.46, 148.47, 148.47, 148.35 (C-4, 4ʹ, 6 and 6ʹ), 138.28, 138.08 (C-5 and 5ʹ), 116.56, 113.90 (C-2 and 2ʹ), 113.79, 111.21 (C-3 and 3ʹ), 109.03, 107.14 (C-1 and 1ʹ), 101.51 (Rha-1), 73.83 (Rha-3), 72.07 (Rha-5), 71.82 (Rha-4), 71.12 (Rha-2), 18.03 (Rha-6).
Ellagic acid-4-O-β-D-xylopyranoside (7) – White amorphous powder. 1H-NMR (500 MHz, methanol-d4): δ 7.62, 7.30 (each 1H, s, H-3 and 3ʹ), 5.70 (1H, dd, J = 8.3, 2.0 Hz, Xyl-1), 4.39–4.06 (3H, m, Xyl-2, 3 and 4), 3.78 (1H, dd, J = 3.5, 12.0 Hz, Xyl-5a), 3.70 (1H, dd, J = 5.0, 12.0 Hz, Xyl-5b); 13C-NMR (125 MHz, methanol-d4): δ 160.96, 160.92 (C-7 and 7ʹ), 149.80, 147.52, 142.70, 140.73 (C-4, 4ʹ, 6 and 6ʹ), 137.51, 137.19 (C-5 and 5ʹ), 115.71, 112.97 (C-2 and 2ʹ), 112.79, 111.82 (C-3 and 3ʹ), 109.65 (Xyl-1), 108.96 (C-1 and 1ʹ), 88.36 (Xyl-3), 82.71 (Xyl-2), 78.56 (Xyl-4), 62.97 (Xyl-5).
Ellagic acid (8) – White amorphous powder. 1H-NMR (500 MHz, methanol-d4): δ 7.47 (each 2H, s, H-3 and 3ʹ); 13C-NMR (125 MHz, methanol-d4): δ 161.44 (C-7 and 7ʹ), 149.43 (C-4 and 4ʹ), 140.91 (C-6 and 6ʹ), 137.60 (C-5 and 5ʹ), 114.00 (C-2 and 2ʹ), 111.72 (C-3 and 3ʹ), 109.45 (C-1 and 1ʹ).
Quercitrin (9) – Yellow amorphous powder. 1H-NMR (500 MHz, DMSO-d6): δ 6.63 (1H, d, J = 2.2 Hz, H-2ʹ), 6.59 (1H, dd, J = 2.2, 8.3 Hz, H-6ʹ), 6.20 (1H, d, J = 8.3 Hz, H-5ʹ), 5.72 (1H, d, J = 2.1 Hz, H-8), 5.54 (1H, d, J = 2.1 Hz, H-6), 4.59 (1H, d, J = 1.4 Hz, Rha-1), 3.31 (1H, brs, Rha-2), 2.84 (1H, dd, J = 2.9, 9.1 Hz, Rha-3), 2.55 (2H, m, Rha-4,5), 0.15 (3H, d, J = 6.1 Hz, CH3); 13CNMR (125 MHz, DMSO-d6): δ 177.79 (C-4), 164.27 (C-7), 161.34 (C-5), 157.35 (C-2), 156.49 (C-9), 148.48 (C-4ʹ), 145.25 (C-3ʹ), 134.25 (C-3), 121.16 (C-6ʹ), 120.77 (C-1ʹ), 115.68 (C-5ʹ), 115.50 (C-2ʹ), 104.10 (C-10), 101.86 (Rha-1), 98.73 (C-6), 93.67 (C-8), 71.20 (Rha-4), 70.64 (Rha-5), 70.38 (Rha-3), 70.10 (Rha-2), 17.54 (Rha-6).
HPLC-DAD analysis – The analysis of P. granatum solutions was performed using HPLC-DAD. The HPLC system used was from Waters, MA, USA, featuring a Waters 1525 Binary HPLC pump, and a Waters 2998 Photodiode Array (PDA) Detector. Each sample was injected and separated using an Agilent Eclipse XD8-C18 column (4.6 × 150 mm, 5 µm) at a temperature of 25°C. The mobile phase consisted of methanol (A) and 0.1% formic acid in water (B). The gradient elution program was set as follows: 95–90% B (0–20 min), 90–60% B (20–50 min), 60–58% B (50–60 min), 58–50% B (60–80 min), 50–0% B (80–90 min), and 0% B (90–95 min), followed by re-equilibration at 95% B from 95 to 100 min. The flow rate was set at 1.0 mL/min, and the injection volume was 10 µL. UV wavelengths were configured to 210, 254, and 280 nm, with chromatograms acquired at 254 nm to effectively capture all peaks simultaneously.
Preparation of standard solutions and analysis samples – Standard stock solutions were prepared by dissolving nine isolated compounds pedunculagin (1), flavogallonic acid (2), casuarinin (3), isocorilagin (4), methyl brevifolincarboxylate (5), eschweilenol C (6), ellagic acid-4-O-β-D-xylopyranoside (7), ellagic acid (8), and quercitrin (9) in HPLC-grade MeOH at a concentration of 1 mg/mL. These stock solutions were then serially diluted with HPLC-grade MeOH to prepare the working solutions. After filtration through a 0.20 μm PTFE filter, the working solutions were used to establish the calibration curve. The PG and PGP extracted were dissolved in HPLC-grade MeOH at a concentration of 10 mg/mL. The P. granatum sample was also filtered through 0.20 μm PTFE filter before being injected into the HPLC system.
Validation of method – Stock solutions of nine compounds (1–9) were diluted to concentration of 5, 10, 50, 100, and 200 μg/mL for HPLC analysis. Calibration curves were constructed by plotting the peak areas (y) against the concentrations of the analytes (x), using regression equations in the form of y = ax + b. Linearity was assessed using the least squares treatment (R2). Each working standard concentration was analyzed in triplicate.
