
5,7-Dimethoxyflavone Augments Oolonghomobisflavan B-Mediated Apoptotic Cell Death in Multiple Myeloma Cells through Activation of the 67LR/cGMP/ASM Signaling Axis
Abstract
Oolonghomobisflavan B (OHBFB) is a prominent and specific polyphenolic constituent of oolong tea. We previously identified a cell membrane receptor for OHBFB and showed that it induces cancer-selective cell death in vitro and in vivo, although its standalone anticancer efficacy is limited. 5,7-Dimethoxyflavone (5,7-DMF), a principal bioactive component of black ginger (Kaempferia parviflora), also exhibits anticancer activity. Here, we demonstrate that 5,7-DMF synergistically enhances the anticancer effect of OHBFB in multiple myeloma cells by boosting OHBFB-induced intracellular cyclic guanosine monophosphate (cGMP) production. Moreover, co-treatment with OHBFB and 5,7-DMF triggers apoptotic cell death in multiple myeloma cells, accompanied by activation of the cGMP/acid sphingomyelinase (ASM) pathway. In summary, 5,7-DMF amplifies the anticancer activity of OHBFB by upregulating cGMP in multiple myeloma cells.
Keywords:
Oolonghomobisflavan B, 5,7-Dimethoxyflavone, Anti-cancer, Multiple myeloma, Cyclic guanosine monophosphateIntroduction
Oolong tea and one of its major polyphenolic constituents, oolonghomobisflavan B (OHBFB), exhibit anticancer properties.1–3 We previously identified the 67-kDa laminin receptor (67LR) as the cell-surface sensor for OHBFB.4–6 67LR is overexpressed across many cancer types, including multiple myeloma.6–10 Consistent with this, OHBFB selectively induces apoptosis in cancer cells while sparing normal cells by targeting 67LR.6–9 OHBFB triggers apoptosis in several malignancies—such as multiple myeloma, pancreatic cancer, and prostate cancer—via cGMP-dependent mechanisms,6 supporting its potential for clinical evaluation.9–11 Nonetheless, at physiological concentrations, OHBFB alone does not elicit sufficiently robust anticancer effects.
cGMP is a second messenger that mediates diverse physiological actions, including anticancer effects.6 Intracellular cGMP levels are elevated by soluble guanylate cyclase (sGC), the receptor for nitric oxide (NO).6,12–17 Cyclic nucleotide phosphodiesterases (PDEs) are key regulators of cGMP and/or cAMP signaling, hydrolyzing and thereby inactivating these messengers in mammalian tissues. PDE5 is a cGMP-specific enzyme that degrades intracellular cGMP.18–21 Inhibition of PDE activity has emerged as a therapeutic strategy in various diseases.18–20; PDE5 inhibitors, for example, are clinically effective for erectile dysfunction22 and have been shown to enhance cGMP-dependent apoptotic pathways in multiple cancer cell types.6,7
Black ginger (Kaempferia parviflora), commonly referred to as black turmeric, contains diverse flavonoids and is particularly rich in polymethoxyflavones. Among them, 5,7-dimethoxyflavone (5,7-DMF) is a major component and a potent PDE5 inhibitor.23 Multiple studies also report health-promoting activities of 5,7-DMF, including anticancer effects.24–26 In this study, we selected 5,7-dimethoxyflavone (5,7-DMF) as a mechanistically aligned partner to potentiate OHBFB-mediated anticancer activity. Because OHBFB activates a 67LR-linked, cGMP-dependent signaling axis, we hypothesized that sustaining intracellular cGMP would amplify downstream pro-apoptotic outputs such as ASM activation; notably, 5,7-DMF (a major polymethoxyflavone from Kaempferia parviflora) has been reported to exhibit PDE5-inhibitory activity and thus may promote cGMP accumulation. Moreover, our prior work showed that 5,7-DMF synergistically enhanced catechin-mediated apoptosis in multiple myeloma cells in association with increased cGMP signaling, supporting its use here as a mechanistic probe. Although many dimethoxyflavone analogs exist, they are not equally characterized for cGMP modulation and synergy in MM models, and systematic structure–activity comparisons will be addressed in future studies.
