Secondary Metabolites of Hemerocallis fulva var. kwanso Regel and Their PTP1B Inhibitory Potential
Abstract
A phytochemical study of the root extract of Hemerocallis fulva var. kwanso Regel resulted in the isolation and structural characterization of four lignans (1‒4), three flavanones (5, 6, and 8), one chalcone (9), and two monoterpenes (7 and 10). Enzyme inhibition assays revealed that two flavanone glucosides (5 and 6) showed significant inhibitory effect against protein tyrosine phosphatase 1B (PTP1B), with IC50 values of 31.38 µM and 51.46 µM, respectively. Further enzyme kinetic studies demonstrated that these two compounds acted as noncompetitive PTP1B inhibitors, with Ki values of 30.88 µM and 50.38 µM, respectively.
Keywords:
Hemerocallis fulva var. kwanso, Asphodeaceae, phenolics, PTP1BIntroduction
The genus Hemerocallis (daylily), belonging to the family Asphodelaceae (Engler system: Liliaceae), comprises perennial herbaceous plants characterized by showy flowers that typically bloom for only one day. The genus is primarily distributed in East Asia, particularly in China, Korea, and Japan, and several species have been introduced worldwide as ornamental plants. Among these, Hemerocallis fulva var. kwanso Regel is one of the most representative and widely distributed species. It occurs naturally and as a naturalized plant throughout the Korean Peninsula and is commonly found along roadsides, in fields, and in low mountain areas. Due to its strong adaptability, it is also widely cultivated in gardens and landscapes in Korea.1
Historically, H. fulva var. kwanso has been extensively used in Asian countries as both a food source and a traditional medicinal plant for thousands of years, with its earliest recorded cultivation dating back to the Chinese Zhou Dynasty (112–255 B.C.). In traditional medicine, it has been widely employed for its antipyretic, antiemetic, anthelmintic, diuretic, and sedative properties and has been used to treat conditions such as inflammation, jaundice, insomnia, rheumatism, and fever. Ethnomedical records from China, Japan, Korea, and India document the use of various plant parts, including flowers, leaves, and roots, for the management of both internal disorders and topical inflammation.1 In recent years, advances in modern pharmacological and phytochemical research have provided scientific evidence supporting its traditional therapeutic applications. Phytochemical investigations of H. fulva var. kwanso have led to the isolation of a wide range of bioactive constituents, including flavonoids,2 saponins,3 alkaloids,4,5 anthraquinones,6 and other secondary metabolites.1 Numerous flavonoids and caffeoylquinic acid derivatives isolated from the flowers and leaves have been reported to exhibit strong antioxidant activities through free radical scavenging, inhibition of lipid peroxidation, and attenuation of oxidative stress in cellular models.2,7 In addition, rare lactam pyrrole alkaloids, such as fulvanines, isolated from the flowers of H. fulva var. kwanso, have demonstrated anti-complement and immunomodulatory activities in vitro, suggesting potential anti-inflammatory effects.4 Furthermore, phenolic-rich extracts and their constituent compounds have been shown to protect human endothelial cells against high glucose-induced oxidative and inflammatory damage, supporting the potential relevance of this species in the context of metabolic disorders.8 Collectively, these findings indicated that H. fulva var. kwanso represents a promising source of bioactive natural products. In the present study, we further explored the chemical and pharmacological diversity of this plant through the isolation and structural elucidation of its constituents and evaluation of their inhibitory activity against protein tyrosine phosphatase 1B (PTP1B).
Experimental
General experimental procedures – One- and two-dimensional NMR experiments were conducted on a Bruker Ascend 500 MHz spectrometer and a Varian Unity Inova 400 MHz spectrometer (Varian, Inc., California, USA), employing tetramethylsilane (TMS) as an internal reference, with chemical shifts expressed in δ (ppm). High-performance liquid chromatography analyses were carried out using a Waters 2487 system equipped with a UV detector (UV/VIS-156). Spectrophotometric measurements for enzyme inhibition assays were performed using an Autobio PHOmo microplate reader.