The limit of detection (LOD) was determined as the minimum concentration of the sample, while the limit of quantification (LOQ) was defined as the lowest concentration of the compound. This was established by injecting a diluted standard solution and achieving a signal-to-noise ratio in the range of 3.3 to 10.
The precision of the method was evaluated through intra-day (n = 5) and inter-day (n = 5) injections. For both assessments, three different concentrations (50, 150, and 300 μg/mL) were tested, as verified by the calibration curves. Repeatability and precision were measured using the relative standard deviation (RSD, %). The accuracy of the method was evaluated through a recovery study. This involved adding known amounts of compound solutions to the P. granatum extract sample. Three different concentrations (50, 150, and 300 μg/mL) of compounds 1–9 were added to the samples. The recovery percentage was calculated using the equation: (detected amount – original amount)/spiked amount × 100.
α-Glucosidase inhibitory assay – The α-glucosidase inhibitory activity was assessed using a spectrophotometric assay with p-nitrophenyl α-D-glucopyranoside (p-NPG) as the substrate, in accordance with a previously described protocol.24 The assay was conducted in 96-well microplates, and enzyme activity was determined by measuring the absorbance of the released p-nitrophenol at 405 nm using a microplate spectrophotometer (Agilent, CA, USA). Reaction mixtures comprised α-glucosidase enzyme, varying concentrations of the test compound, and the p-NPG substrate. Acarbose was included as the positive control. The reactions were performed under optimized experimental conditions. The extent of enzyme inhibition was calculated by comparing absorbance values of the tested samples with those of the control. Inhibitory activity was expressed as IC50 values, representing the concentration of each compound required to inhibit 50% of α-glucosidase activity. The percentage inhibition was calculated using the following equation:
where ΔC is the control intensity and ΔS is the intensity of the inhibitor.
α-Glucosidase kinetic analysis – To clarify the mode of α-glucosidase inhibition, enzyme kinetic analyses were conducted using Lineweaver–Burk plots, following established protocols.24 The enzymatic reactions were conducted with varying concentrations of p-nitrophenyl α-D-glucopyranoside (p-NPG) substrate (0.625, 1.25, and 2.5 mM) in the presence of different concentrations of the test compound. The inhibition mechanism was determined by examining the intersection patterns of the Lineweaver–Burk plots. Intersections occurring on the y-axis indicated competitive inhibition, whereas intersections on the x-axis were characteristic of non-competitive inhibition. Parallel lines in the plots suggested uncompetitive inhibition, reflecting inhibitor binding exclusively to the enzyme-substrate complex. Mixed-type inhibition was identified when the trendlines intersected within the x–y quadrant. The inhibition constants (Ki) were subsequently calculated using secondary plots and Dixon plots analysis to evaluate the binding affinity between the enzyme and the inhibitor. All kinetic data and graphical representations were generated and analyzed using GraphPad Prism 10.
Molecular docking simulations – Molecular docking simulations were carried out using AutoDock Vina 4.2 to predict binding affinity and interaction modes between the active compounds and α-glucosidase. Since the crystal structure of Saccharomyces cerevisiaeα-glucosidase has not been reported, a three-dimensional protein model was generated by homology modeling using the SWISS-MODEL, with the α-glucosidase amino acid sequence (PDB ID: 3AJ7) as a template. The generated model was energy-minimized using the GROMOS96 force field, and its structural quality was evaluated using the MolProbity web server. The 3D structure of compounds 1, 2 and 3 were constructed from ChemDraw 3D Professional 19 and subsequently optimized using Avogadro 2 (Kitware, Inc., USA). Protein preparation steps, including the removal of water molecules, addition of hydrogen atoms, and assignment of rotatable bonds, were performed using AutoDock Tools version 1.5.7. Blind docking simulations were conducted using a grid box with dimensions of 126 × 126 × 126 points, centered at coordinates X = 24.227, Y = –2.294, and Z = 17.943. The resulting docking poses were analyzed and visualized using PyMOL, while two-dimensional ligand-protein interaction diagrams were generated with Discovery Studio Visualizer 21.1 (Accelrys, Inc., USA).24
