
Dillapiole, a Phenylpropanoid from Piper aduncum and Dill Weed, Attenuates H2O2-Induced Muscle Atrophy in C2C12 Myotubes
Abstract
Skeletal muscle atrophy is closely associated with oxidative stress-induced inflammatory responses, which lead to the activation of catabolic pathways and the loss of myotube integrity. Dillapiole, a phenylpropanoid compound derived from several aromatic plants, has been reported to exhibit anti-inflammatory and antioxidant properties, suggesting its potential benefits in conditions characterized by muscle wasting. This study aimed to investigate whether dillapiole attenuates hydrogen peroxide (H2O2)-induced muscle atrophy in C2C12 myotubes and to elucidate its regulatory effects on key atrogenes. C2C12 myoblasts were differentiated into myotubes and then exposed to H2O2 to induce oxidative stress-mediated atrophy in the presence or absence of dillapiole. These findings indicate that dillapiole protects C2C12 myotubes against H2O2-induced muscle atrophy primarily by attenuating oxidative stress–associated inflammatory responses and suppressing the expression of Atrogin-1 and MuRF1.
Keywords:
Dillapiole, C2C12 myotubes, Hydrogen peroxide (H2O2), Skeletal muscle atrophy, Oxidative stressIntroduction
Skeletal muscle atrophy is characterized by an imbalance between protein synthesis and degradation, leading to a net loss of muscle mass and function.1–3 It commonly occurs under various conditions such as aging, prolonged immobilization, denervation, and chronic inflammation.4–7 Accumulating evidence has shown that reactive oxygen species (ROS) levels are markedly elevated in atrophic muscles, acting as critical mediators that trigger the expression of muscle-specific ubiquitin ligases such as MuRF1 and Atrogin-1.8,9 These molecules accelerate protein degradation and ultimately promote muscle atrophy. Consequently, strategies aimed at suppressing ROS generation and reducing pro-inflammatory cytokine expression through antioxidant or anti-inflammatory agents have gained increasing attention as potential therapeutic approaches.10,11
Dillapiole, found in plants such as giant peppermint, dill, and anise, has been reported to exhibit both anti-inflammatory and antioxidant properties.12,13 In particular, previous studies have shown that dillapiole or dillapiole-rich essential oils can attenuate inflammatory mediator production and scavenge reactive oxygen species in various cell types, suggesting that this compound may effectively modulate redox-sensitive signaling pathways.14,15 Given that oxidative stress and inflammation are tightly interconnected processes that synergistically drive proteolytic signaling and myofiber degeneration in skeletal muscle, targeting both ROS accumulation and pro-inflammatory cytokine expression represents a rational strategy for preventing or attenuating muscle atrophy.10,16 Although several natural antioxidants and plant-derived compounds have been investigated for their capacity to mitigate oxidative stress–induced muscle damage, the impact of dillapiole on skeletal muscle cells remains poorly characterized, and its potential to counteract oxidative stress-driven atrophic signaling has not been clarified. In particular, it remains unclear whether dillapiole can attenuate muscle atrophy by suppressing ROS accumulation and downstream catabolic signaling pathways, including the activation of muscle-specific E3 ubiquitin ligases such as MuRF1 and Atrogin-1. Therefore, the present study aimed to elucidate whether dillapiole exerts anti-atrophic and anti-inflammatory effects by modulating ROS production, inflammatory responses, and atrogene expression under oxidative stress-induced muscle atrophy conditions.
Experimental
Cell culture and differentiation – C2C12 myoblasts (mouse skeletal muscle myoblast cell line, CRL-1772TM) were purchased from the American Type Culture Collection (ATCC, Manassas, VA, USA). C2C12 cultures were expanded in Dulbecco modified Eagle medium (Welgene, Gyeongsan, South Korea) supplemented with 10% (v/v) fetal bovine serum (FBS; Gibco, Thermo Fisher Scientific, Waltham, MA, USA) and 1% Penicillin-Streptomycin (Welgene, Gyeongsan, South Korea) at 5% CO2 and 37℃. For C2C12 differentiation, 5 × 104 cells were seeded in six-well plates and cultured in growth media until reaching 70–80% confluence (day 0). Media were then replaced with Dulbecco modified Eagle medium supplemented with 2% (v/v) horse serum (differentiation medium, Gibco, Thermo Fisher Scientific, Waltham, MA, USA). Cells were kept in differentiation medium until the end of the assay, typically between 4 and 7 days.