Chemicals and reagents – Silica gel (Merck, 0.040−0.063 mm), Sephadex LH-20 (Pharmacia Company), and RPC-18 (Merck, 17 mesh) were used in column chromatography (CC). Thin-layer chromatography (TLC) analyses were performed using RPC-18 F254s and silica gel F254 plates supplied by Merck. Spots were detected under ultraviolet irradiation at 254 and 360 nm and were further visualized by spraying with 10% H2SO4 in water, followed by heating. All solvents applied in the experiments were of first-grade quality and were used without additional purification. Reagents employed for the enzyme inhibitory assay were purchased from Sigma-Aldrich (St. Louis, MO, USA), including p-nitrophenyl phosphate (p-NPP), sodium hydroxide, ursolic acid, sodium citrate, and dimethyl sulfoxide (DMSO). The recombinant human PTP1B enzyme (1.8 mg/mL) was obtained from Enzo Life Sciences (Farmingdale, New York, USA), while dithiothreitol (DTT) was acquired from Bio-Rad Laboratories (Hercules, CA, USA).
Plant materials – The roots of H. fulva var. kwanso were harvested in March 2023 from the medicinal garden of the College of Pharmacy, Daegu Catholic University, Republic of Korea. The plant material was taxonomically authenticated by Prof. Byung Sun Min, and a voucher specimen (CUD-803) was deposited in the Herbarium of the College of Pharmacy, Daegu Catholic University, Republic of Korea.
Extraction and isolation – The air-dried roots of H. fulva var. kwanso (18.6 kg) were exhaustively extracted with methanol (20 L × 3 × 4 h). The combined methanol extracts were then evaporated under vacuum to obtain a crude extract (2.8 kg). The resulting residue was subsequently suspended in distilled water and partitioned sequentially with n-hexane, methylene chloride (CH2Cl2), and ethyl acetate (EtOAc), affording the n-hexane (HH, 198.6 g), CH2Cl2 (MH, 46.8 g), and EtOAc (EH, 52.6 g) fractions, in addition to the remaining aqueous layer.
The CH2Cl2 fraction (MH, 46.8 g) was subjected to silica gel column chromatography (CC) using a stepwise gradient of CH2Cl2:MeOH (100:1 to 100% MeOH), yielding multiple fractions designated MH1–MH17. Fraction MH5 (5.1 g) was further separated by silica gel CC eluted with CH2Cl2:acetone (5:1) to afford ten subfractions (MH5.1–MH5.10). Subfraction MH5.5 (965.1 mg) was purified by RP C-18 CC with MeOH:H2O (1:2), resulting in the isolation of compound 1 (5.0 mg). MH7 (3.5 g) and MH8 (4.6 g) were individually fractionated on normal phase columns using CH2Cl2:MeOH (12:1 for MH7 and 10:1 for MH8), affording subfractions MH7.1–MH7.6 and MH8.1–MH8.10, respectively. Subfraction MH7.6 (467.5 mg) was further separated by Sephadex CC with MeOH-H2O (1:1) and subsequently purified by HPLC using 50% aqueous MeOH to yield compound 2 (10.5 mg, tR = 26.8 min). In a similar manner, compound 3 (10.5 mg) was obtained from subfraction MH8.9 (421.8 mg). Fraction MH9 (3.3 g) was chromatographed over silica gel using CH2Cl2-MeOH (8:1) to give five subfractions (MH9.1–MH9.5). Subfraction MH9.1 (232.7 mg) was further purified by RP C-18 CC with MeOH:H2O (1:1), affording compound 4 (4.5 mg).
The EtOAc fraction (EH, 52.6 g) was fractionated by silica gel CC employing a stepwise gradient of CH2Cl2:MeOH (10:1 to 100% MeOH), yielding ten fractions (EH1–EH10). Among these, fraction EH9 (2.6 g) was further separated on a RP C-18 column using MeOH:H2O (1:3) to afford EH9.1–EH9.11. Subfraction EH9.6 (555.5 mg) was subsequently purified by RP C-18 CC with MeOH:H2O (1:3), resulting in the isolation of compound 5 (6.1 mg). In parallel, subfractions EH9.7 (968.9 g) and EH9.9 (5.5 g) were further resolved by RP C-18 CC eluted with MeOH:H2O (1:4), giving a series of smaller subfractions EH9.7.1–EH9.7.3 and EH9.9.1–EH9.9.10, respectively. Compound 6 (2.8 mg) was obtained from subfraction EH9.7.3 (134.1 mg), while compounds 8 (5.6 mg) and 9 (2.6 mg) were isolated from subfraction EH9.9.10 (343.0 mg) following purification on RP C-18 columns using MeOH:H2O (1:1). Subfraction EH9.7.8 (211.1 mg) was subjected to Sephadex CC with MeOH:H2O (1:1) and subsequently purified by HPLC using 35% aqueous MeOH to afford compound 7 (4.6 mg, tR = 22.0 min). Fraction EH10 (3.7 g) was further fractionated by RP C-18 CC with MeOH:H2O (1:4), yielding subfractions EH10.1–EH10.4. Subfraction EH10.2 (1.1 g) was successively purified by RP C-18 CC (MeOH:H2O, 1:4) and Sephadex CC (MeOH:H2O, 1:1), resulting in the isolation of compound 10 (4.5 mg).
(‒)-Tanegool (1) ‒ Colorless solid. 1H-NMR (400 MHz, CD3OD): δ 6.90 (1H, s, H-2), 6.85 (1H, s, H-2ʹ), 6.79 (1H, d, J = 8.2 Hz, H-6), 6.74 (1H, d, J = 8.2 Hz, H-6ʹ), 6.72 (2H, m, H-5, H-5ʹ), 4.60 (1H, d, J = 7.4 Hz, H-7), 4.46 (1H, d, J = 8.5 Hz, H-7ʹ), 4.23 (1H, d, J = 8.9, 4.3 Hz, H-9ʹa), 3.92 (1H, t, J = 8.2 Hz, H-9ʹb), 3.82 (3H, s, 3-OCH3), 3.79 (3H, s, 3ʹ-OCH3), 3.27 (1H, m, H-9a), 3.19 (1H, m, H-9b), 2.52 (1H, m, H-8ʹ), 1.86 (1H, m, H-8); 13C-NMR (100 MHz, CD3OD): δ 147.5 (C-4ʹ), 147.5 (C-4), 145.6 (C-3ʹ), 145.6 (C-3), 134.6 (C-1ʹ), 133.2 (C-1), 119.2 (C-6ʹ), 118.8 (C-6), 114.5 (C-5ʹ), 114.4 (C-5), 109.9 (C-2ʹ), 109.5 (C-2), 83.6 (C-7), 75.2 (C-7ʹ), 70.1 (C-9ʹ), 60.7 (C-9), 54.9 (3-OCH3), 54.9 (3ʹ-OCH3), 51.2 (C-8), 49.2 (C-8ʹ).9