Statistics analysis – All data are expressed as the mean ± standard error of the mean (SEM) based on at least three independent experiments. Statistical significance among groups was evaluated using one-way analysis of variance (ANOVA), followed by Duncan’s test (Systat Inc., Evanston, IL, USA). Differences were considered statistically significant at p < 0.05.
Results and Discussion
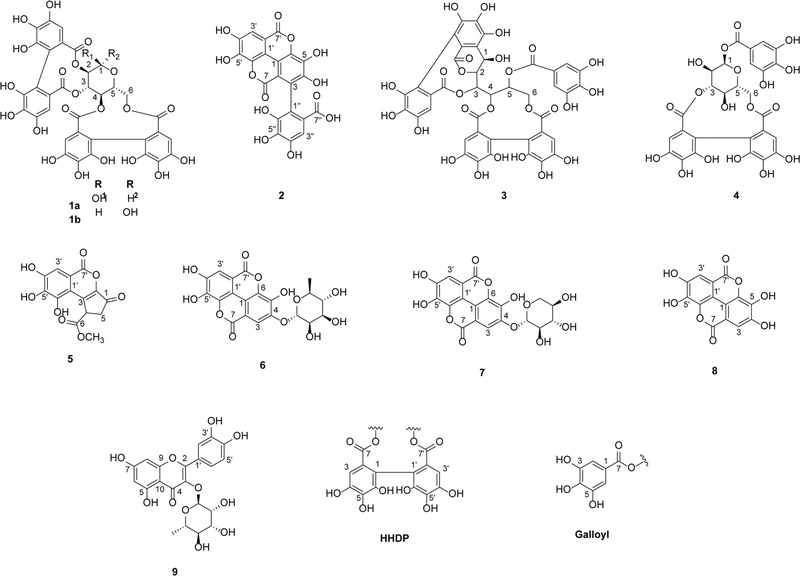
The reference standards pedunculagin (1),25 flavogallonic acid (2),26 casuarinin (3),27 isocorilagin (4),28 methyl brevifolincarboxylate (5),29 eschweilenol C (6),30 ellagic acid-4-O-β-D-xylopyranoside (7),31 ellagic acid (8),32 and quercitrin (9)33 (Fig. 1) were isolated and purified from the PG and PGP employing various chromatographic methods. By comparing the NMR data with previously reported values in the literature, these structures were identified.
Chromatographic conditions were developed for the simultaneous determination of major compounds of PG and PGP extracts. Preliminary tests were conducted to determine effective HPLC-DAD conditions, including selection a suitable Agilent Eclipse XD8-C18 reversed-phase column. The chosen mobile phase consisted of a multi-step gradient solution system of MeOH (A) and 0.1% formic acid in water (B). The DAD detector wavelength was tested at 210, 254, and 280 nm, with chromatograms selected based on the UV spectrum at 254 nm. Each compound peak was verified by comparing retention time and UV spectrum with those of the corresponding reference standards. The chromatograms encompass all identified compounds in P. granatum (Fig. 2), the separated compounds showed retention times of 9.8 and 19.1 minutes for pendunculagin (1), 28.2 minutes for flavogallonic acid (2), 31.6 minutes for casuarinin (3), 40.8 minutes for isocorilagin (4), 47.2 minutes for methyl brevifolincarboxylate (5), 54.7 minutes for eschweilenol C (6), 55.7 minutes for ellagic acid-4-O-β-D-xylopyranoside (7), 56.7 minutes for ellagic acid (8) and 60.7 minutes for quercitrin (9). Although identical chromatographic conditions were applied, the MeOH extracts of pericarpium and stems were analyzed and validated separately due to differences in their chemical composition and peak distribution patterns. Chromatographic profiling revealed variations in the relative abundance of major constituents and matrix complexity between the two plant parts, which may influence analytical performance parameters such as sensitivity, precision, and recovery. Therefore, separate validation was conducted to ensure accurate and reliable quantification within each specific plant matrix.
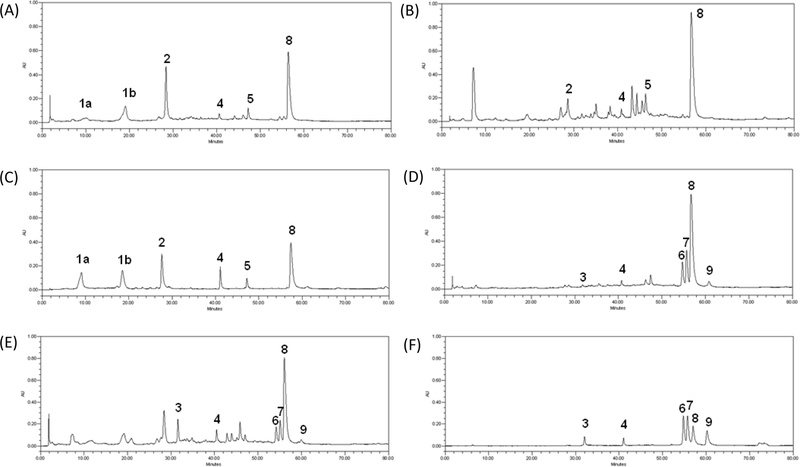
HPLC chromatograms of the MeOH extract (A), EtOAc extract (B), and the standard mixture (C) of PGP; and the MeOH extract (D), EtOAc extract (E), and the standard mixture (F) of PG.
To validate this method, experiments were conducted to verify linearity, detection and quantification limits, precision and accuracy, and recovery. Regression equations were established based on five concentration trials of each standard, performed in triplicate. High correlation coefficients (R² > 0.9991) indicate excellent linearity of all calibration curves within the specified test ranges. According to this linear regression, the limits of detection (LOD) ranged from 2.75 to 7.44 μg/mL, and the limits of quantification (LOQ) were within the range of 8.33 to 22.54 μg/mL, respectively (Table 1). This suggests that the method can detect and quantify compounds at minimal concentrations.
Calibration curves, correlation coefficients (R2), limits of detection (LOD) and limits of quantification (LOQ) of compounds 1–9