Dillapiole treatment – Due to its poor water solubility, a 50 mM stock solution was prepared by dissolving dillapiole in absolute ethanol and stored at −20°C protected from light. Working concentrations (6.25, 12.5, and 25 μM) were freshly diluted in differentiation medium immediately prior to cell treatment. For co-treatment experiments, differentiated C2C12 myotubes were preincubated with dillapiole for 2 h, followed by co-exposure to H2O2 (hydrogen peroxide, Sigma-Aldrich, St. Louis, MO, USA) for 24 h to induce oxidative stress-mediated atrophy.
Western blotting – For protein immunoblot analysis, polypeptides in whole cell lysates were resolved by SDS-PAGE and transferred to PVDF membrane filters (Bio-Rad, Hercules, CA, USA). Proteins were detected with a 1:100 or 1:5000 dilution of primary antibody using an enhanced chemiluminescence (ECL Solution, Dogen, Seoul) system. The images were acquired using AmershamTM ImageQuantTMTM 800 Western blot imaging systems (Cytiva, Marlborough, MA). The MuRF1 antibody was purchased from Santacruz (sc-398608), the Atrogin-1/MAFbx antibody from Santacruz (sc-166806), and the Beta-Actin antibody from Abm (G043). Band intensities were quantified using ImageJ and normalized to Beta-Actin (n = 3 independent experiments).
Quantitative RT-PCR – Cells were harvested and RNA was extracted using Trizol (Invitrogen, Carlsbad, CA, USA) in accordance with the manufacturer’s instructions, and subjected to reverse transcription-PCR (RT-PCR) using CFX Connect Real-Time PCR Detection System (Bio-Rad, Hercules, CA, USA). The mMuRF1 (mouse origin MuRF1) mRNA was amplified using the forward primer 5′-GGAGAAGCTGGACTTCATCG-3′ and reverse primer 5′-CTTGGCACTCAAGAGGAAGG-3′. The mAtrogin-1(mouse origin Atrogin-1/MAFbx) mRNA was amplified using the forward primer 5′-GAA CAT CAT GCA GAG GCT GA-3′ and reverse primer 5′-GTA GCC GGT CTT CAC TGA GC-3′. The input RNA was normalized via the amplification of mGAPDH (mouse origin GAPDH) RNA with the forward primer 5′-ACA TCA TCC CTG CAT CCA CT-3′ and reverse primer 5′-CAC ATT GGG GGT AGG AAC AC-3′.
ROS detection assay – C2C12 myoblast cells were seeded in 48-well plates at a cell density of 3 × 104 cells/well and differentiated. The ROS levels were measured using the 2′,7′-dichlorofluorescein diacetate (DCFDA) method (Catalog # ab113851, Abcam, MA, USA). The media from the differentiated cells were removed and washed with 1X buffer solution. Next, the cells were stained with 100 µl of diluted DCFDA (20 µM) for 45 min at 37°C in the dark. Then the cells were washed three times with 1X buffer solution to remove the DCFDA solution. The cells were then treated with 1X supplemented buffer for 24 h. The fluorescence intensity of the cells was measured using a fluorescence spectrophotometer, with excitation and emission wavelengths of 485 nm and 535 nm, respectively, following the H2O2 incubation period.
Statistical analysis – All experiments were performed in at least three independent experiments, with each experiment conducted in triplicate (n = 3). Data are presented as mean ± standard deviation (SD) from three biological replicates. Statistical significance was determined by a 2-tailed Student’s t-test using GraphPad Prism software (GraphPad, La Jolla, CA).