Here, we demonstrate that 5,7-DMF synergistically potentiates the anticancer activity of OHBFB in multiple myeloma cells, concomitant with enhanced OHBFB-induced upregulation of intracellular cGMP.
Experimental
Cell culture – 5,7-dimethoxyflavone (5,7-DMF; purity ≥98.0% by HPLC) was purchased from Wako (Tokyo, Japan). Bay 41-2272 was obtained from Enzo Life Sciences (Crescent, Ex, UK). oolonghomobisflavan B (OHBFB) was obtained from Nagara science (Kyoto, Japan). Catalase, and superoxide dismutase (SOD) were obtained from Sigma-Aldrich (St Louis, MO, USA). The human multiple myeloma cell line U266, at a density of 5 × 104 cells/mL, was cultured in RPMI 1640 supplemented with 1% (v/v) foetal bovine serum, 200 U/mL catalase, and 5 U/mL superoxide dismutase (SOD) from Sigma-Aldrich (St Louis, MO, USA) at 37°C, 100% humidity, and 5% CO2. After 72 h, cell viability was determined using trypan blue staining assay.
Analyzed apoptotic cell death – 2 × 104 cells/mL of U266 cells were cultured in 1% FBS-RPMI 1640 at 100% humidity and 5% CO2, at 37°C. Apoptotic cells were determined using a flow cytometric test. Annexin-V+ cells were evaluated by combining early Annexin-V+ propidium iodide− and late Annexin-V+ propidium iodide+ cells after treatment with OHBFB or/and 5,7-DMF. The analysis was performed after 72 hours using a VerseTM system from BD.
Western blot analysis – U266 cells were seeded onto a 12-well plate at a density of 1 × 106 cells/well or 5 × 104 cells/well and treated with 1 μM OHBFB or/and 10 μM 5,7-DMF for 3 h or 72 h. After treatment, cells were lysed in a solution containing 50 mM Tris-HCl (pH 7.5), 150 mM NaCl, 1% Triton X-100, 2 mg/mL aprotinin, 50 mM NaF, 1 mM phenylmethanesulfonyl fluoride, 30 mM Na4P2O7, 1 mM pervanadate, and 1 mM ethylenediaminetetraacetic acid. SDS-PAGE (sodium dodecyl sulphate-polyacrylamide gel electrophoresis) was performed as previously described.9 Approximately 50 μg of protein was suspended in Laemmli sample buffer (0.1 M Tris-HCl buffer, pH 6.8; 0.05% mercaptoethanol; 1% SDS; 0.001% bromophenol blue; and 10% glycerol) and boiled before electrophoresis on SDS-polyacrylamide gels. The gels were then transferred to Trans-Blot nitrocellulose membranes (Bio-Rad) using electrob-lotting. Primary antibodies were incubated in Tween 20-TBS containing 1% BSA. The blots were washed with Tween 20-TBS and then incubated in anti-rabbit HRP conjugates (secondary antibody). The primary antibody used at a 1:3000 dilution was incubated overnight at 4°C, followed by the secondary antibody at a 1:10000 dilution for 1 hour. The anti-eNOS (H-159) antibody was purchased from Santa Cruz Biotechnology, the anti-Laminin-R (MLuC5) antibody was purchased from Santa Cruz Biotechnology, the anti-IgM (A-7) antibody was purchased from Santa Cruz Biotechnology, the anti-67 kDa Laminin Receptor (RPSA/2699) antibody was purchased from abcam (Cambridge, UK), and the anti-β actin antibody was purchased from Sigma-Aldrich (St Louis, MO, USA).
Quantitative reverse transcription PCR (qRT-PCR) – U266 cells were cultured with the presence of 1 μM OHBFB or/and 10 μM 5,7-DMF. After 3 h, the culture medium was removed, and the cells were washed 2 times with PBS. After then, cells were collected by centrifugation at 500 × g for 5 min. Total RNA was isolated using the Qiagen RNeasy Mini Kit from QIAGEN (Hilden, Germany) according to the manufacturer’s instructions. Primers were used for homo sapiens ribosomal protein SA (RPSA, 67LR), Forward: 5′- GCAGCAGGAACCCACTTAGG-3′, Reverse: 5′-GCAGCAGCAAACTTCAGCAC-3′; Human GAPDH primer, Forward: 5′-CCACTCCTCCACCTTTGACG-3′ (upstream), Reverse: 5′-CCACCACCCTGTTGCTGTAG -3′. cDNA was synthesized from total RNA using the cDNA Master Mix from Applied Biosystems (CA, USA). qRT-PCR was conducted using 2 mL cDNA and SYBR Green PCR 2 × Master Mix from Applied Biosystems (CA, USA) with 40 cycles of 15 seconds at 95°C, 60 seconds at 60°C according to the manufacturer’s instructions. The data were analyzed using v2.1 StepOne software from Applied Biosystems (CA, USA).