Erythro-syringylglycerol-8-O-4′-(synapyl alcohol) ether (2) ‒ Colorless solid. 1H-NMR (500 MHz, CD3OD): δ 6.77 (2H, s, H-2ʹ, H-6ʹ), 6.70 (2H, s, H-2, H-6), 6.58 (1H, d, J = 15.9 Hz, H-7ʹ), 6.35 (1H, dd, J = 15.8, 5.6 Hz, H-8ʹ), 4.95 (1H, d, J = 5.0 Hz, H-7), 4.26 (2H, dd, J = 5.6, 1.4 Hz, H2-9ʹ), 3.93 (1H, dd, J = 12.0, 5.5 Hz, H-9b), 3.86 (6H, s, 3ʹ-OCH3, 5ʹ-OCH3), 3.85 (6H, s, 3-OCH3, 5-OCH3), 3.60 (1H, dd, J = 12.0, 3.6 Hz, H-9a); 13C-NMR (100 MHz, CD3OD): δ 153.2 (C-3ʹ, C-5ʹ), 147.6 (C-3, C-5), 135.1 (C-4ʹ), 134.4 (C-1), 133.4 (C-4), 131.6 (C-1ʹ), 129.9 (C-7ʹ), 128.5 (C-8ʹ), 103.8 (C-2ʹ, C-6ʹ), 103.6 (C-2, C-6), 86.2 (C-8), 72.8 (C-7), 62.2 (C-9ʹ), 60.2 (C-9), 55.4 (3-OCH3,5-OCH3), 55.3 (3ʹ-OCH3,5ʹ- OCH3).10
Threo-guaiacylglycerol-β-caffeic acid ether (3) ‒ Colorless solid. 1H-NMR (500 MHz, CD3OD): δ 7. 39 (1H, d, J = 15.8 Hz, H-7ʹ), 7.05 (1H, d, J = 1.7 Hz, H-2ʹ), 7.02 (1H, d, J = 8.4 Hz, H-6ʹ), 6.92 (1H, d, J = 1.9 Hz, H-2), 6.88 (1H, d, J = 8.3 Hz, H-5ʹ), 6.81 (1H, dd, J = 8.2, 1.9 Hz, H-6), 6.75 (1H, d, J = 8.2 Hz, H-5), 6.24 (1H, d, J = 15.8 Hz, H-8ʹ), 4.82 (1H, d, J = 8.2 Hz, H-7), 4.00 (1H, ddd, J = 8.0, 4.4, 2.5 Hz, H-8), 3.78 (3H, s, 3-OCH3), 3.61 (1H, dd, J = 12.4, 2.5 Hz, H-9a), 3.38 (1H, dd, J = 12.4, 4.4 Hz, H-9b); 13C-NMR (100 MHz, CD3OD): δ 166.1 (C-9ʹ), 147.8 (C-4ʹ), 147.0 (C-3), 146.6 (C-3ʹ), 145.4 (C-4), 144.2 (C-7ʹ), 128.5 (C-1), 127.9 (C-1ʹ), 121.5 (C-2ʹ), 120.3 (C-2), 117.0 (C-8ʹ), 115.8 (C-3), 114.9 (C-6ʹ), 110.7 (C-6), 78.9 (C-8), 76.3 (C-7), 60.6 (C-9), 55.1 (3-OCH3).11
(+)-Neo-olivil (4) ‒ Colorless solid. 1H-NMR (500 MHz, CD3OD): δ 7.06 (2H, d, J = 1.9 Hz, H-2, H-2ʹ), 6.90 (2H, d, J = 8.1, 1.9 Hz, H-6, H-6ʹ), 6.82 (2H, d, J = 8.1 Hz, H-5, H-5ʹ), 4.96 (2H, d, J = 8.6 Hz, H-7, H-7ʹ), 3.91 (6H, s, 3-OCH3, 3ʹ-OCH3), 3.73 (2H, dd, J = 11.3, 3.7 Hz, H-9a, H-9ʹa), 3.64 (2H, dd, J = 11.3, 5.4 Hz, H-9b, H-9ʹb), 2.35 (2H, m, H-8, H-8ʹ); 13C-NMR (100 MHz, CD3OD): δ 147.5 (C-3, C-3ʹ), 146.0 (C-4, C-4ʹ), 133.6 (C-1, C-1ʹ), 119.2 (C-6, C-6ʹ), 114.7 (C-5, C-5ʹ), 109.8 (C-2, C-2ʹ), 83.0 (C-7, C-7ʹ), 60.4 (C-9, C-9ʹ), 55.1 (3-OCH3, 3ʹ-OCH3), 54.0 (C-8, C-8ʹ).12
(2S)-Naringenin 7-O-β-D-glucopyranoside (5) ‒ White amorphous powder. 1H-NMR (400 MHz, CD3OD): δ 7.27 (2H, d, J = 7.8 Hz, H-2ʹ, H-6ʹ), 6.79 (2H, d, J = 7.8 Hz, H-3ʹ, H-5ʹ), 6.16 (2H, m, H-6, H-8), 5.31 (1H, d, J = 12.6 Hz, H-2), 4.96 (1H, d, J = 7.8, H-1ʺ), 3.85 (1H, d, J = 12.1 Hz, H-6ʺa), 3.66 (1H, d, J = 11.8 Hz, H-6ʺb), 3.43–3.37 (4H, m, H-2ʺ–5ʺ), 3.11 (1H, dd, J = 16.4, 14.0 Hz, Heq-3), 2.69 (1H, d, J = 17.2 Hz, Hax-3); 13C-NMR (100 MHz, CD3OD): δ 198.5 (C-4), 166.8 (C-7), 164.8 (C-5), 164.5 (C-9), 159.0 (C-4ʹ), 130.8 (C-1ʹ), 129.1 (C-2ʹ, C-6ʹ), 116.3 (C-3ʹ, C-5ʹ), 104.8 (C-10), 101.1 (C-1ʺ), 98.0 (C-6), 96.8 (C-8), 80.6 (C-2), 78.1 (C-3ʺ), 77.7 (C-5ʺ), 74.6 (C-2ʺ), 71.0 (C-4ʺ), 62.2 (C-6ʺ), 43.8 (C-3).13
(2S)-Naringenin-5-O-β-D-glucopyranoside (6) ‒ White amorphous powder. 1H-NMR (400 MHz, CD3OD): δ 7.28 (2H, d, J = 8.0 Hz, H-2ʹ, H-6ʹ), 6.79 (2H, d, J = 8.0 Hz, H-3ʹ, H-5ʹ), 6.46 (1H, s, H-8), 6.11 (1H, s, H-6), 5.29 (1H, d, J = 13.1 Hz, H-2), 4.74 (1H, d, J = 7.8, H-1ʺ), 3.94–3.43 (6H, m, H-2ʺ–6ʺ), 2.96 (1H, dd, J = 17.2, 13.2 Hz, Heq-3), 2.68 (1H, d, J = 16.9 Hz, Hax-3); 13C-NMR (100 MHz, CD3OD): δ 191.6 (C-4), 165.7 (C-7), 165.1 (C-9), 160.9 (C-5), 157.5 (C-4ʹ), 129.6 (C-1ʹ), 127.6 (C-2ʹ, C-6ʹ), 114.9 (C-3ʹ, C-5ʹ), 105.6 (C-10), 103.6 (C-1ʺ), 99.0 (C-6), 98.0 (C-8), 78.8 (C-2), 77.2 (C-3ʺ), 75.7 (C-5ʺ), 73.2 (C-2ʺ), 69.8 (C-4ʺ), 61.1 (C-6ʺ), 44.9 (C-3).14
(–)-Angelicoidenol-2-O-β-D-glucopyranoside (7) ‒ Colorless oil. 1H-NMR (400 MHz, CD3OD): δ 4.20 (1H, d, J = 7.8 Hz, H-1ʹ), 4.03 (1H, d, J = 8.5 Hz, H-2β), 3.83 (2H, m, H-5α, H-6ʹ), 3.65 (1H, dd, J = 11.8, 5.4, H-6ʹ), 3.32 (1H, m, H-3ʹ), 3.26 (1H, m, H-4ʹ), 3.19 (1H, m, H-5ʹ), 3.13 (1H, t, J = 8.1 Hz, H-2ʹ), 2.47 (1H, dd, J = 13.2, 8.0 Hz, H-6α), 2.16 (1H, m, H-3β), 1.68 (1H, d, J = 5.0 Hz, H-4α), 1.29 (1H, d, J = 12.9 Hz, H-6β), 1.06 (3H, s, H3-8), 1.00 (1H, dd, J = 13.4, 2.5 Hz, H-3α), 0.90 (3H, s, H3-10), 0.83 (3H, s, H3-9); 13C-NMR (100 MHz, CD3OD): δ 101.4 (C-1ʹ), 81.5 (C-2), 76.7 (C-3ʹ), 76.4 (C-5ʹ), 74.4 (C-5), 73.6 (C-2ʹ), 70.2 (C-4ʹ), 61.3 (C-6ʹ), 52.1 (C-4), 49.6 (C-7), 47.3 (C-1), 38.3 (C-6), 32.8 (C-3), 19.9 (C-8), 18.9 (C-9), 11.9 (C-10).15
