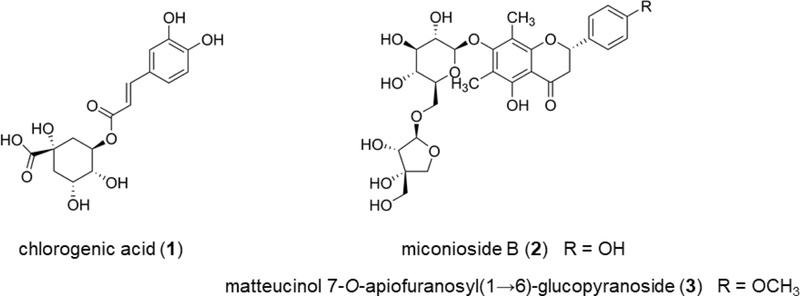
Development and Validation of an Analytical Method for the Simultaneous Determination of Three Marker Compounds in Wikstroemia trichotoma
Abstract
The Wikstroemia genus is highly regarded in traditional Asian medicine for its diverse therapeutic applications, including the treatment of inflammatory and infectious conditions. Among its members, Wikstroemia trichotoma (Thunb.) Makino remains a promising medicinal resource which is yet to be chemically characterized. To ensure the chemical consistency of W. trichotoma, we developed and validated the first HPLC method for the simultaneous quantification of three major marker compounds: chlorogenic acid (1), miconioside B (2), and matteucinol 7-O-apiofuranosyl(1→6)-glucopyranoside (3). Chromatographic separation was achieved on a C18 column using a gradient elution system of 0.1% formic acid in water and 0.1% formic acid in acetonitrile. Detection was optimized using a photodiode array (PDA) detector at 280 and 325 nm, based on the absorption maxima of the markers. The method was validated in accordance with the International Council for Harmonisation (ICH) and the Ministry of Food and Drug Safety (MFDS) guidelines. The results demonstrated high linearity (r2 > 0.999), with limits of detection and quantitation ranging from 4.28–6.42 and 12.97–19.47 μg/mL, respectively. Intra- and inter-day precision (% RSD ≤ 1.83%) and accuracy (recoveries of 92.5–101.7%) were within acceptable limits. Quantitative analysis revealed the contents of 1, 2, and 3 in the W. trichotoma extract to be 19.9, 139.8, and 264.9 mg/g, respectively. This study provides a reliable analytical framework for the standardization, quality control, and future pharmacological evaluation of W. trichotoma.
Keywords:
Wikstroemia trichotoma (Thunb.) Makino, HPLC-PDA, Simultaneous determination, Method validation, Quality control, Phenolic compoundsIntroduction
Plants possess a remarkable ability to biosynthesize an extensive variety of structurally diverse bioactive compounds, including alkaloids, phenolics, terpenoids, and glycosides.1–3 This chemical diversity arises from complex secondary metabolic pathways, which have evolved to help plants adapt to environmental stresses such as pathogens, UV radiation, and herbivores.4,5 Many compounds derived from natural sources exhibit significant pharmacological properties, including anti-inflammatory, anti-oxidant, anti-microbial, and anticancer activities.6–8 The genus Wikstroemia (Thymelaeaceae) has been used in Asian traditional medicine for managing conditions such as pneumonia, parotitis, and pertussis, because of its anti-inflammatory, detoxifying, and antimicrobial effects.9 While several Wikstroemia species have been studied phytochemically and pharmacologically, Wikstroemia trichotoma (Thunb.) Makino remains underexplored, despite its taxonomic proximity to bioactive species.10–12 Recent studies have identified phenolics and flavonoids in W. trichotoma, which suggests that this species has pharmacological potential.13 However, challenges in reproducibility and analytical standardization often limit the development of plant-derived products.14
Herbal extracts are complex multi-component systems, often with pharmacological effects resulting from synergistic effects of major bioactive compounds. Thus, quality control based on a single marker cannot ensure batch-to-batch consistency or therapeutic efficacy.15 Therefore, establishing a method for the simultaneous determination of multiple key constituents is essential for reliable quality control of W. trichotoma extract. However, no simultaneous analytical method has yet been reported. Given the need for comprehensive quality assurance, establishing a validated analytical method for the simultaneous quantification of three marker compounds in W. trichotoma extract using HPLC-PDA offers significant advantages, due to the broad applicability, reliability, and regulatory acceptance of the technique for quality control and standardization, which is required to ensure consistent quality, safety, and efficacy across production batches.16