cGMP assays – Measurements of intracellular cGMP production were carried out using the TRFRET cGMP assay kit (Perkin-Elmer) following the manufacturer’s protocol. The cells were treated with OHBFB and/or 5,7-DMF for 3 hours in 96-well plates, and the plate assessment was performed using the EnVisionTM Plate Reader (Perkin-Elmer).
ASM activity measurement – U266 cells were lysed with lysis buffer and incubated for 1 hour at 4°C, followed by centrifugation for 20 minutes at 15,000 × g. The resulting supernatant was incubated with substrate buffer, including 200 mM sodium acetate, 1% Triton X-100, and 400 pmol BODIPY-C12 sphingomyelin in dH2O, for 18 hours at 37°C. BODIPY-C12-sphingomyelin was obtained from Sigma-Aldrich (St. Louis, MO, USA).
Statistical analysis – Our data are indicated as mean ± SEM. Calculation the IC50 values and isobologram methods were determined by using the Graphpad software. Tukey’s test was used to assess the significance of differences between experimental variables. Statistical analyses were performed using KyPlot software (Kyens Lab, Tokyo, Japan).
Results and Discussion
We used commercially purchased standards for both compounds, 5,7-dimethoxyflavone (5,7-DMF) and oolong-homobisflavan B (OHBFB) (Fig. 1). OHBFB (Fig. 1A) is a dimer of (−)-epigallocatechin-3-O-gallate (EGCG), a major catechin-derived polyphenol, and 5,7-DMF (Fig. 1B) is a polymethoxyflavone abundant in Kaempferia parviflora. Establishing these structures provides context for their complementary actions on 67LR and cGMP signaling described below.
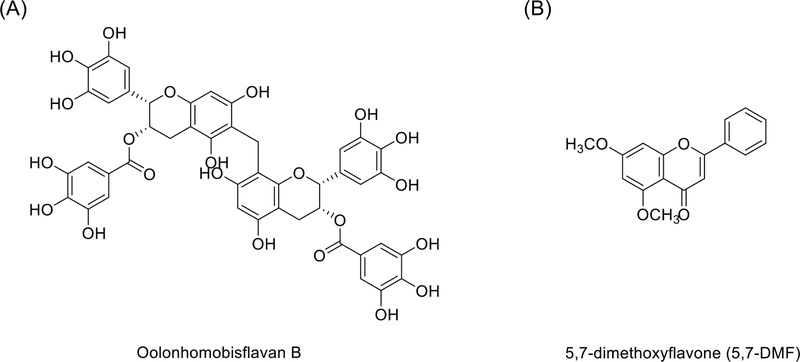
Molecular structure of oolonghomobisflavan B (OHBFB) and 5,7-dimethoxyflavone (5,7-DMF). Based on chemical structure of (A) OHBFB and (B) 5,7-DMF.
Consistent with prior reports that OHBFB selectively induces apoptosis via the 67-kDa laminin receptor (67LR),8,9,27 its efficacy at physiological concentrations is limited.28 We therefore evaluated combination effects with 5,7-DMF in U266 and RPMI 8226 multiple myeloma (MM) cells. In U266 cells, OHBFB and 5,7-DMF displayed IC50 values of 5.16 µM and 65.45 µM, respectively, whereas their combination yielded markedly lower apparent IC50 values (3.70 µM or 2.16 µM depending on the mixing ratio), indicating enhanced potency (Fig. 2A–D). Isobologram analysis, constructed from single-agent IC50s (x-axis: OHBFB 5.16 µM; y-axis: 5,7-DMF 65.45 µM), showed combination data points falling below the line of additivity, demonstrating synergy in both U266 and RPMI 8226 cells (Fig. 2E, F). Importantly, the same concentrations of OHBFB and/or 5,7-DMF did not reduce viability of normal PBMCs over 72 h (Fig. 2G–I), supporting cancer selectivity of the combination.
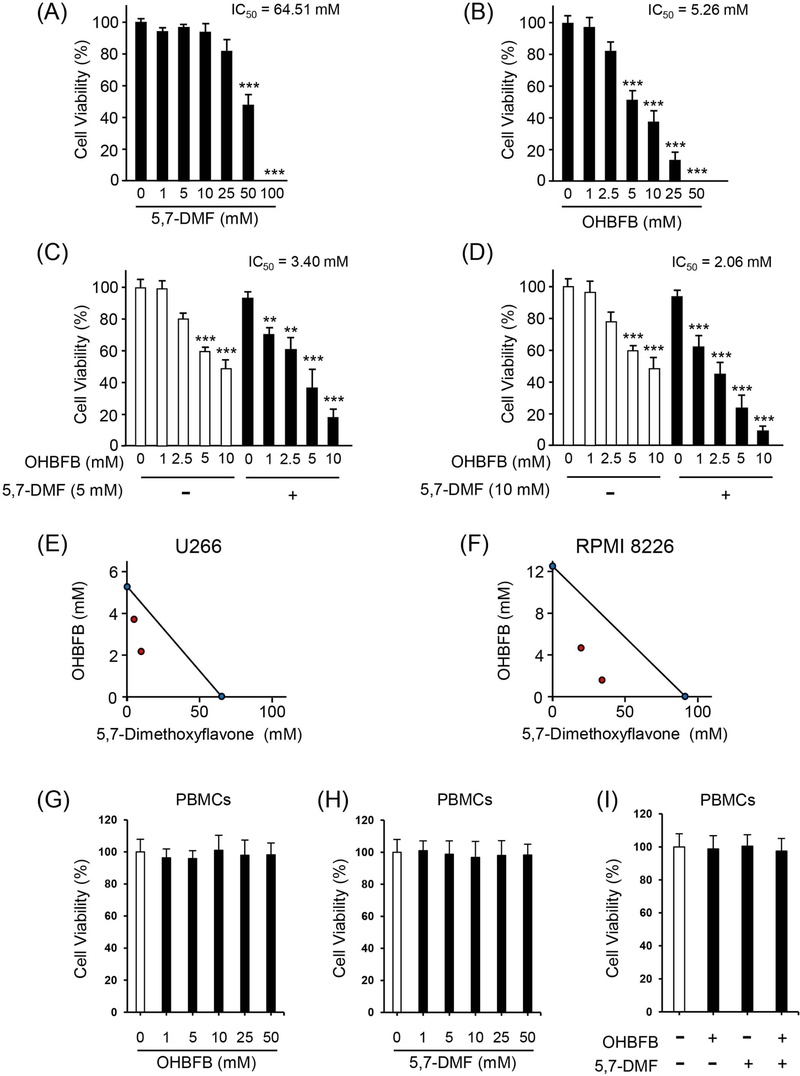
5,7-DMF synergistically potentiates cell death effect of OHBFB in multiple myeloma cells. (A, B) Based on chemical structure of OHBFB and 5,7-DMF. (A–D) The human multiple myeloma cell line U266 was cultured with or without 5,7-DMF and/or OHBFB at the indicated concentration for 72 h, and the viability of U266 was measured by trypan blue staining assay. (E, F) The combination effect of OHBFB and 5,7-DMF was measured by isobologram analysis in U266 (E) and RPMI 8226 (F) cells. Data are presented as mean ± SEM (n = 3), Tukey’s test, *p < 0.05, **p < 0.01, ***p < 0.001.
OHBFB has been reported to induce anticancer effects via apoptosis in cancer cells.6,9 In U266 cells, neither 1 µM OHBFB nor 10 µM 5,7-DMF alone induced significant apoptosis; however, the combination (1 µM OHBFB + 10 µM 5,7-DMF) robustly increased apoptotic cell death (Fig. 3). These data indicate that 5,7-DMF effectively lowers the apoptotic threshold for OHBFB in MM cells, converting subeffective single-agent doses into a pro-apoptotic regimen.