(2S)-5,7-Di-O-β-D-glucopyranosyl-naringenin (8) – White amorphous powder. 1H-NMR (400 MHz, pyridine-d5): δ 7.46 (2H, d, J = 8.3 Hz, H-2ʹ, H-6ʹ), 7.19 (2H, ovl, H-3ʹ, H-5ʹ), 7.30 (1H, s, H-8), 6.64 (1H, s, H-6), 5.74 (1H, d, J = 7.6 Hz, H-1ʺ), 5.39 (1H, d, J = 7.0, H-1‴), 5.31 (1H, dd, J = 13.1, 2.4 Hz, H-2), 4.58 (2H, m, H-6ʺa, H-6‴a), 4.37–4.22 (4H, m, H-2ʺ–5ʺ, H-6ʺb, H-2‴–5‴, H-6‴b), 3.08 (1H, dd, J = 17.2, 13.4 Hz, Heq-3), 2.82 (1H, dd, J = 17.0, 2.7 Hz, Hax-3); 13C-NMR (100 MHz, pyridine-d5): δ 191.1 (C-4), 165.1 (C-7), 165.0 (C-5), 161.9 (C-9), 159.8 (C-4ʹ), 130.0 (C-1ʹ), 129.1 (C-2ʹ, C-6ʹ), 116.7 (C-3ʹ, C-5ʹ), 108.4 (C-10), 105.3 (C-1ʺ), 101.4 (C1‴), 99.9 (C-6), 99.9 (C-8), 79.8 (C-2), 79.6 (C-3ʺ), 79.3 (C-3‴), 78.7 (C-5ʺ), 77.9 (C-5‴), 75.5 (C-2ʺ), 75.0 (C-2‴), 71.8 (C-4ʺ), 71.6 (C-4‴), 63.0 (C-6ʺ), 62.8 (C-6‴), 46.2 (C-3).13
Chalcononaringenin 2ʹ,4ʹ-di-O-β-D-glucoside (9) – Colorless solid. 1H-NMR (400 MHz, DMSO-d6): δ 8.01 (1H, d, J = 15.4 Hz, H-7ʹ), 7.75 (1H, d, J = 15.4 Hz, H8ʹ), 7.66 (2H, d, J = 8.5 Hz, H-2ʹ, H-6ʹ), 6.89 (2H, d, J = 8.5 Hz, H-3ʹ, H-5ʹ), 6.56 (1H, s, H-5), 6.33 (1H, s, H-3), 5.25 (1H, d, J = 7.3 Hz, H-1ʺ), 5.08 (1H, d, J = 7.1, H-1‴), 3.99 (1H, d, J = 11.8 Hz, H-6ʺa, H-6‴a), 3.77–3.38 (4H, m, H-2ʺ–5ʺ, H-6ʺb, H-2‴–5‴, H-6‴b); 13C-NMR (100 MHz, CD3OD): δ 195.0 (C-9ʹ), 166.5 (C-6), 164.4 (C-2), 161.3 (C-4ʹ), 161.1 (C-4), 145.1 (C-7ʹ), 131.9 (C-2ʹ, C-6ʹ), 128.3 (C-1ʹ), 125.6 (C-8ʹ), 116.9 (C-3ʹ, C-5ʹ), 109.3 (C-1), 101.7 (C-1ʺ), 101.2 (C-1‴), 99.7 (C-3), 95.9 (C-5), 78.5 (C-3ʺ, C-3‴), 78.4 (C-5ʺ), 77.9 (C-5‴), 75.0 (C-2ʺ), 74.7 (C-2‴), 71.6 (C-4ʺ), 71.5 (C-4‴), 62.8 (C-6ʺ), 62.7 (C-6‴).16
Hemerocalloside (10) – Colorless solid. 1H-NMR (500 MHz, CD3OD): δ 5.33 (1H, s, H-2), 4.99 (1H, d, J = 2.1 Hz, H-1ʺ), 4.44 (1H, d, J = 7.7 Hz, H-1ʹ), 3.97 (1H, d, J = 7.5 Hz, H-4ʺ), 3.95 (1H, dd, J = 10.5, 2.0 Hz, H-6ʹb), 3.90 (1H, d, J = 2.1 Hz, H-2ʺ), 3.78 (1H, d, J = 9.6 Hz, H-4ʺ), 3.61 (2H, s, H-3‴), 3.55 (1H, dd, J = 11.0, 6.4 Hz, H-6ʹa), 3.32 (1H, t, J = 3.2 Hz, H-3ʹ), 3.25 (1H, dd, J = 9.6, 8.9 Hz, H-4ʹ), 3.17 (1H, dd, J = 9.1, 7.7 Hz, H-5ʹ), 2.37 (1H, d, J = 10.0 Hz, H-6), 2.26 (2H, q, J = 18.3 Hz, H-3), 1.96 (1H, dt, J = 13.7, 6.8 Hz, H-10), 1.89 (1H, m, H-5a), 1.67 (3H, s, H-5b), 0.99 (3H, d, J = 5.2 Hz, H3-8), 0.97 (3H, d, J = 5.2 Hz, H3-9); 13C-NMR (100 MHz, CD3OD): δ 136.1 (C-1), 120.3 (C-2), 111.5 (C-1ʺ), 99.4 (C-1ʹ), 81.4 (C-4), 81.1 (C-3ʺ), 78.9 (C-2ʺ), 78.7 (C-3ʹ), 76.9 (C-2ʹ), 75.6 (C4ʺ), 72.6 (C-4ʹ), 69.5 (C-6ʹ), 66.5 (C-3‴), 35.9 (C-10), 32.3 (C-3), 30.8 (C-5), 29.2 (C-6), 23.9 (C-7), 18.4 (C-8), 18.2 (C-9).17
PTP1B inhibitory assay – The PTP1B inhibitory assay was performed according to a previously reported method with slight modifications.5,18 The reaction mixture was prepared using a buffer solution (50 mM citrate buffer, pH 6.0) containing 0.1 M NaCl, 1 mM EDTA, and 1 mM DTT. PTP1B enzyme (0.5 units), the substrate p-nitrophenyl phosphate (p-NPP, 2.0 mM), and the test compounds (1–100 µM) were diluted with this reaction buffer prior to the assay. The experiment was carried out in a 96-well microplate by sequentially adding 10 µL of sample solution, 30–40 µL of reaction buffer, 10 µL of enzyme solution, and 50 µL of p-NPP to each well. The plate was gently shaken and incubated at 37°C for 20 min. The reaction was terminated by adding 10 µL of 10 M NaOH. Under alkaline conditions, dephosphorylation of p-NPP produces the yellow p-nitrophenolate ion, allowing the enzymatic activity of PTP1B to be measured spectrophotometrically at 405 nm. Ursolic acid served as the positive control.
PTP1B enzyme kinetics – To investigate the inhibition mechanism of the most active compounds (5 and 6), enzyme kinetic analyses were performed using Lineweaver–Burk and Dixon plot approaches.18 The inhibitory activity of PTP1B was evaluated at different concentrations of the substrate p-nitrophenyl phosphate (p-NPP; 1, 2, and 4 mM) in the presence of varying concentrations of compound 5 (10.2, 25.6, and 55.1 µM) and compound 6 (21.5, 45.5, and 80.5 µM). The enzymatic assays for PTP1B were carried out following the procedure described above. Kinetic data were processed using SigmaPlot 15.0 (SPCC Inc., Chicago, IL, USA), and the inhibition constants (Ki) were determined from Dixon plot analysis.
Results and Discussions
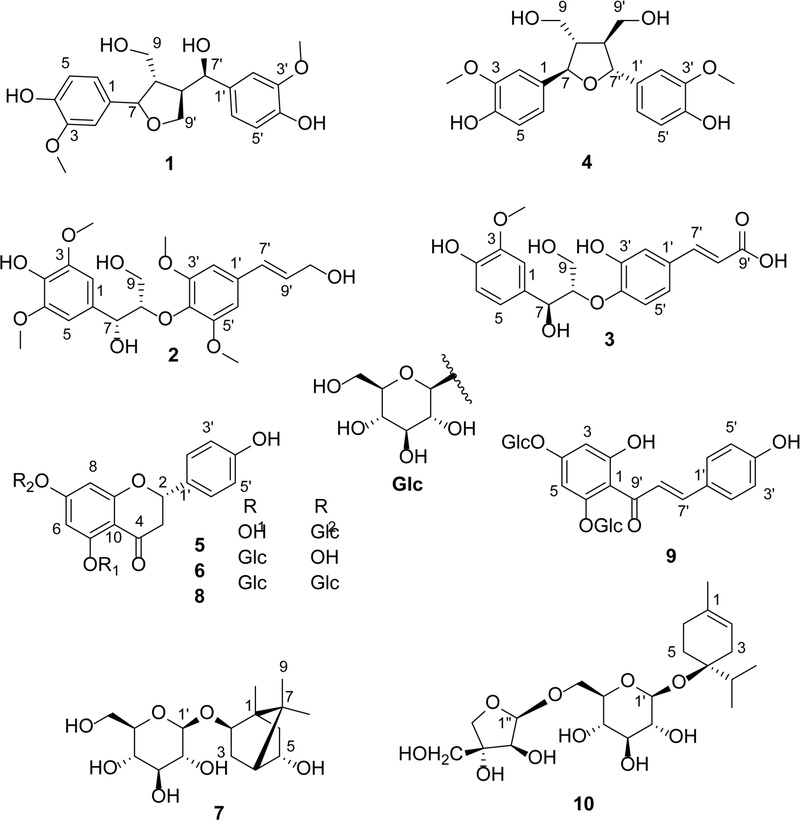
Ten compounds including (‒)-tanegool (1),9erythro-syringylglycerol-8-O-4′-(synapyl alcohol) ether (2),10threo-guaiacylglycerol-β-caffeic acid ether (3),11 (+)-neo-olivil (4),12 naringenin 7-O-β-D-glucopyranoside (5),13 naringenin-5-O-β-D-glucopyranoside (6),14 (–)-angelicoidenol-2-O-β-D-glucopyranoside (7),15 5,7-di-O-β-D-glucopyranosyl-naringenin (8),13 chalcononaringenin 2ʹ,4ʹ-di-O-β-D-glucoside (9),16 and hemerocalloside (10)17 were isolated from H. fulva root extract (Fig. 1). The structures of these compounds were determined by comparing their spectroscopic characteristics with previously published data. To the best of our knowledge, compounds 1, 3–7, and 9 are reported here for the first time from the genus Hemerocallis. In addition, compounds 1 and 3–9 are reported here for the first time from H. fulva var. kwanso.