To provide a comprehensive quality profile, three marker compounds were selected to represent different chemical classes and abundance levels in the extract. Chlorogenic acid (1) was chosen as a general quality marker because its potent pharmacological properties are structurally linked to its o-diphenolic backbone, making it a representative of the phenolic acid class in this extract.17,18 Furthermore, miconioside B (2) and matteucinol 7-O-apiofuranosyl(1→6)-glucopyranoside (3) were included as the principal quantitative markers, as they represent over 50% of the total peak area in the preliminary chromatographic profile.13 Collectively, their selection encompasses the two major phytochemical classes, specifically, phenolic acids (1) and flavonoid glycosides (2 and 3), which characterize the chemical profile of this extract. Therefore, the primary objective of this investigation was to develop and validate an HPLC-PDA method that simultaneously and accurately quantifies these three marker compounds in W. trichotoma extract. The method was developed in accordance with ICH Q2(R2) guidelines and Korean MFDS guidelines, with an emphasis on quality assurance and industrial applicability.19,20
Experimental
Plant material – Wikstroemia trichotoma (Thunb.) Makino used in this study was obtained from Sineosan (Gimhae, Republic of Korea) in September 2020. The plant specimens were dried and stored in a storage cabinet (GT SCIEN, Daejeon, Republic of Korea) maintained at 25°C. Its taxonomic identity was confirmed by Dr. Jin-Hyub Paik at the International Biological Material Research Center (IBMRC) in the Korea Research Institute of Bioscience and Biotechnology (KRIBB). A voucher specimen (PNU-0041) was archived at the College of Pharmacy, Pusan National University.
General experimental procedures – Nuclear magnetic resonance (NMR) analysis was performed using a JEOL JNM-ECZ400S 400 MHz NMR spectrometer (JEOL, Tokyo, Japan). High-resolution electrospray ionization mass spectrometry (HR-ESI-MS) was conducted using an Agilent 6530 Accurate-Mass Q/TOF-LC/MS system (Agilent Technologies, Santa Clara, CA, USA). The high-performance liquid chromatography-photodiode array (HPLC-PDA) analysis was performed using a Waters HPLC system with an e2695 separation module, a 2998 PDA detector, and Empower® 3 Software (Waters Corp., Milford, MA, USA). Separation was carried out on a reversed-phase column (Aegispak C18-L, 5 µm, 4.6 × 250 mm, Young Jin Biochrom., Seongnam, Republic of Korea). Semi-preparative HPLC was performed using a Gilson HPLC system (Gilson®, Middleton, WI, USA) equipped with two pumps (305 pump, 307 pump) and a mixer (811C dynamic mixer), in series with Shimadzu HPLC system (Shimadzu Co., Ltd., Kyoto, Japan) equipped with a pump (LC-20AT), and a UV/vis detector (SPD-20A). A Watchers reversed-phase column (120 ODS-BP, S-10 µm, 150 × 10 mm, Isu Industry Corp., Seoul, Republic of Korea) was used. Open-column chromatography was performed using Silica gel 60 (particle size 63-200 µm, Merck, Darmstadt, Germany) and SephadexTM LH-20 (bead size 25-100 µm; GE Healthcare, Uppsala, Sweden) as stationary phases. Thin-layer chromatography (TLC) was performed using silica gel 60 F254 plates (Merck, Darmstadt, Germany). Spots were visualized under UV light after treatment with anisaldehyde-sulfuric acid reagent, and the technique was used to monitor the separation progress during column chromatography.
Extraction and isolation – For the subsequent quantitative validation study, dry powdered W. trichotoma (1.004 g), obtained from the aerial parts of the plant, was extracted using 95% ethanol (EtOH) by ultrasonic-assisted extraction for 90 minutes and then allowed to stand at room temperature (RT) for 12 hours. This procedure was performed twice under the same experimental conditions. Filtration of the resulting crude extract through filter paper was followed by pooling, concentration in vacuo at 40°C, and freeze-drying. W. trichotoma EtOH extract (131.9 mg) obtained was stored at 4°C.