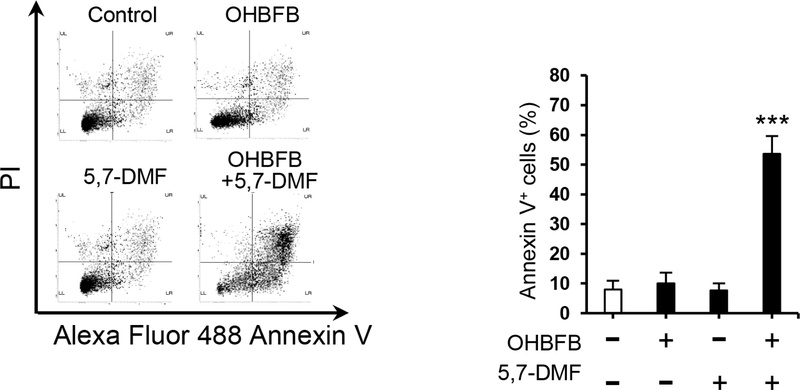
Combination of OHBFB and 5,7-DMF induces apoptotic cell death in U266 cells. U266 cells were treated with OHBFB (1 μM) and/or 5,7-DMF (10 μM) for 72 h. Apoptotic cells were stained with propidium iodide and Annexin V-Alexa Fluor 488. Data are presented as mean ± SEM (n = 3). ***p < 0.001.
Mechanistic studies implicated potentiation of cGMP signaling and downstream acid sphingomyelinase (ASM) activation. OHBFB increased intracellular cGMP at 5 µM but not at 0–2.5 µM (Fig. 4A), establishing a concentration threshold for second-messenger engagement. Addition of 5,7-DMF significantly boosted OHBFB-induced cGMP (Fig. 4B) and synergistically enhanced ASM activity (Fig. 4C), consistent with 5,7-DMF functioning as a PDE5 inhibitor that preserves the cGMP pool. To further probe the pathway, we combined 5,7-DMF with the soluble guanylate cyclase (sGC) stimulator Bay 41-2272. The combination increased cGMP production (Fig. 4D) and augmented Bay 41-2272–elicited ASM activation (Fig. 4E), reinforcing that PDE5 inhibition amplifies cGMP-dependent signaling downstream of sGC. Notably, while OHBFB engages cGMP signaling, our data indicate that combination with 10 µM 5,7-DMF does augments cGMP availability, thereby facilitating ASM-mediated, ceramide-linked apoptosis.
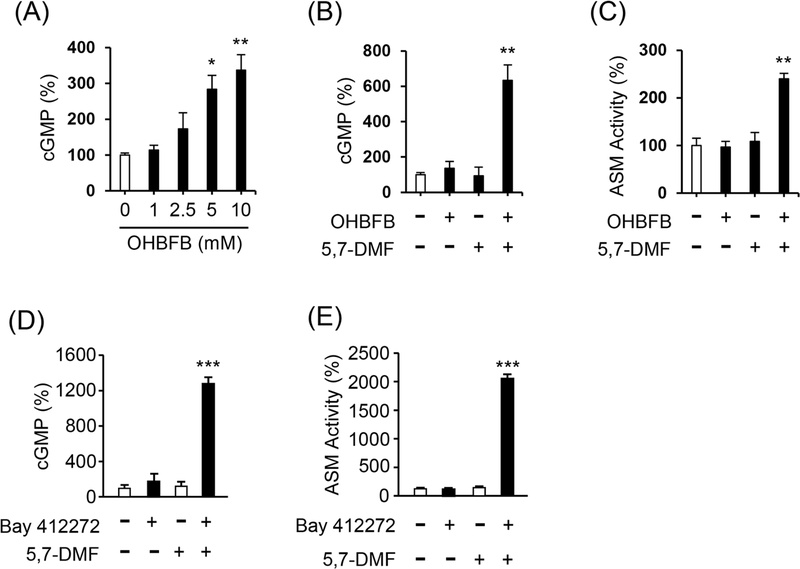
5,7-DMF potentiated OHBFB-induced apoptosis via activation of cGMP/ASM pathway. (A) U266 cells were treated with OHBFB (1 μM) and/or 5,7-DMF (10 μM). After one hour, intracellular cGMP levels were measured. (B–D) U266 cells were treated with OHBFB, Bay 412272 and/or 5,7-DMF. After 1 h, (B, D) intracellular cGMP levels or (C) ASM activity were determined. (E) U266 cells were treated with Bay 412272 (1 μM) and/or 5,7-DMF (10 μM). After 72 h, ASM activity was assessed using TLC. Data are presented as mean ± SEM (n = 3). *p < 0.05, **p < 0.01, ***p < 0.001.
Because anti-67LR antibody treatment is widely used to assess the role of 67LR in OHBFB-induced cell death,6 we tested its necessity for the combination effect. Pretreatment with an anti-67LR antibody significantly attenuated the combination’s effects on viability (Fig. 5A), intracellular cGMP (Fig. 5B), and ASM activity (Fig. 5C), establishing 67LR as an essential upstream node. In contrast, qRT-PCR showed that OHBFB and/or 5,7-DMF did not alter 67LR mRNA levels (Fig. 5D), indicating that synergy arises from functional amplification of 67LR-linked signaling rather than transcriptional upregulation of the receptor.
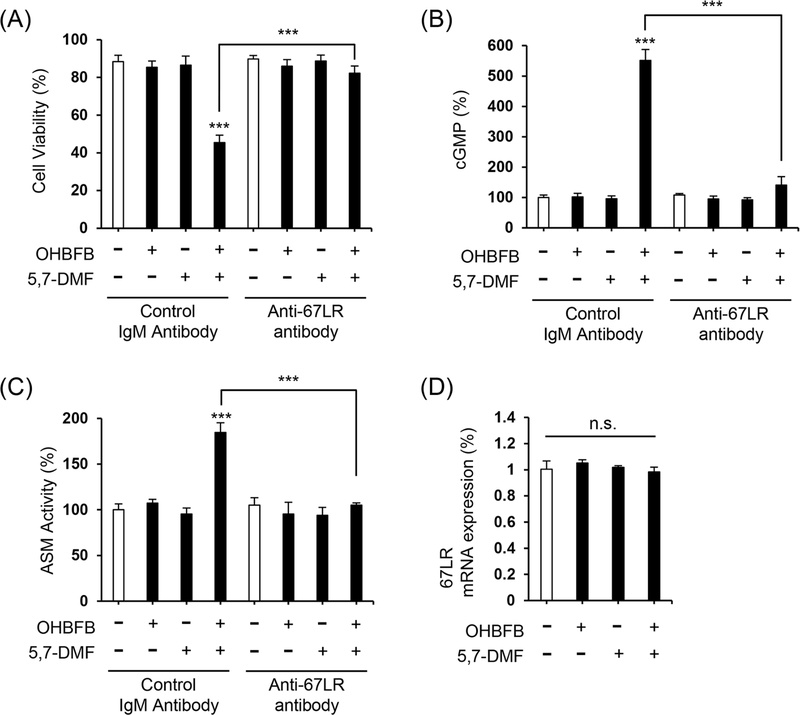
Combined OHBFB and 5,7-DMF induce cell death via 67LR in u266 cells without affecting normal cells. (A) After a 2 h pre-treatment with control IgM antibody or MLuC5 (anti-67LR antibody), U266 cells were exposed to 1 µM OHBFB and/or 10 µM 5,7-DMF for 72 h. Cell viability was evaluated using the trypan blue method (n = 3). (B) Primary peripheral blood mononuclear cells (PBMC) were subjected to treatment with 1 µM OHBFB and/or 10 µM 5,7-DMF for 72 h, and the mRNA expression of 67LR was quantified using real-time PCR analysis (n = 3). n.s.: not significant. Data are presented as mean ± SEM (n = 3). ***p < 0.001.
Together, these findings define a 67LR → cGMP → ASM axis as a tractable vulnerability in multiple myeloma and show that preserving cGMP signaling can enhance cancer-selective apoptosis via downstream ASM activation.29–32 The combination triggers apoptosis at doses where single agents are ineffective and spares normal PBMCs, supporting a favorable therapeutic window. These results align with prior reports that pharmacologic PDE5 inhibition augments polyphenol-driven, 67LR-dependent cGMP signaling and apoptosis; for example, vardenafil synergizes with EGCG in myeloma models.6 We previously reported that 5,7-dimethoxyflavone (5,7-DMF) potentiates EGCG-induced apoptosis in U266 