Interpretation of the NMR spectroscopic data allowed compound 5 to be assigned as naringenin glucoside.13 The presence of a carbonyl signal at δC 198.5, a characteristic C-2 methine at δH 5.31 (1H, d, J = 12.6 Hz)/δC 80.6, and a C-3 methylene at δH 3.11(1H, m, Ha-3) and 2.69 (1H, d, J = 17.2 Hz)/δC 43.8 confirmed the flavanone skeleton.13 The A-ring exhibited two meta-coupled protons overlapped at δH 6.16, consistent with a 5,7-dioxylation benzene ring, while the B-ring showed an AA′BB′ pattern indicative of a para-substituted phenyl group at δH/δC 7.27 (2H, d, J = 7.8 Hz, H-2ʹ, H-6ʹ)/129.1 and 6.79 (2H, d, J = 7.8 Hz, H-3ʹ, H-5ʹ)/116.3. In addition, typical signals for a β-D-glucose moiety were observed at δH 4.96 (1H, d, J = 7.8, H-1ʺ), 3.85 (1H, d, J = 12.1 Hz, H-6ʺa), 3.66 (1H, d, J = 11.8 Hz, H-6ʺb), 3.43–3.37 (4H, m, H-2ʺ–5ʺ) and δC 101.1 (C-1ʺ), 78.1 (C-3ʺ), 77.7 (C-5ʺ), 74.6 (C-2ʺ), 71.0 (C-4ʺ), 62.2 (C-6ʺ). Key HBMC correlations from anomeric proton H-1ʹ to C-7 confirmed the glucose moiety linked with C-7. The NOE correlation between H-3eq (δH 3.11) and the aromatic protons H-2′/H-6′ (δH 7.27) indicated an α-orientation of the B ring, allowing assignment of the (2S)-isoform of 5.19 In conclusion, compound 5 was identified as (2S)-naringenin 7-O-β-D-glucopyranoside (prunin) based on comprehensive spectroscopic analyses, comparison with previously published data,13 and consideration of the natural biosynthetic pathway of flavanones.19 Similarly, compounds 6 and 8 were determined as (2S)-naringenin-5-O-β-D-glucopyranoside (6)14 and (2S)-5,7-di-O-β-D-glucopyranosyl-naringenin (8),13 respectively.
Analysis of the NMR spectroscopic data for compound 7 revealed characteristic features of a bornan-2,5-diol monoterpene glycoside.15 The aglycone moiety was first recognized from the 1D NMR data by the presence of sp3 protons corresponding to two methylene groups at δH/δC 2.47 (1H, dd, J = 13.2, 8.0 Hz, H-6a) and 1.29 (1H, d, J = 12.9 Hz, H-6b)/38.3 (C-6) and 2.16 (1H, m, H-3a) and 1.00 (1H, dd, J = 13.4, 2.5 Hz, H-3b)/32.8 (C-3), one methine at δH/δC 1.68 (1H, d, J = 5.0 Hz, H-4)/52.1 (C-4), together with three singlet methyls at δH/δC 1. 06 (3H, s, H3-8)/19.9 (C-8), 0.90 (3H, s, H3-10)/11.9 (C-10), and 0.83 (3H, s, H3-9)/18.9 (C-9), which are diagnostic features of the bornane skeleton. Notably, two oxygenated protons were observed in the downfield region at δH/δC 4.03 (1H, d, J = 8.5 Hz, H-2)/81.5 (C-2) and 3.83 (1H, m, H-5)/74.4 (C-5), indicating the presence of hydroxyl and/or glycosylated functionalities on the aglycone. The remaining signals were attributed to one β-D-glucopyranosyl, based on the anomeric proton signal at δH/δC 4.20 (d, J = 7.8 Hz, H-1ʹ)/101.4 (C-1ʹ), while the other sugar resonances appeared in the ranges δH 3.83–3.13 and δC 76.7–61.3. Importantly, C-2 (δC 81.5) was significantly deshielded relative to C-5 (δC 74.4), indicating that C-2 was the site of glycosidic linkage. The glucosylation shift observed for C-2 (δC 81.5) relative to the corresponding carbon in the aglycone (Δδ ≈ 6–7 ppm), together with the upfield shift of the adjacent C-3 (δC 32.8), is characteristic of an R-configured secondary alcohol, indicating a 2R,5S configuration for the aglycone.15,20 On the basis of the above spectroscopic evidence and by comparison with reported data in the literature,15 compound 7 was identified as (–)-angelicoidenol-2-O-β-D-glucopyranoside.
The 1D NMR spectra of compound 9 displayed signals closely resembling those of compound 8. The main difference arose from the replacement of the sp3 protons in C ring by a trans-double bond at δH/δC 8.01 (1H, d, J = 15.4 Hz, H-7ʹ)/145.1 (C-7ʹ) and 7.75 (1H, d, J = 15.4 Hz, H-8ʹ)/125.6 (C-8ʹ), suggested that compound 9 was a naringenin-derived chalcone glucoside. Based on analysis of its NMR data and comparison with published literature values,16 9 was determined to be chalcononaringenin 2ʹ,4ʹ-di-O-β-D-glucopyranoside.
Protein tyrosine phosphatase 1B (PTP1B) is a cytosolic enzyme involved in the negative regulation of insulin and leptin signaling pathways. It acts by dephosphorylating the insulin receptor and related downstream signaling molecules, which leads to reduced insulin sensitivity. Overexpression or hyperactivity of PTP1B has been strongly linked to insulin resistance, a key feature of type 2 diabetes. In addition, PTP1B plays a critical role in leptin resistance, which contributes to increased food intake and the development of obesity. Consequently, inhibition of PTP1B has been widely explored as a potential strategy for the management of type 2 diabetes and obesity.21