For the isolation of marker compounds, shade-dried, powdered aerial parts (3.4 kg) of W. trichotoma were extracted with 34 L of methanol (MeOH) via ultra-sonication for 90 minutes, and allowed to stand for 12 hours at RT. The solvent was then removed by filtering through a double-layered gauze and evaporation under low pressure. The extraction procedure was repeated twice on the residue. The crude MeOH extract (629.3 g) was suspended in 4 L of distilled water and sequentially partitioned with 8 L of each solvent (hexane, chloroform, ethyl acetate (EtOAc), and n-butanol (n-BuOH)), yielding four fractions. All organic fractions were concentrated to dryness under reduced pressure, and the resulting four fractions weighed 96.2 g, 41.5 g, 216.5 g, and 197.9 g, respectively. Polarity-based separation was performed using chromatographic techniques. The three marker compounds (1–3) were isolated from the EtOAc and n-BuOH fractions and identified spectroscopically.
The EtOAc fraction (186.8 g) of the MeOH extract was dissolved in a minimum volume of MeOH and then allowed to stand at 4°C. The resulting crystals were collected by vacuum filtration and washed with a small amount of MeOH. This collected material (1.1316 g) was dissolved in MeOH at a concentration of 100 mg/mL and then purified by semi-prep HPLC (Shimadzu system, UV wavelength at 254 and 330 nm, 2 mL/min, Watchers 120 ODS-BP column, 10 μm, 10.0 mm × 250 mm) using 0.1% formic acid in acetonitrile (CH3CN)-0.1% formic acid in H2O (35:65, v/v) as eluent. The resulting eluent was concentrated under reduced pressure to yield compound 2 (tR = 11.2 min, 90.3 mg, 0.014% from crude MeOH extract). The residue recovered from the mother liquor after recrystallization of the EtOAc fraction was fractionated by silica gel column chromatography (CC) using CH2Cl2-MeOH (7:1 → 0:1, v/v) to obtain six fractions (WTE1 ~ WTE6). Fraction WTE3 (31.6 g) was separated by silica gel CC using EtOAc-MeOH (10:1 → 0:1, v/v) into five subfractions (WTE3-1–WTE3-5). Subfraction WTE3-2 (8.0 g) was recrystallized from MeOH to yield compound 3 (3.2 g).
The n-BuOH fraction (182.2 g) was subjected to silica gel CC using EtOAc-MeOH (10:1 → 0:1, v/v), which resulted in seven fractions (WTB1–WTB7). Fraction WTB6 (11.4 g) was separated by semi-prep HPLC (Gilson system, 2 mL/min, INNO C18 column, 10 µm, 10.0 mm × 250 mm) using MeOH-H2O (10:90 → 100:0, v/v, 60 min) to give six subfractions (WTB6-1–WTB6-6). Subfraction WTB6-1 (596.7 mg) was purified by semi-prep HPLC (Shimadzu system, UV wavelength at 254 and 330 nm, 2 mL/min, Watchers 120 ODS-BP column, 10 µm, 10.0 mm × 250 mm) using 0.1% formic acid in CH3CN-0.1% formic acid in H2O (7:93, v/v). The resulting eluent was concentrated under reduced pressure to yield compound 1 (tR = 86.7 min, 4.0 mg).
Preparation of samples and standard solutions for HPLC – The EtOH extract was selected for method validation based on previous findings that it exhibited activity against inflammatory skin disease and had an active constituent profile similar to that of the MeOH extract.21 A stock solution (10 mg/mL) was prepared by solubilizing the dried EtOH extract in MeOH. This solution was then purified by passing it through a 0.45 µm PTFE syringe filter. Compound 1 was purchased as a commercial standard. Compounds 2 and 3 were isolated from the W. trichotoma MeOH extract and purified to serve as reference standards by semi-prep HPLC. Their exact purity levels, determined by HPLC-PDA analysis, were 97.42% for compound 2 and 99.78% for compound 3. Standard stock solutions of the three compounds were prepared in MeOH at 1000 μg/mL, and working solutions were generated by stepwise dilution using MeOH. All sample and standard solutions were filtered through a 0.45 µm PTFE syringe filter immediately before HPLC injection.
Determination of HPLC-PDA conditions – Chromatographic conditions for the quantitative evaluation of the three markers were determined through a step-by-step optimization process based on the single factor approach. Initial mobile phases screening utilized combinations of water, CH3CN, and MeOH. Gradient elution was performed with the addition of acidic modifiers, such as acetic acid, formic acid, or trifluoroacetic acid to the mobile phase. The solvent flow rate was set to 0.5–1.0 mL/min, the column oven temperature to 25, 30, or 35°C, and the detection wavelength to 210 and 400 nm. The injection volume was 10 µL.