To evaluate the method's repeatability and precision, both intra-day analysis (n = 5) and inter-day (n = 5) analyses were conducted. Intra-day testing was repeated three times within a single day, while inter-day testing was performed on three different days. The Relative Standard Deviation (RSD) values for intra-day testing varied from 0.54% to 2.85%, while for inter-day testing, they ranged from 0.11% to 2.72% (Table 2). The intraday accuracy was in the range of 97.54 to 106.61% and the inter-day accuracy was 97.50 to 107.72%. These results demonstrate the method's high reproducibility and precision. All analyzed compounds exhibited recovery within the optimal range, and RSDs remained consistently low. This data highlights the method's high accuracy. In summary, the comprehensive results confirm the method's suitability for the quantitative analysis of P. granatum samples.
As shown in Table 3, the chromatographic separation of compounds 1–9 in the extracts of PG and PGP was well attained by the developed method. The comparison between the MeOH extract of PGP and the MeOH extract of PG highlights significant differences in compound diversity and concentration. The MeOH extract of PGP contains compounds 1a (3.47%) and 1b (13.91%), which are not determined in the MeOH extract of PG. Additionally, it has a significantly higher concentration of compound 2 (23.00%), whereas the MeOH extract of PG shows the absence of this compound. Another difference is the presence of compounds 3 (0.69%) and 9 (2.19%) in the MeOH extract of PG, which cannot be observed in the MeOH extract of PGP. Both extracts share compounds 4–8, but the MeOH extract of PG generally exhibits higher concentrations of these compounds, except for compounds 5 which is more abundant in the MeOH extract of PGP. Compound 8 is founded in both extracts with the highest levels, particularly in MeOH extract of PG (59.21%), while in the MeOH extract of PGP, compound 8 is present at a lower level (41.23%). Moreover, compounds 6 and 7 are also present in both extracts, but their concentration is significantly higher in MeOH extract of PG (8.81% and 14.27%, respectively) compared to MeOH extract of PGP (0.70% and 0.32%, respectively). Compounds 1a, 1b and 2 were detected exclusively in PGP, suggesting that they may serve as characteristic markers for this plant part. Meanwhile, compounds 3, 6 and 7 may be more suitable markers for PG. The comparison between the MeOH and EtOAc extracts of both PG and PGP reveals that the major compounds in the MeOH extract are primarily distributed in the EtOAc extract and at high concentrations. The solvent dependent distribution provides important insight into the phytochemical characteristics of the plant materials and confirms that the EtOAc fraction contains a high concentration of key bioactive compounds. Therefore, this fraction represents a suitable and rational choice for subsequent bioactivity evaluation and further pharmacological investigation.
The inhibitory effects of the test compounds on α-glucosidase were evaluated using acarbose as a reference inhibitor, which exhibited an IC50 value of 195.92 ± 1.65 µM. Except for compound 5, most compounds displayed strong inhibitory activity against α-glucosidase (Table 4). Compounds 1–4 showed pronounced inhibitory effects, with IC50 values of 0.46 ± 0.01 µM, 1.89 ± 0.07 µM, 1.51 ± 0.06 µM and 1.32 ± 0.03 µM, respectively, markedly surpassing the inhibitory potency of acarbose. Given that the α-glucosidase inhibitory activities of compounds 4 and 6–9 have been previously reported,34–38 further kinetic investigations were focused on compounds 1–3. The inhibition mechanisms were examined using Lineweaver-Burk plots, which enable classification of inhibition types based on the intersection patterns of the plotted lines. As shown in Fig. 3A, compound 1 produced intersecting lines in the x–y quadrant, indicating a mixed-type inhibition mechanism. In contrast, increasing concentrations of compound 2 resulted in a series of parallel lines (Fig. 3D), characteristics of uncompetitive inhibition. Meanwhile, the Lineweaver-Burk plot for compound 3 exhibited line intersections at the y-axis (Fig. 3F), consistent with a competitive inhibition mechanism.