Protein tyrosine phosphatase 1B (PTP1B) is a cytosolic enzyme involved in the negative regulation of insulin and leptin signaling pathways. It acts by dephosphorylating the insulin receptor and related downstream signaling molecules, which leads to reduced insulin sensitivity. Elevated expression or increased activity of PTP1B has been strongly linked to insulin resistance, a hallmark of type 2 diabetes. Moreover, PTP1B is also implicated in leptin resistance, a condition that promotes excessive food intake and contributes to the development of obesity. For these reasons, PTP1B has attracted considerable attention as a potential therapeutic target for managing both type 2 diabetes and obesity.
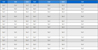
The inhibitory activities of the isolated compounds (1‒10) against PTP1B were evaluated using a colorimetric assay. Among them, compound 5 exhibited the strongest inhibitory effect, with an IC50 value of 31.38 µM, whereas its isomer, compound 6, showed weaker activity, with an IC50 value of 51.46 µM (Table 1). Compound 8, a naringenin derivative bearing two glucose moieties, displayed no inhibitory activity at 100 μM. These results suggest that simultaneous glycosylation at both C-5 and C-7 positions negatively affects the PTP1B inhibitory activity of naringenin, and that glycosylation at C-7 is more favorable than at C-5. The others (1–4 and 7–10) were inactive at 100 μM.
Compounds 5 and 6 were further evaluated to enzyme kinetic analysis using Lineweaver–Burk and Dixon plots. In the Lineweaver–Burk plots, the regression lines for both compounds intersected at the x-axis, indicating a non-competitive mode of inhibition toward PTP1B. The inhibition constants (Ki) were determined from Dixon plots, giving Ki values of 30.88 and 50.38 µM for compounds 5 and 6, respectively (Fig. 2).
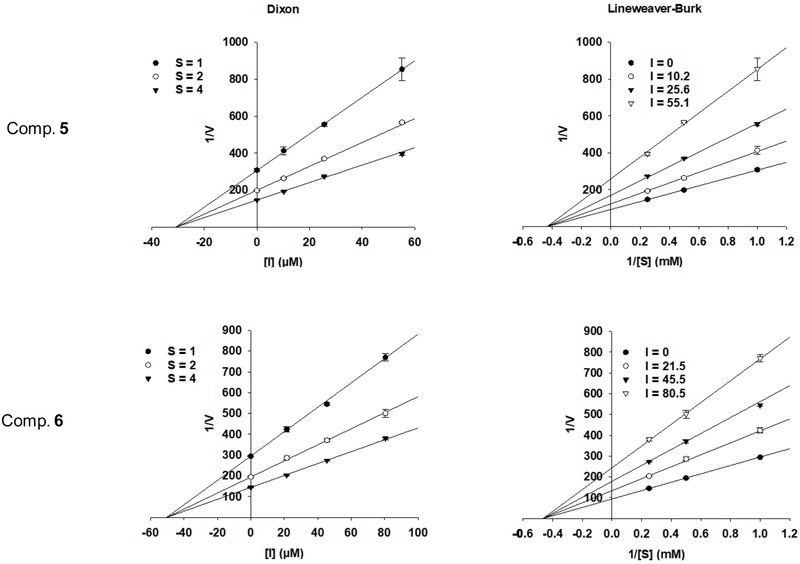
Lineweaver-Burk plot and Dixon analyses of the inhibitory activity of compounds 5 and 6. [S] (mM) and [I] (μM) correspond to the concentrations of p-NPP and inhibitor, respectively. Plots were prepared using SigmaPlot 15.0.
Flavonoids represent a large class of plant-derived natural products that have attracted considerable attention due to their pronounced inhibitory activity against PTP1B, a well-established therapeutic target for type 2 diabetes and other metabolic disorders. In the present study, two glucosylated naringenin derivatives (5 and 6) exhibited significant inhibitory activity against PTP1B. Notably, this was the first report describing the PTP1B inhibitory activity of naringenin 5-O-β-D-glucopyranoside (6), whereas the activity of naringenin 7-O-β-D-glucopyrano-side (5)22 has been reported previously. Overall, these findings suggest that mono-glucosylated naringenin derivatives represent promising natural PTP1B inhibitors and provide further support for their potential development as lead compounds for antidiabetic drug discovery.
In conclusion, comprehensive phytochemical analysis of H. fulva var. kwanso roots led to the isolation and structural characterization of ten secondary metabolites, including four lignans (1‒4), three flavanones (5, 6, and 8), one chalcone (9), and two monoterpenes (7 and 10). Compounds 3–9 are reported here for the first time from this species, and compounds 3–7 and 9 are newly identified from the genus Hemerocallis, thereby expanding the known chemical diversity for this medicinal plant. Among the isolated constituents, two flavanone glucosides (5 and 6) demonstrated notable PTP1B inhibitory activity through a non-competitive mechanism. These findings highlight H. fulva var. kwanso as a valuable natural source of bioactive flavonoids and provide a scientific foundation for its potential application in the discovery and development of therapeutic agents targeting metabolic disorders.
Acknowledgments
This research was supported by the Korea Ministry of Food and Drug Safety (23212KFDA204) and Gyeongsangbuk-do RISE (Regional Innovation System & Education; 2025-RISE-15-107). We gratefully acknowledge Korea Basic Science Institute (KBSI) for performing the mass spectrometry measurements.
Conflicts of Interest
The authors have declared that there are no conflicts of interest.
References
-
Li, X.; Jiang, S.; Cui, J.; Qin, X.; Zhang, G. J. Hortic. Sci. Biotechnol. 2022, 97, 298–314.
[https://doi.org/10.1080/14620316.2021.1988728]