Results and Discussion
Dillapiole is a hydrophobic phenylpropanoid, and its chemical structure is shown in Fig. 1A. To treat cells with dillapiole, a stock solution was prepared in DMSO at a concentration of 25.0 mg/mL. First, the cytotoxicity of dillapiole was evaluated by measuring cell viability using the MTT assay. C2C12 cells were treated with dillapiole at 0, 6.25, 12.5, 25, and 50 μM, and a marked decrease in cell viability was observed at 50 μM. Based on these results, subsequent experiments were performed using dillapiole at concentrations up to 25 μM (Fig. 1B).
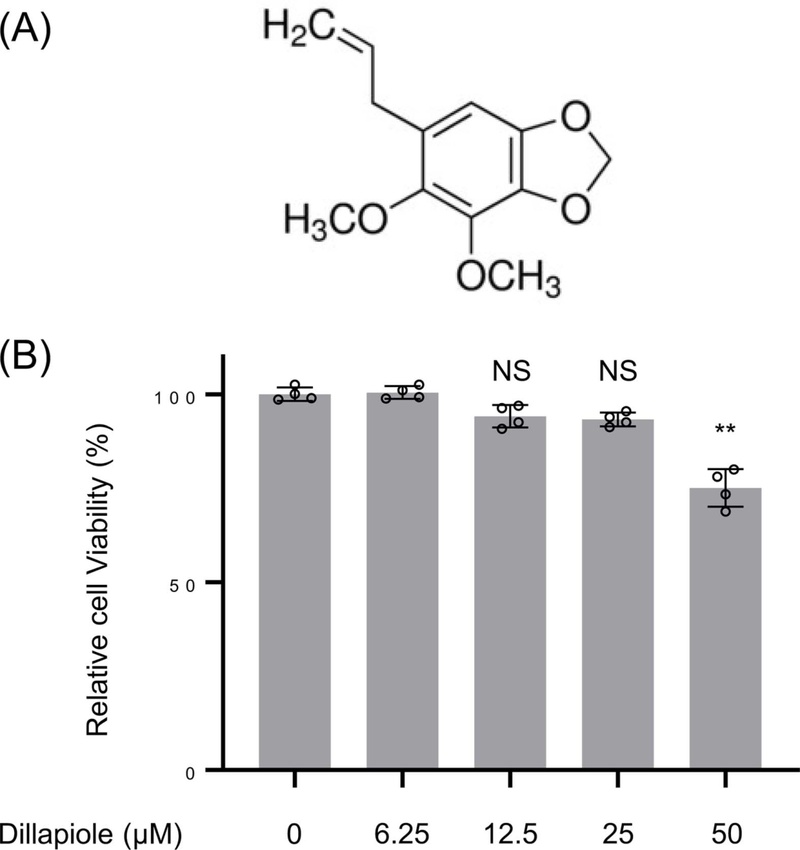
Structure of dillapiole and cell viability of C2C12 myotube cells under dillapiole. (A) Dillapiole is an alkenyl benzene compound featuring a 1,3-benzodioxole ring substituted with two methoxy groups and an allyl group. (B) Effects of dillapiole on the viability of C2C12 cells. C2C12 cells were treated with increasing concentrations of dillapiole (0–50 μM), and cell viability was evaluated using an MTT assay. NS; Not Significant, **p < 0.01 vs. control.
Dillapiole has been suggested as a potential anti-inflammatory agent, and thus it was hypothesized that it could exert anti-inflammatory effects in skeletal muscle cells. ROS levels were measured using DCFDA under muscle atrophy-inducing conditions (100 μM H2O2, 24 h). H2O2 treatment markedly increased intracellular ROS levels compared with the control group, whereas dose-dependent treatment with dillapiole significantly reduced ROS accumulation (Fig. 2). These findings indicate that dillapiole attenuates oxidative stress induced by ROS in C2C12 myotubes.