HPLC-PDA method validation – The validation procedure followed the ICH Q2(R2) guideline issued by the International Council for Harmonisation of Technical Requirements for Pharmaceuticals for Human Use and the Pharmaceutical Analytical Method Validation Guideline issued by the Ministry of Food and Drug Safety (MFDS) of the Republic of Korea. The process was conducted using recommended parameters, viz., specificity, linearity, limit of detection (LOD) and quantitation (LOQ), precision including repeatability and intermediate precision, and accuracy based on recovery tests.
To confirm the specificity of the developed method, chromatograms of the standard solutions and W. trichotoma extract sample were compared to assess the possibility of interference from co-eluting peaks. The retention times and UV patterns of target compounds in the EtOH extract sample were compared with those of the reference standards to ensure accurate identification.
Linearity was assessed using six concentrations (8.0, 20.0, 51.2, 128.0, 320.0, and 800.0 μg/mL) of each standard compound in MeOH, and each concentration was injected six times. Calibration curves were generated by plotting recorded peak areas against known concentrations. Linear regression analysis was performed to derive the regression equation and coefficient of determination (r2), and to assess model significance using ANOVA (p < 0.001). LOD and LOQ were calculated from calibration curves using the formulas LOD = 3.3 × (σ/S) and LOQ = 10 × (σ/S), where σ denotes the standard deviation (SD) of the intercepts of multiple calibration curves, and S is the slope. LOD represents the lowest concentration detectable above baseline noise, while LOQ denotes the lowest concentration quantifiable with acceptable accuracy and precision.
Validation of precision was performed by gauging intra-day (repeatability) and inter-day (intermediate precision) variation. Repeatability was determined by analyzing the EtOH extract sample at three different concentrations, low (25.0 μg/mL), medium (100.0 μg/mL), and high (400.0 μg/mL), using five replicate injections (n = 5) at each concentration. To determine repeatability, relative standard deviation (RSD, %) was calculated using the results obtained at each concentration. Intermediate precision was assessed by repeating the same procedure over three different days (days 1, 3, and 5) using identical sample preparation conditions, and % RSD was calculated using the results obtained.
Accuracy was assessed by spiking known amounts of the three compounds into W. trichotoma extract at three concentrations (30.0, 150.0, and 360.0 μg/mL). Original concentration refers to the measured level of each analyte in the extract, which was determined by calibration curves. Analysis was conducted in triplicate (n = 3) at each level, and percentage recoveries were calculated using:
Stability testing was conducted to evaluate changes in solution concentration under different storage conditions and durations. Stock solutions of W. trichotoma extract and the three compounds were stored at RT or 4°C for 0, 6, 24, 48, or 72 hours. Five replicate injections were performed at each time point (n = 5). Stability was assessed by calculating the percentage change in peak area and % RSD values versus the peak area of the respective compound at 0 hours.
Statistical analysis – The analysis was conducted using ANOVA in GraphPad Prism (version 9.1.0; GraphPad Software Inc., San Diego, CA, USA). Results are expressed as means ± SDs, and statistical significance was accepted for p values of < 0.05.
Results and Discussion
Chlorogenic acid, miconioside B, and matteucinol 7-O-apiofuranosyl(1→6)-glucopyranoside were isolated from W. trichotoma (Fig. 1). Their structures were determined by comparing spectroscopic data with previous reports, and compounds were identified in the extract by matching retention times and spectral data with those of reference standards.22,23 Based on their relatively high abundances in W. trichotoma extract, as well as their excellent chromatographic resolution and sensitivity, all three compounds were considered appropriate markers for the chemical standardization and quality control. Specifically, these three markers collectively account for over 50% of the total detectable peak area in the W. trichotoma chromatogram, thereby confirming their suitability as principal quantitative representatives for the overall quality control of the extract.