-
Taguchi, K.; Yamasaki, K.; Maesaki, H.; Tokuno, M.; Okazaki, S.; Moriuchi, H.; Takeshita, K.; Otagiri, M.; Seo, H. Nat. Prod. Res. 2014, 28, 2211–2213.
[https://doi.org/10.1080/14786419.2014.919285]

-
Konishi, T.; Fujiwara, Y.; Konoshima, T.; Kiyosawa, S.; Nishi, M.; Miyahara, K. Chem. Pharm. Bull. 2001, 49, 318–320.
[https://doi.org/10.1248/cpb.49.318]

-
Lei, Y.-E.; Wang, Q.; Yaermaimaiti, S.; Ma, Z.-H.; Li, M.-M.; Lu, Y.; Chen, D.-F.; Wang, Q. Chem. Biodiver. 2024, 21, e202301672.
[https://doi.org/10.1002/cbdv.202301672]

-
Le, T. T.; Ha, M. T.; Tran, T. T.; Min, S. E.; Han, K.-H.; Kim, J. A.; Min, B. S. Phytochemistry 2026, 246, 114800.
[https://doi.org/10.1016/j.phytochem.2026.114800]

-
Cichewicz, R. H.; Lim, K.-C.; McKerrow, J. H.; Nair, M. G. Tetrahedron 2002, 58, 8597–8606.
[https://doi.org/10.1016/S0040-4020(02)00802-5]