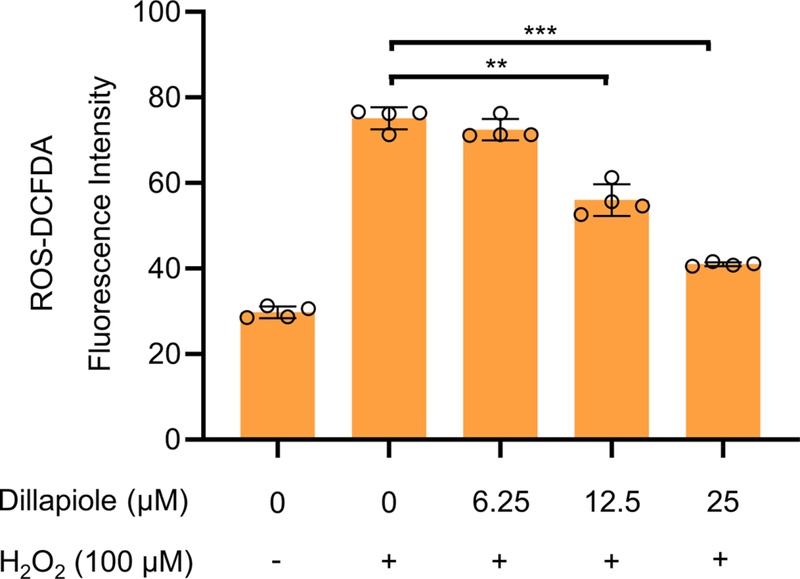
Dillapiole attenuates H2O2-induced reactive oxygen species (ROS) production in C2C12 myotubes. C2C12 myotubes were treated with various concentrations of dillapiole (0–25 μM) in the absence or presence of H2O2 (100 μM). To measure H2O2-induced ROS production, the cells were then labeled with 2′,7′-dichlorodihydrofluorescein diacetate (H2DCFDA). Quantitative analysis was performed by measuring the fluorescence intensity. **p < 0.01, ***p < 0.005 vs. control.
To determine whether dillapiole also affects muscle atrophy, the protein expression of the muscle atrophy markers MuRF1 and Atrogin-1 was analyzed by Western blotting (Fig. 3A). Densitometric quantification using ImageJ revealed that dillapiole treatment dose-dependently decreased H2O2-induced MuRF1 and Atrogin-1 protein levels (Fig. 3B).
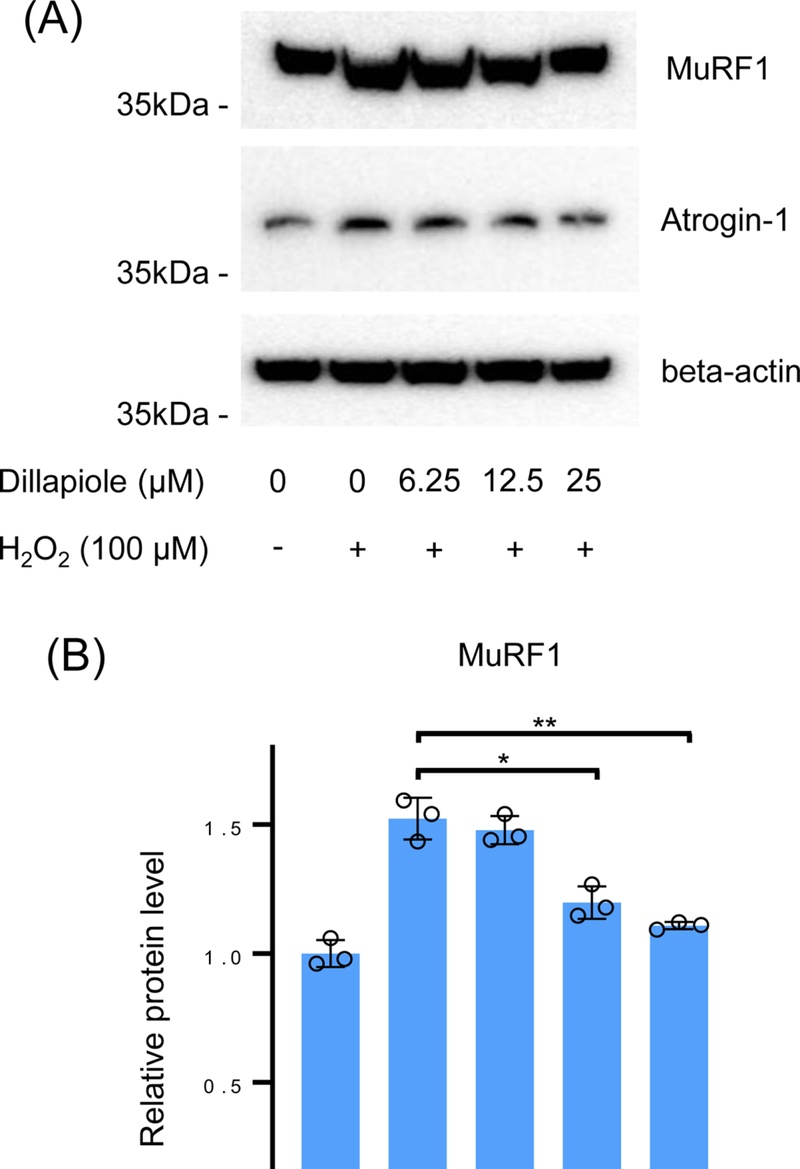
Protein expression levels of MuRF1 and Atrogin-1 are downregulated by dose-dependent treatment with dillapiole in C2C12 myotubes. (A) Representative western blot images showing the protein expression levels of MuRF1 and Atrogin-1 in C2C12 myotubes. (B) The level of each protein in panel. The means and standard deviations are shown. Control vs. Dose-dependent treatment with dillapiole: *p < 0.05, **p < 0.01, ***p < 0.005.