Several preliminary tests were conducted using the single factor approach to optimize chromatographic parameters for the simultaneous determination of the three marker compounds. The use of acidic modifiers in the mobile phase enhances the separation of phenolics by suppressing the ionization of hydroxyl groups, thus improving peak resolution and symmetry.24 Among the acids tested, formic acid was selected because it offered more stable pH control in the acidic range. To promote adequate retention on the reversed-phase column used, the pH was adjusted to remain below the dissociation constant (pKa) of the target compounds, to stabilize their non-ionized forms.25
The UV absorbance profiles of the marker compounds were evaluated. Scanning between 210 and 400 nm showed 325 nm was the optimal detection wavelength for compound 1, whereas compounds 2 and 3 absorbed most at 280 nm. Based on these results, dual-wavelength detection at 280 and 325 nm was employed, which enhanced sensitivity, baseline clarity, and signal-to-noise ratios.
Optimization of the column temperature and flow rate was conducted at 25, 30, and 35°C and from 0.5 to 1.0 mL/min. A temperature of 30°C at a flow rate of 1.0 mL/min was selected, as these settings improved analyte solubility and reduced total run times. The Aegispak C18-L column (4.6 mm × 250 mm, 5 µm) under these conditions provided efficient separation and a shorter analysis time, making it suitable for the routine quantification of the target compounds.
Initial gradient trials were conducted on the W. trichotoma extract using a 5–95% MeOH gradient and a run time of 60 minutes. Although all three marker compounds were detected, these conditions resulted in an excessive run time and inadequate separation. Further optimization led to the selection of a mobile phase consisting of 0.1% formic acid in CH3CN (solvent A) and 0.1% formic acid in water (solvent B) and the following elution gradient conditions: 0 min, 14% A; 3 min, 14% A; 5 min, 30% A; 15 min, 35% A; 20 min, 70% A; and 30 min, 70% A.
The chromatographic conditions finally used were an Aegispak C18-L column (4.6 × 250 mm, 5 µm), column oven temperature 30°C, flow rate 1.0 mL/min, detection wavelengths at 280 and 325 nm, and an injection volume of 10 µL. Under these conditions, the three marker compounds were successfully separated and detected within 20 minutes (Fig. 2).
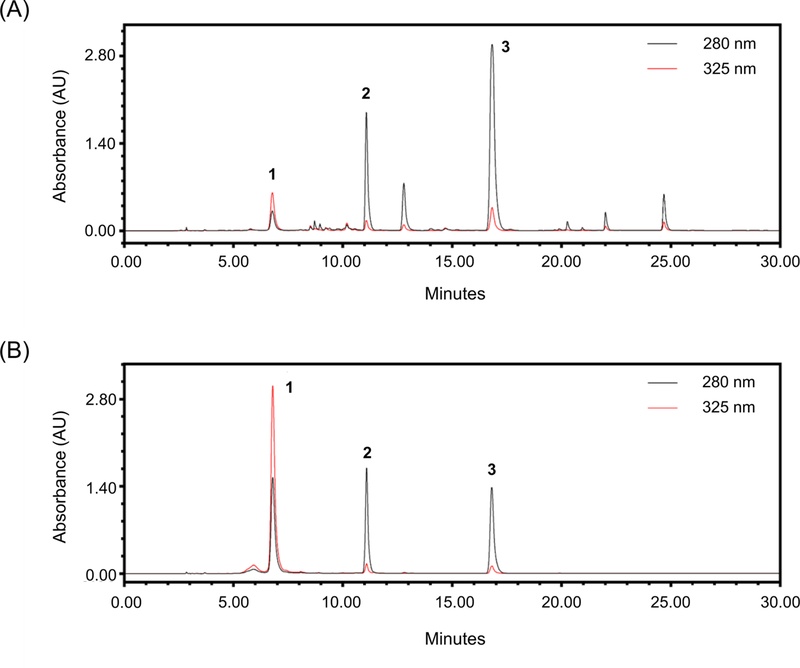
HPLC chromatogram of W. trichotoma extract (A) and a standard mixture (B), detected at 280 and 325 nm, showing chlorogenic acid (1), miconioside B (2), and matteucinol 7-O-apiofuranosyl(1→6)-glucopyranoside (3) peaks.
Specificity was assessed by comparing the HPLC-PDA chromatograms of extract sample and the reference mixture. The three marker compounds showed consistent retention times (tR), maximum absorption wavelengths (λmax), and PDA spectra. Their values were as follows: compound 1 (tR 6.77 min and λmax 240.1 and 325.8 nm), compound 2 (tR 11.08 min and λmax 281.7 and 360.9 nm), and compound 3 (tR 16.83 min and λmax 278.1 and 360.9 nm). The PDA spectra were consistent with the identified compounds (Fig. 3). These results confirm that the developed method enables the selective and accurate identification of the three marker compounds, and show that no interfering peaks were observed.
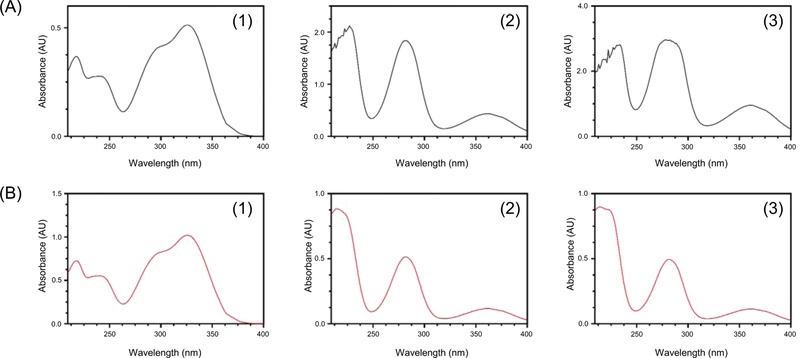
PDA spectra of W. trichotoma extract (A) and standard solutions (B): chlorogenic acid (1), miconioside B (2), and matteucinol 7-O-apiofuranosyl(1→6)-glucopyranoside (3).
Linearity was assessed for all three compounds using six measurements at six concentration levels within the range of 8.0 to 800.0 μg/mL. All regression equations yielded r2 > 0.999, confirming excellent linearity (Table 1). Calculated LODs and LOQs ranged from 4.28 to 6.42 μg/mL and 12.97 to 19.47 μg/mL, respectively. These results demonstrate that the method provides reliable linear responses and sufficient sensitivity for routine quantification.