multiple myeloma cells by enhancing intracellular cGMP signaling and activating downstream ASM/caspase-3 pathways. In the present study, we extend this cGMP-centered synergy framework to a distinct, oolong-tea–derived polyphenol ligand, oolonghomobisflavan B (OHBFB), which—despite structural relatedness to EGCG—constitutes a different natural product entity and therefore warrants independent validation. Importantly, we demonstrate synergistic cytotoxicity in two multiple myeloma cell lines (U266 and RPMI 8226) and show that the effective combination doses do not reduce the viability of normal PBMCs, supporting cancer selectivity. Mechanistically, we further establish that 67LR is required not only for the viability phenotype but also for combination-induced cGMP accumulation and ASM activation, thereby defining a functional 67LR → cGMP → ASM signaling axis as the key vulnerability engaged by OHBFB/5,7-DMF co-treatment. However, a key limitation of this study is that PDE5 involvement was not validated by direct enzymatic activity measurements or pharmacological benchmarking with a selective PDE5 inhibitor. Future work should include PDE5 activity assays and/or comparative experiments using PDE5-specific inhibitors (and, where feasible, genetic approaches such as PDE5A knockdown/overexpression) to confirm whether PDE5 is the primary mediator of the cGMP-related effects observed with 5,7-DMF. Mechanistically, ASM activation promotes ceramide-dependent apoptosis in cancer cells, supporting the central role of the cGMP/ASM axis in this context.33 Although our combination data were generated in MM cell lines, in vivo validation will be important; notably, OHBFB binds 67LR with micromolar affinity and suppresses tumor growth in vivo in melanoma, underscoring the translational potential of 67LR-targeted polyphenols.4 The broader relevance of 67LR/cGMP-mediated apoptosis has been demonstrated in other cancers (e.g., coptisine in hepatocellular carcinoma), further supporting this axis as a therapeutically actionable pathway.34 Beyond biology, combination optimization and pharmacodynamics will require careful modeling; recent MM studies provide frameworks to quantify and predict clinical synergy from single-agent and fixed-ratio data.35 Finally, prospective work should characterize pharmacokinetics, potential drug–drug interactions, and dosing schedules in vivo to translate these mechanistic synergies into effective therapies.
In conclusion, 5,7-DMF synergistically potentiates OHBFB-induced apoptosis in multiple myeloma by functionally amplifying 67LR/cGMP/ASM signaling. By overcoming the efficacy ceiling of OHBFB at physiological concentrations while sparing normal PBMCs, this natural product-based combination offers a promising, cancer-selective approach. Future work should evaluate efficacy in vivo, delineate ceramide-mediated apoptotic mechanisms, and optimize translational dosing paradigms.
Acknowledgments
This research was funded by the National Research Foundation of Korea (NRF) grant funded by Korea government (MSIT) (NRF-2021R1C1C2095006). This research was supported by the Daegu Haany University Regional Innovation System & Education (RISE) Glocal project program [Global Joint Research on Traditional Medicine and K-Beauty] through the Gyeongbook RISE center, funded by the Ministry of Education (MOE) and the Gyeongsangbookdo, Republic of Korea. This research was supported by the Regional Innovation System & Education (RISE) Local Customized R&D (University Autonomous Local Customized R&D) program through the Gyeongbuk RISE CENTER, funded by the Ministry of Education (MOE) and the Gyeongsangbuk-do, Republic of Korea (2025-rise-15-110).
Conflicts of Interest
The authors declare that they have no conflicts of interest.
References
-
Chen, B.-H.; Hsieh, C.-H.; Tsai, S.-Y.; Wang, C.-Y.; Wang, C.-C. Sci Rep. 2020, 10, 5163.
[https://doi.org/10.1038/s41598-020-62136-2]