-
Lin, Y.-L.; Lu, C.-K.; Huang, Y.-J.; Chen, H.-J. J. Agric. Food Chem. 2011, 59, 8789–8795.
[https://doi.org/10.1021/jf201166b]

-
Wu, W.-T.; Mong, M.-C.; Yang, Y.-C.; Wang, Z.-H.; Yin, M.-C. J. Food Sci. 2018, 83, 1463–1469.
[https://doi.org/10.1111/1750-3841.14137]

-
Cutillo, F.; D'Abrosca, B.; DellaGreca, M.; Fiorentino, A.; Zarrelli, A. J. Agric. Food Chem. 2003, 51, 6165–6172.
[https://doi.org/10.1021/jf034644c]

- Trang, B. T. T.; Anh, B. T. M.; Mai, N. T. JMEST 2023, 10, 15823–15827.
-
Ichikawa, M.; Ryu, K.; Yoshida, J.; Ide, N.; Kodera, Y.; Sasaoka, T.; Rosen, R. T. J. Agric. Food Chem. 2003, 51, 7313–7317.
[https://doi.org/10.1021/jf034791a]

-
Schöttner, M.; Reiner, J.; Tayman, F. S. K. Phytochemistry 1997, 46, 1107–1109.
[https://doi.org/10.1016/S0031-9422(97)00401-9]

-
Shimoda, K.; Kubota, N.; Taniuchi, K.; Sato, D.; Nakajima, N.; Hamada, H.; Hamada, H. Phytochemistry 2010, 71, 201–205.
[https://doi.org/10.1016/j.phytochem.2009.09.035]

-
Ibrahim, L. F.; El-Senousy, W. M.; Hawas, U. W. Chem. Nat. Compd. 2007, 43, 659–662.
[https://doi.org/10.1007/s10600-007-0222-y]

-
Inoshiri, S.; Saiki, M.; Kohda, H.; Otsuka, H.; Yamasaki, K. Phytochemistry 1988, 27, 2869–2871.
[https://doi.org/10.1016/0031-9422(88)80678-2]

-
Iwashina, T.; Kitajima, J.; Shiuchi, T.; Itou, Y. Biochem. Syst. Ecol. 2005, 33, 571–584.
[https://doi.org/10.1016/j.bse.2004.12.005]

-
Yang, Z.-D.; Chen, H.; Li, Y.-C. Helv. Chim. Acta 2003, 86, 3305–3309.
[https://doi.org/10.1002/hlca.200390271]

-
Le, T. T.; Ha, M. T.; Lee, G. S.; Nguyen, V. P.; Kim, C. S.; Kim, J. A.; Min, B. S. Phytochemistry 2025, 229, 114269.
[https://doi.org/10.1016/j.phytochem.2024.114269]

-
Maltese, F.; Erkelens, C.; van der Kooy, F.; Choi, Y. H.; Verpoorte, R. Food Chem. 2009, 116, 575–579.
[https://doi.org/10.1016/j.foodchem.2009.03.023]

-
Jiang, Z.-Y.; Liu, W.-F.; Zhang, X.-M.; Luo, J.; Ma, Y.-B.; Chen, J.-J. Bioorg. Med. Chem. Lett. 2013, 23, 2123–2127.
[https://doi.org/10.1016/j.bmcl.2013.01.118]

-
Johnson, T. O.; Ermolieff, J.; Jirousek, M. R. Nat. Rev. Drug Discov. 2002, 1, 696–709.
[https://doi.org/10.1038/nrd895]

-
Jung, H. A.; Paudel, P.; Seong, S. H.; Min, B.-S.; Choi, J. S. Bioorg. Med. Chem. Lett. 2017, 27, 2274–2280.
[https://doi.org/10.1016/j.bmcl.2017.04.054]