The precision of the method was evaluated by performing intra-day and inter-day assays on the three marker compounds at three concentration levels (25.0, 100.0, and 400.0 μg/mL). For intra-day determinations, values ranged from 0.41 to 0.86% (1), 0.70 to 1.14% (2), and 0.55 to 1.70% (3). For inter-day determinations, values ranged from 0.68 to 1.55% (1), 1.12 to 1.83% (2), and 0.72 to 1.40% (3). The highest observed values were 1.70% (3, intra-day at 400.0 µg/mL) and 1.83% (2, inter-day at 25.0 μg/mL). Although these values represent the highest variability observed in the study, they remain well within the ICH acceptance limit of 2%. These results collectively demonstrate a high level of precision, confirming the reliability and reproducibility of the developed method. Intra-day accuracy values ranged from 91.90% to 102.53% (1), 99.74% to 100.82% (2) and 98.67% to 104.24% (3). Inter-day accuracy values ranged from 91.55% to 100.77% (1), 100.32% to 103.16% (2) and 99.99% to 104.42% (3), respectively (Table 2).
The accuracy of the devised method was assessed using recoveries at three spiking concentrations (30.0, 150.0, and 360.0 μg/mL). Recoveries and % RSDs are summarized in Table 3. The average recovery values of three marker compounds were 97.9%, 95.7%, and 99.1%, respectively, and % RSD values were < 2%, suggesting minimal variation and reliable quantification. The lowest recovery value obtained was 92.54% for compound 2 at 150.0 g/mL, while the highest was 101.70% for compound 3 at 360.0 g/mL. Together with low % RSD values, these findings support the precision and reproducibility of the method.
Marker stability was evaluated over 72 hours under two storage conditions (RT and 4°C), based on changes in peak area versus initial values and % RSD values. In extract sample solutions, all three compounds maintained consistent peak areas and % RSD values remained below 2.0% over 72 h under both storage conditions. In reference standard solutions, compounds 1–3 generally exhibited greater stability at 4°C than at RT. While compounds 1 and 3 in the standard solutions showed a minor increase in peak area after 24 h at RT, their % RSD values remained below 2.0%. As this trend was not observed in the extract samples, it is likely attributable to minor analytical variability or differences in the sample matrix rather than intrinsic instability of the compounds. Accordingly, it is recommended that W. trichotoma extract samples be kept under refrigeration and used within 72 hours of preparation. Detailed stability results are presented in Table 4.
The validated HPLC-PDA method was applied to the quantitative determination of the three marker compounds in the EtOH extract samples. Analyses were performed in quintuplicate. Compound contents were determined using calibration curves. The concentrations of the three markers 1–3 in the extract were found to be 19.9, 139.8, and 264.9 mg/g, respectively (Table 5). The most abundant marker was compound 3 with a concentration of 264.9 mg/g. By accurately quantifying, this method establishes a critical link between the chemical profile and the pharmacological potential of W. trichotoma.
In conclusion, this study establishes a robust basis for the standardization of W. trichotoma extract through the selection of three major markers from the phenolic and flavonoid classes. Simultaneously quantifying these markers enables a more comprehensive and holistic management of the extract’s chemical profile compared to single-analyte detection methods. Furthermore, the reliance of the method on the widely available HPLC-PDA system ensures high technical accessibility and cost-effectiveness for industrial quality control. This validated platform will be helpful for future structure-activity relationship studies and for correlating batch-to-batch chemical variation with observed in vivo efficacy.
Acknowledgments
This work was supported by a 2-Year Research Grant of Pusan National University.
Conflicts of Interest
The authors declare that they have no conflicts of interest.
References
-
Huang, W.; Wang, Y.; Tian, W.; Cui, X.; Tu, P.; Li, J.; Shi, S.; Liu, X. Antibiotics 2022, 11, 1380.
[https://doi.org/10.3390/antibiotics11101380]