-
Naganuma, T.; Kuriyama, S.; Kakizaki, M.; Sone, T.; Nakaya, N.; Ohmori-Matsuda, K.; Hozawa, A.; Nishino, Y.; Tsuji, I. Am. J. Epidemiol. 2009, 170, 730–738.
[https://doi.org/10.1093/aje/kwp187]

-
Bettuzzi, S.; Brausi, M.; Rizzi, F.; Castagnetti, G.; Peracchia, G.; Corti, A. Cancer Res. 2006, 66, 1234–1240.
[https://doi.org/10.1158/0008-5472.CAN-05-1145]

-
Bae, J.; Kumazoe, M.; Lee, K.-W.; Fujimura, Y.; Tachibana, H. Phytomedicine 2023, 118, 154970.
[https://doi.org/10.1016/j.phymed.2023.154970]

-
Tachibana, H.; Koga, K.; Fujimura, Y.; Yamada, K. Nat. Struct. Mol. Biol. 2004, 11, 380–381.
[https://doi.org/10.1038/nsmb743]

- Kumazoe, M.; Sugihara, K.; Tsukamoto, S.; Huang, Y.; Tsurudome, Y.; Suzuki, T.; Suemasu, Y.; Ueda, N.; Yamashita, S.; Kim, Y.; Yamada, K.; Tachibana, H. J. Clin. Invest. 2013, 123, 787–799.
-
Kumazoe, M.; Tsukamoto, S.; Lesnick, C.; Kay, N. E.; Yamada, K.; Shanafelt, T. D.; Tachibana, H. Br. J. Haematol. 2015, 168, 610–613.
[https://doi.org/10.1111/bjh.13135]