Inhibitory activity of compounds 1–9 from stems and pericarpium of P. granatum against α-glucosidase
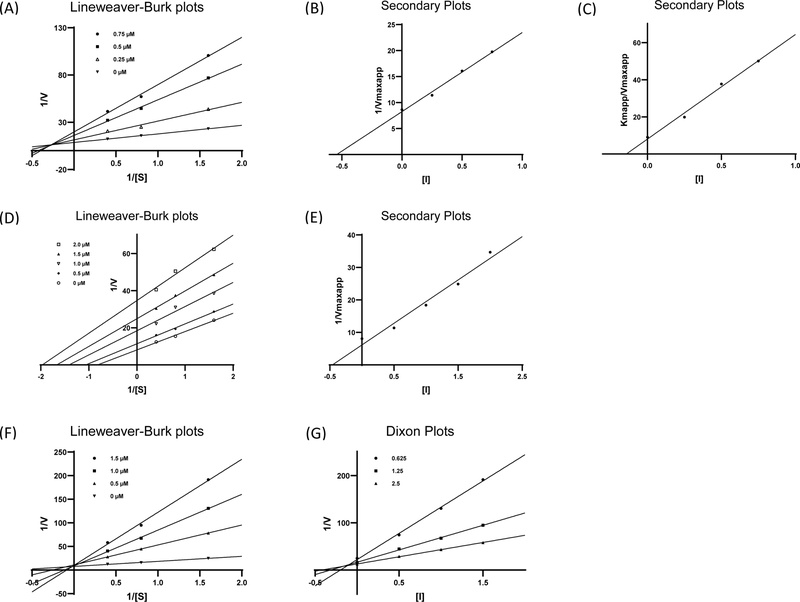
Lineweaver-Burk plots of compounds 1 (A), 2 (D) and 3 (F) with α-glucosidase. Secondary plots of compounds 1 (B, C), 2 (E) and Dixon Plots of compounds 3 (F) with α-glucosidase.
Based on the kinetic analyses, Dixon plots and secondary plots were further employed to calculate the inhibition constants (Ki) of the tested compounds. For compound 2, only the uncompetitive inhibition constant (Kiu) was determined, whereas both the competitive (Kic) and uncompetitive (Kiu) constants were obtained for compound 1 and 3. Compound 1 was identified as a mixed-type inhibitor, exhibiting a Kic value of 0.14 µM and a Kiu value of 0.54 µM (Fig. 3B and C). The higher Kiu compared with Kic indicates that compound 1 preferentially binds to the free enzyme rather than the enzyme-substrate complex. As shown in Fig. 3E and G, compounds 2 and 3 displayed Kiu and Kic values of 0.46 µM and 0.10 µM, respectively. Collectively, the low inhibition constant values of compounds 1, 2, and 3 highlight their strong inhibitory potential against α-glucosidase.
Molecular docking results further supported the kinetic findings (Fig. 4A). As summarized in Table 5, compounds 1 and 3 exhibited binding energy –7.9 and –7.3 kcal/mol, respectively, comparable to that of acarbose, a well-known competitive inhibitor, indicating strong interactions with α-glucosidase.39 Compound 1 formed multiple hydrogen bonds between its hydroxyl groups and residues Glu304, Gly306, Thr307, Pro309, Gln322, suggesting direct interaction within the catalytic pocket. These residues are located in the substrate recognition region and are closely associated with catalytic function. Strong hydrogen bonding within this region may stabilize the ligand inside the active cavity and interfere with substrate orientation. In addition, compounds 1 and 3 shared interactions with Asn241, His279, and Glu304, which are considered key gatekeeping residues regulating ligand access to the active site (Fig. 4B and D). Occupation of this region may restrict substrate entry. Aromatic interactions, including π-π T shaped and π-alkyl contacts with residues such as His279 and Trp242, further contributed to binding stability within the hydrophobic environment of the active pocket. These combined interactions suggest that compounds 1 and 3 effectively compete with the natural substrate. This structural behavior is consistent with the competitive inhibition patterns observed in kinetic analysis.
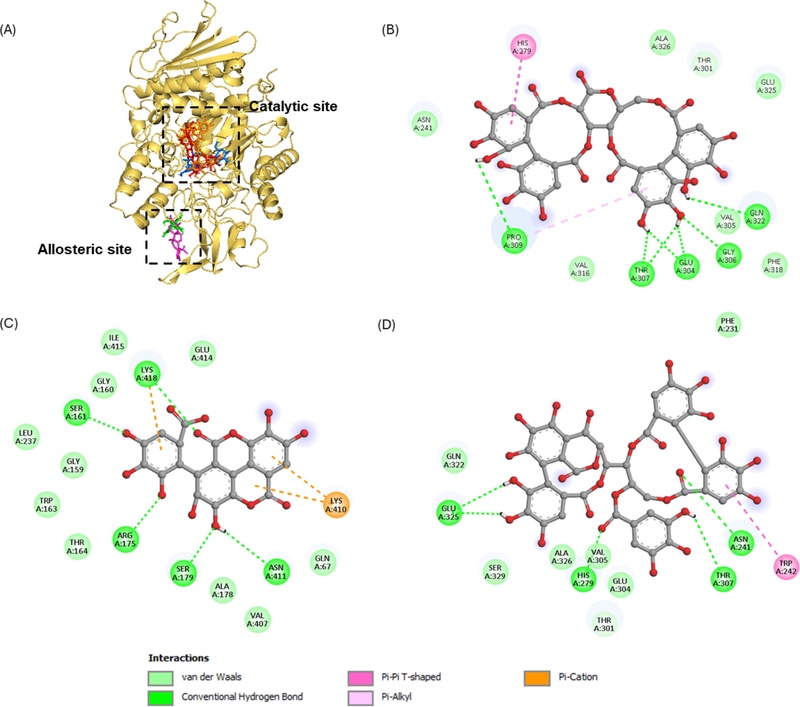
Molecular docking related to α-glucosidase inhibition by acarbose (blue stick), ursolic acid (pink stick), compound 1 (red stick), 2 (green stick), and 3 (orange stick) (A). 2D diagram of α-glucosidase inhibition by compound 1 (B), 2 (C) and 3 (D). The figure was generated using PyMOL and Discovery Studio Visualizer.