-
Dar, R. A.; Shahnawaz, M.; Ahanger, M. A.; Majid, I. U. J. Phytopharm. 2023, 12, 189–195.
[https://doi.org/10.31254/phyto.2023.12307]

-
Roy, A.; Khan, A.; Ahmad, I.; Alghamdi, S.; Rajab, B. S.; Babalghith, A. O.; Alshahrani, M. Y.; Islam, S.; Islam, M .R. Biomed Res. Int. 2022, 2022, 5445291.
[https://doi.org/10.1155/2022/5445291]

-
Salam, U.; Ullah, S.; Tang, Z.-H.; Elateeq, A. A.; Khan, Y.; Khan, J.; Khan, A.; Ali, S. Life 2023, 13, 706.
[https://doi.org/10.3390/life13030706]

-
Al-Khayri, J. M.; Rashmi, R.; Toppo, V.; Chole, P. B.; Banadka, A.; Sudheer, W. N.; Nagella, P.; Shehata, W. F.; Al-Mssallem, M. Q.; Alessa, F. M.; Almaghasla, M. I.; Rezk, A. A-S. Metabolites 2023, 13, 716.
[https://doi.org/10.3390/metabo13060716]

-
Zhang, L.; Ravipati, A. S.; Koyyalamudi, S. R.; Jeong, S. C.; Reddy, N.; Smith, P. T.; Bartlett, J.; Shanmugam, K.; Münch, G.; Wu, M. J. J. Agric. Food Chem. 2011, 59, 12361–12367.
[https://doi.org/10.1021/jf203146e]