-
Britschgi, A.; Simon, H.-U.; Tobler, A.; Fey, M. F.; Tschan, M. P. Br. J. Haematol. 2010, 149, 55–64.
[https://doi.org/10.1111/j.1365-2141.2009.08040.x]

-
Tsukamoto, S.; Hirotsu, K.; Kumazoe, M.; Goto, Y.; Sugihara, K.; Suda, T.; Tsurudome, Y.; Suzuki, T.; Yamashita, S.; Kim, Y.; Huang, Y.; Yamada, K.; Tachibana, H. Biochem. J. 2012, 443, 525–534.
[https://doi.org/10.1042/BJ20111837]

-
Tsukamoto, S.; Huang, Y.; Umeda, D.; Yamada, S.; Yamashita, S.; Kumazoe, M.; Kim, Y.; Murata, M.; Yamada, K.; Tachibana, H. J. Biol. Chem. 2014, 289, 32671–32681.
[https://doi.org/10.1074/jbc.M114.604983]

-
Yamada, S.; Tsukamoto, S.; Huang, Y.; Makio, A.; Kumazoe, M.; Yamashita, S.; Tachibana, H. Sci. Rep. 2016, 6, 19225.
[https://doi.org/10.1038/srep19225]

-
Linder, A. E.; McCluskey, L. P.; Cole, K. R., 3rd; Lanning, K. M.; Webb, R. C. J. Pharmacol. Exp. Ther. 2005, 314, 9–15.
[https://doi.org/10.1124/jpet.105.083634]

-
Russwurm, M.; Wittau, N.; Koesling, D. J. Biol. Chem. 2001, 276, 44647–44652.
[https://doi.org/10.1074/jbc.M105587200]

-
Schoser, B. G.; Behrends, S. Neuroreport 2001, 12, 979–981.
[https://doi.org/10.1097/00001756-200104170-00023]

-
Zabel, U.; Kleinschnitz, C.; Oh, P.; Nedvetsky, P.; Smolenski, A.; Müller, H.; Kronich, P.; Kugler, P.; Walter, U.; Schnitzer, J. E.; Schmidt, H. H. W. Nat. Cell Biol. 2002, 4, 307–311.
[https://doi.org/10.1038/ncb775]

-
Warner, T. D.; Mitchell, J. A.; Sheng, H.; Murad, F. Adv. Pharmacol. 1994, 26, 171–194.
[https://doi.org/10.1016/S1054-3589(08)60054-X]

-
Buechler, W. A.; Ivanova, K.; Wolfram, G.; Drummer, C.; Heim, J. M.; Gerzer, R. Ann. N. Y. Acad. Sci. 1994, 714, 151–157.
[https://doi.org/10.1111/j.1749-6632.1994.tb12039.x]

-
Lugnier, C. Pharmacol. Ther. 2006, 109, 366–398.
[https://doi.org/10.1016/j.pharmthera.2005.07.003]

-
Keravis, T.; Lugnier, C. Br. J. Pharmacol. 2012, 165, 1288–1305.
[https://doi.org/10.1111/j.1476-5381.2011.01729.x]

-
Porst, H.; Rosen, R.; Padma-Nathan, H.; Goldstein, I.; Giuliano, F.; Ulbrich, E.; Bandel, T. Int. J. Impot. Res. 2001, 13, 192–199.
[https://doi.org/10.1038/sj.ijir.3900713]