Binding site residues and docking scores of compounds 1, 2 and 3 in α-glucosidase, obtained using Autodock 4.2
In contrast, compound 2 exhibited a different binding site. Its interactions were located at a different region involving residues such as Ser161, Arg175, Ser179, Asn411, and Lys418, which are positioned outside the catalytic pocket. Compound 2 interacted with residues that largely overlap with those reported for ursolic acid, a previously described allosteric inhibitor of α-glucosidase. This similarity in interacting residues suggests that compound 2 shares a comparable binding region with ursolic acid.40 The interacting residues correspond to the allosteric site designated as AS-4 in reported structural studies.41 Binding at this allosteric site may induce conformational changes that alter the geometry or dynamics of the catalytic center, thereby reducing enzymatic activity without directly blocking substrate access. This structural observation aligns well with the experimentally determined uncompetitive inhibition mechanism.
In conclusion, nine phenolic compounds: pedunculagin (1), flavogallonic acid (2), casuarinin (3), isocorilagin (4), methyl brevifolincarboxylate (5), eschweilenol C (6), ellagic acid-4-O-β-D-xylopyranoside (7), ellagic acid (8), and quercitrin (9) were isolated from P. granatum stems and pericarpium. Additionally, an HPLC-DAD method was developed and applied for the qualitative and quantitative analysis of the major compounds present in stem and pericapium of P. granatum. The establishment of linear calibration curves enabled the precise quantification of these compounds across a range of concentrations. The validated method demonstrated high sensitivity, repeatability, and precision. Furthermore, significant differences were observed in the content of the main components between PG and PGP extracts, highlighting the chemical distinction between the stem and pericarpium of P. granatum. The major compounds in both samples were ellagitannins. The major constituents identified in both samples were ellagitannins. In the stem extract, the total ellagitannin content was mainly attributed to ellagic acid and two ellagic acid glucoside isomers. Meanwhile, in the pericarpium extract, ellagic acid and flavogallonic acid were the predominant constituents. These compositional differences provide a reliable chemical basis for distinguishing between the stems and pericarpium of P. granatum. The isolated compounds exhibited significant α-glucosidase inhibitory activity, with compounds 1–3 showing markedly stronger potency than acarbose. Kinetic analyses revealed distinct inhibition mechanisms, with compound 1 acting as a mixed-type inhibitor, compound 2 as an uncompetitive inhibitor, and compound 3 as a competitive inhibitor. Molecular docking results were consistent with the kinetic analysis. Compounds 1 and 3 were predicted to bind within the catalytic pocket, interacting with key residues that regulate substrate access and suppress enzymatic activity. Compound 2 binded at the reported AS-4 allosteric site, where binding may induce conformational changes that reduce catalytic efficiency. These findings support distinct inhibition mechanisms and reinforce the potential of the isolated compounds as α-glucosidase inhibitors. Overall, the proposed method provides a valuable tool for the quality control of P. granatum products and offers important insights into future investigations into P. granatum constituents. In addition, the experiment results highlight compounds 1–3 as promising lead candidates for the development of novel α-glucosidase inhibitors.
Acknowledgments
This research was supported by the National Research Foundation of Korea (NRF) funded by the Korea government (MSIT) (No. NRF-2020R1A5A2017323) and Korea Basic Science Institute (National research Facilities and Equipment Center) grant funded by the Ministry of Education (No. RS-2025-02308336). The authors would like to thank Korea Basic Science Institute (KBSI) Daegu Center for the mass spectra measurement service.
Conflicts of Interest
The authors have declared that there are no conflicts of interest.
References
-
Yisimayili, Z.; Chao, Z. Food Chem. 2022, 395, 133600.
[https://doi.org/10.1016/j.foodchem.2022.133600]