-
Kumar, M. S.; Adki, K. M. Biomed. Pharmacother. 2018, 105, 233–245.
[https://doi.org/10.1016/j.biopha.2018.05.142]

-
Nunes, C. dos R.; Barreto Arantes, M.; Menezes de Faria Pereira, S.; Leandro da Cruz, L.; de Souza Passos, M.; Pereira de Moraes, L.; Vieira, I. J. C.; Barros de Oliveira, D. Molecules 2020, 25, 3726.
[https://doi.org/10.3390/molecules25163726]

-
Huan, D. Q.; Hop, N. Q.; The Son, N. Curr. Pharm. Biotechnol. 2024, 25, 563–598.
[https://doi.org/10.2174/1389201024666230606122116]

-
Lei, J.-P.; Yuan, J.-J.; Pi, S.-H.; Wang, R.; Tan, R.; Ma, C.-Y.; Zhang, T.; Jiang, H.-Z. Pharmacogn. Mag. 2017, 13, 488–491.
[https://doi.org/10.4103/pm.pm_275_16]

-
Wu, M.; Su, X.; Wu, Y.; Luo, Y.; Guo, Y.; Xue, Y. Beilstein J. Org. Chem. 2022, 18, 200–207.
[https://doi.org/10.3762/bjoc.18.23]

-
Lu, C.-L.; Zhu, L.; Piao, J.-H.; Jiang, J.-G. Pharm. Biol. 2012, 50, 225–231.
[https://doi.org/10.3109/13880209.2011.596207]

-
Keem, M.-J.; Kim, T.-Y.; Park, N.-J.; Choi, S.; Paik, J.-H.; Jo, B.-G.; Kwon, T.-H.; Kim, S.-N.; Lee, S. R.; Yang, M. H. Nutrients 2025, 17, 1552.
[https://doi.org/10.3390/nu17091552]

-
Balekundri, A.; Mannur, V. Future J. Pharm. Sci. 2020, 6, 67.
[https://doi.org/10.1186/s43094-020-00091-5]

-
Dar, A. A.; Raina, A.; Kumar, A. Biomed. Chromatogr. 2022, 36, e5408.
[https://doi.org/10.1002/bmc.5408]

-
Zhang, C.; Zheng, X.; Ni, H.; Li, P.; Li, H.-J. J. Pharm. Biomed. Anal. 2018, 159, 296–304.
[https://doi.org/10.1016/j.jpba.2018.07.006]

-
He, Y.; Mao, S.; Zhao, Y.; Yang, J. Foods 2025, 14, 1914.
[https://doi.org/10.3390/foods14111914]

-
Chandimala, U. R.; Ajtony, Z. LWT-Food Sci. Technol. 2025, 232, 118416.
[https://doi.org/10.1016/j.lwt.2025.118416]

- International Council for Harmonisation of Technical Requirements for Pharmaceuticals for Human Use (ICH). Validation of analytical procedures: Text and methodology Q2(R2). Geneva, Switzerland, 2023.
- Ministry of Food and Drug Safety (MFDS). Guideline on Validation of Analytical Procedures for Drugs; Ministry of Food and Drug Safety: Cheongju, Republic of Korea, 2015.
-
Keem, M.-J.; Jo, B.-G.; Lee, S. H.; Kim, T.-Y.; Jung, Y. S.; Jeong, E.-J.; Kim, K. H.; Kim, S.-N.; Yang, M. H. J. Ethnopharmacol. 2024, 333, 118398.
[https://doi.org/10.1016/j.jep.2024.118398]

-
Han, T.; Li, H.; Zhang, Q.; Zheng, H.; Qin, L. Chem. Nat. Compd. 2006, 42, 567–570.
[https://doi.org/10.1007/s10600-006-0215-2]

-
Gimenez, V. M. M.; e Silva, M. L. A.; Cunha, W. R.; Januario, A. H.; Costa, E. J. X.; Pauletti, P. M. Biochem. Syst. Ecol. 2020, 91, 104049.
[https://doi.org/10.1016/j.bse.2020.104049]

-
Klimek-Turek, A.; Dzido, T.H. Adsorption 2010, 16, 287–294.
[https://doi.org/10.1007/s10450-010-9241-2]

-
Kubik, Ł.; Struck-Lewicka, W.; Kaliszan, R.; Wiczling, P. J. Chromatogr. A 2015, 1416, 31–37.
[https://doi.org/10.1016/j.chroma.2015.09.003]