-
Loughney, K.; Hill, T. R.; Florio, V. A.; Uher, L.; Rosman, G. J.; Wolda, S. L.; Jones, B. A.; Howard, M. L.; McAllister-Lucas, L. M.; Sonnenburg, W. K.; Francis, S. H.; Corbin, J. D.; Beavo, J. A.; Ferguson, K. Gene 1998, 216, 139–147.
[https://doi.org/10.1016/S0378-1119(98)00303-5]

-
Huang, S. A.; Lie, J. D. P. T. 2013, 38, 407–419.
[https://doi.org/10.1007/s12046-013-0143-3]

-
Temkitthawon, P.; Hinds, T. R.; Beavo, J. A.; Viyoch, J.; Suwanborirux, K.; Pongamornkul, W.; Sawasdee, P.; Ingkaninan, K. J. Ethnopharmacol. 2011, 137, 1437–1441.
[https://doi.org/10.1016/j.jep.2011.08.025]

-
Song, Y.; Kim, M.-B.; Kim, C.; Kim, J.; Hwang, J.-K. J. Med. Food 2016, 19, 1111–1119.
[https://doi.org/10.1089/jmf.2016.3800]

-
Horigome, S.; Yoshida, I.; Tsuda, A.; Harada, T.; Yamaguchi, A.; Yamazaki, K.; Inohana, S.; Isagawa, S.; Kibune, N.; Satoyama, T.; Katsuda, S.-I.; Suzuki, S.; Watai, M.; Hirose, N.; Mitsue, T.; Shirakawa, H.; Komai, M. Biosci. Biotechnol. Biochem. 2014, 78, 851–860.
[https://doi.org/10.1080/09168451.2014.905177]

-
Li, H.; Zhang, X.; Wang, W. Afr. J. Tradit. Complement. Altern. Med. 2017, 14, 213–220.
[https://doi.org/10.21010/ajtcam.v14i4.24]

-
Shammas, M. A.; Neri, P.; Koley, H.; Batchu, R. B.; Bertheau, R. C.; Munshi, V.; Prabhala, R.; Fulciniti, M.; Tai, Y. T.; Treon, S. P.; Goyal, R. K.; Anderson, K. C.; Munshi, N. C. Blood. 2006, 108, 2804–2810.
[https://doi.org/10.1182/blood-2006-05-022814]

-
Shanafelt, T. D.; Call, T. G.; Zent, C. S.; LaPlant, B.; Bowen, D. A.; Roos, M.; Secreto, C. R.; Ghosh, A. K.; Kabat, B. F.; Lee, M.-J.; Yang, C. S.; Jelinek, D. F.; Erlichman, C.; Kay, N. E. J. Clin. Oncol. 2009, 27, 3808–3814.
[https://doi.org/10.1200/JCO.2008.21.1284]

-
Bae, J.; Kumazoe, M.; Park, S.-J.; Fujimura, Y.; Tachibana, H. FEBS Open Bio 2023, 13, 2147–2156.
[https://doi.org/10.1002/2211-5463.13708]

-
Bae, J.; Kumazoe, M.; Yamashita, S.; Tachibana, H. Sci. Rep. 2017, 7, 6665.
[https://doi.org/10.1038/s41598-017-06879-5]

-
Mullen, T. D.; Hannun, Y. A.; Obeid, L. M. Biochem. J. 2012, 441, 789–802.
[https://doi.org/10.1042/BJ20111626]

-
Bae, J.; Kumazoe, M.; Takeuchi, C.; Hidaka, S.; Fujimura, Y.; Tachibana, H. Biochem. Biophys. Res. Commun. 2019, 520, 186–191.
[https://doi.org/10.1016/j.bbrc.2019.09.102]

-
Tsukamoto, S.; Huang, Y.; Kumazoe, M.; Lesnick, C.; Yamada, S.; Ueda, N.; Suzuki, T.; Yamashita, S.; Kim, Y. H.; Fujimura, Y.; Miura, D.; Kay, N. E.; Shanafelt, T. D.; Tachibana, H. Mol. Cancer Ther. 2015, 14, 2303–2312.
[https://doi.org/10.1158/1535-7163.MCT-15-0185]

-
Zhou, L.; Yang, F.; Li, G.; Huang, J.; Liu, Y.; Zhang, Q.; Tang, Q.; Hu, C.; Zhang, R. Front. Pharmacol. 2018, 9, 517.
[https://doi.org/10.3389/fphar.2018.00517]

-
Sudalagunta, P.; Silva, M. C.; Canevarolo, R. R.; Alugubelli, R. R.; DeAvila, G.; Tungesvik, A.; Perez, L.; Gatenby, R.; Gillies, R.; Baz, R.; Meads, M. B.; Shain, K. H.; Silva, A. S. EBioMedicine. 2020, 54, 102716.
[https://doi.org/10.1016/j.ebiom.2020.102716]