-
Li, H.-M.; Kouye, O.; Yang, D.-S.; Zhang, Y.-Q.; Ruan, J.-Y.; Han, L.-F.; Zhang, Y.; Wang, T. Molecules 2022, 27, 4622.
[https://doi.org/10.3390/molecules27144622]

-
Wang, R.-F.; Xie, W.-D.; Zhang, Z.; Xing, D.-M.; Ding, Y.; Wang, W.; Ma, C.; Du, L.-J. J. Nat. Prod. 2004, 67, 2096–2098.
[https://doi.org/10.1021/np0498051]

-
Yuan, T.; Wan, C.; Ma, H.; Seeram, N. P. Planta Med. 2013, 79, 1674–1679.
[https://doi.org/10.1055/s-0033-1350925]

-
Maphetu, N.; Unuofin, J. O.; Masuku, N. P.; Olisah, C.; Lebelo, S. L. Biomed. Pharmacother. 2022, 153, 113256.
[https://doi.org/10.1016/j.biopha.2022.113256]

-
Ruan, J.-H.; Li, J.; Adili, G.; Sun, G.-Y.; Abuduaini, M.; Abdulla, R.; Maiwulanjiang, M.; Aisa, H. A. J. Agric. Food Chem. 2022, 70, 3678–3686.
[https://doi.org/10.1021/acs.jafc.1c08341]

-
Diamanti, A. C.; Igoumenidis, P. E.; Mourtzinos, I.; Yannakopoulou, K.; Karathanos, V. T. Food Chem. 2017, 214, 61–66.
[https://doi.org/10.1016/j.foodchem.2016.07.072]

-
Pagliarulo, C.; De Vito, V.; Picariello, G.; Colicchio, R.; Pastore, G.; Salvatore, P.; Volpe, M. G. Food Chem. 2016, 190, 824–831.
[https://doi.org/10.1016/j.foodchem.2015.06.028]

-
Sun, H.-Y.; Ma, N.; Pan, T.; Du, C.-L.; Sun, J.-Y. Tetrahedron Lett. 2019, 60, 1231–1233.
[https://doi.org/10.1016/j.tetlet.2019.03.056]

-
Singh, R. P.; Chidambara Murthy, K. N.; Jayaprakasha, G. K. J. Agric. Food Chem. 2002, 50, 81–86.
[https://doi.org/10.1021/jf010865b]

-
Wei, X.; Li, S.; Li, T.; Liu, L.; Liu, Y.; Wang, H.; Zhou, Y.; Liang, F.; Yu, X.; Zang, W.; Zhao, M.; Zhao, Z. J. Funct. Food. 2020, 65, 103712.
[https://doi.org/10.1016/j.jff.2019.103712]

-
Kam, A.; Li, K. M.; Razmovski-Naumovski, V.; Nammi, S.; Shi, J.; Chan, K.; Li, G. Q. Phytother. Res. 2013, 27, 1614–1620.
[https://doi.org/10.1002/ptr.4913]

-
Trabelsi, A.; El Kaibi, M. A.; Abbassi, A.; Horchani, A.; Chekir-Ghedira, L.; Ghedira, K. Scientifica 2020, 2020, 8271203.
[https://doi.org/10.1155/2020/8271203]

-
Bekir, J.; Mars, M.; Vicendo, P.; Fterrich, A.; Bouajila, J. J. Med. Food 2013, 16, 544–550.
[https://doi.org/10.1089/jmf.2012.0275]

-
Xiang, Q.; Li, M.; Wen, J.; Ren, F.; Yang, Z.; Jiang, X.; Chen, Y. J. Food Biochem. 2022, 46, e14105.
[https://doi.org/10.1111/jfbc.14105]

-
Yang, W.; Ma, C. J. Nat. Prod. Sci. 2023, 29, 305–311.
[https://doi.org/10.20307/nps.2023.29.4.305]

-
Park, J.; Jang, H. S.; Hong, I.-K.; Yang, H. Nat. Prod. Sci. 2023, 29, 276–280.
[https://doi.org/10.20307/nps.2023.29.4.276]

-
Ma, C. J. Nat. Prod. Sci. 2024, 30, 52–58.
[https://doi.org/10.20307/nps.2024.30.1.52]

-
Ryu, G.; Ma, C. J. Nat. Prod. Sci. 2022, 28, 187–193.
[https://doi.org/10.20307/nps.2022.28.4.187]

-
Phong, N. V.; Lee, Y. M.; Min, B. S.; Kim, J. A. Nat. Prod. Sci. 2024, 30, 65–71.
[https://doi.org/10.20307/nps.2024.30.2.65]

-
Brighenti, V.; Groothuis, S. F.; Prencipe, F. P.; Amir, R.; Benvenuti, S.; Pellati, F. J. Chromatogr. A 2017, 1480, 20–31.
[https://doi.org/10.1016/j.chroma.2016.12.017]

-
Legua, P.; Melgarejo, P.; Abdelmajid, H.; Martinez, J. J.; Martinez, R.; Ilham, H.; Hafida, H.; Hernández, F. J. Food Sci. 2012, 77, C115–C120.
[https://doi.org/10.1111/j.1750-3841.2011.02516.x]

-
Gundogdu, M.; Yilmaz, H. Sci. Hortic. 2012, 143, 38–42.
[https://doi.org/10.1016/j.scienta.2012.05.029]

-
Pham, T. L.; Nguyen, V. P.; Nguyen, T. T. A.; Min, B. S.; Kim, J. A. J. Enzyme Inhib. Med. Chem. 2025, 40, 2584940.
[https://doi.org/10.1080/14756366.2025.2584940]

-
Tanaka, T.; Kirihara, S.; Nonaka, G.-I.; Nishioka, I. Chem. Pharm. Bull. 1993, 41, 1708–1716.
[https://doi.org/10.1248/cpb.41.1708]

-
Tanaka, T.; Nonaka, G.-I.; Nishioka, I. Chem. Pharm. Bull. 1986, 34, 1039–1049.
[https://doi.org/10.1248/cpb.34.1039]

-
Yoshimura, M.; Ito, H.; Miyashita, K.; Hatano, T.; Taniguchi, S.; Amakura, Y.; Yoshida, T. Phytochemistry 2008, 69, 3062–3069.
[https://doi.org/10.1016/j.phytochem.2008.04.004]

-
Latté, K. P.; Kolodziej, H. Phytochemistry 2000, 54, 701–708.
[https://doi.org/10.1016/S0031-9422(00)00176-X]

-
Wolniak, M.; Tomczyk, M.; Gudej, J.; Wawer, I. J. Mol. Struct. 2006, 825, 26–31.
[https://doi.org/10.1016/j.molstruc.2006.04.010]

-
Yang, S.-W.; Zhou, B.-N.; Wisse, J. H.; Evans, R.; van der Werff, H.; Miller, J. S.; Kingston, D. G. I. J. Nat. Prod. 1998, 61, 901–906.
[https://doi.org/10.1021/np980046u]

-
Tanaka, T.; Jiang, Z.-H.; Kouno, I. Phytochemistry 1998, 47, 851–854.
[https://doi.org/10.1016/S0031-9422(97)00670-5]

-
Bai, N.; He, K.; Roller, M.; Zheng, B.; Chen, X.; Shao, Z.; Peng, T.; Zheng, Q. J. Agric. Food Chem. 2008, 56, 11668–11674.
[https://doi.org/10.1021/jf802152z]

- Park, J. C.; Park, J. G.; Hur, J. S.; Choi, M. R.; Yoo, E. J.; Kim, S. H.; Son, J. C.; Kim, M. S. Nat. Prod. Sci. 2004, 10, 244–247.
-
Yuan, T.; Ding, Y.; Wan, C.; Li, L.; Xu, J.; Liu, K.; Slitt, A.; Ferreira, D.; Khan, I. A.; Seeram, N. P. Org. Lett. 2012, 14, 5358–5361.
[https://doi.org/10.1021/ol302548c]

-
Nguyen, V. B.; Wang, S.-L.; Phan, T. Q.; Pham, T. H.; Huang, H.-T.; Liaw, C.-C.; Nguyen, A. D. Pharmaceuticals, 2023, 756.
[https://doi.org/10.3390/ph16050756]

-
Liu, F.; Ma, H.; Wang, G.; Liu, W.; Seeram, N. P.; Mu, Y.; Xu, Y.; Huang, X.; Li, L. Food Funct. 2018, 9, 4246–4254.
[https://doi.org/10.1039/C8FO00583D]

-
Shi, R.; Zhou, N.; Zhang, H.; Gong, M.; Han, L. Front. Nutr. 2022, 9, 1014862.
[https://doi.org/10.3389/fnut.2022.1014862]

-
Gong, Y.; Li, J.; Li, J.; Wang, L.; Fan, L. Foods 2023, 12, 715.
[https://doi.org/10.3390/foods12040715]

-
Peytam, F.; Adib, M.; Shourgeshty, R.; Firoozpour, L.; Rahmanian-Jazi, M.; Jahani, M.; Moghimi, S.; Divsalar, K.; Faramarzi, M. A.; Mojtabavi, S.; Safari, F.; Mahdavi, M.; Foroumadi, A. Sci. Rep. 2020, 10, 2595.
[https://doi.org/10.1038/s41598-020-59079-z]

-
Ding, H.; Hu, X.; Xu, X.; Zhang, G.; Gong, D. Int. J. Biol. Macromol. 2018, 107, 1844–1855.
[https://doi.org/10.1016/j.ijbiomac.2017.10.040]

-
Ur Rehman, N.; Halim, S. A.; Al-Azri, M.; Khan, M.; Khan, A.; Rafiq, K.; Al-Rawahi, A.; Csuk, R.; Al-Harrasi, A. Biomolecules 2020, 10, 751.
[https://doi.org/10.3390/biom10050751]