In addition, the mRNA expression of MuRF1 and Atrogin-1 was examined by quantitative real-time PCR. Consistent with the protein data, dillapiole treatment reduced the H2O2-induced upregulation of MuRF1 and Atrogin-1 mRNA levels (Fig. 4).
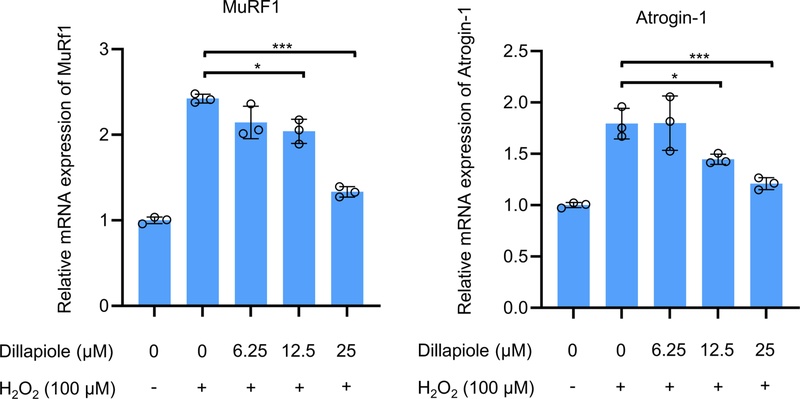
Dillapiole reduces MuRF1 and Atrogin-1 mRNA expression in C2C12 myotubes. mRNA expression levels of MuRF1 and Atrogin-1 were measured using qRT-PCR in C2C12 myotubes. The means and standard deviations are shown. Control vs. Dose-dependent treatment with dillapiole: *p < 0.05, ***p <0.005.
Finally, to assess the functional recovery from muscle atrophy, H2O2-induced atrophy was evaluated by measuring C2C12 myotube diameter. H2O2 treatment resulted in a significant reduction in myotube diameter, whereas co-treatment with dillapiole restored myotube thickness (Fig. 5A). Quantitative analysis showed that myotube diameter was significantly recovered at 12.5 and 25 μM dillapiole (Fig. 5B).
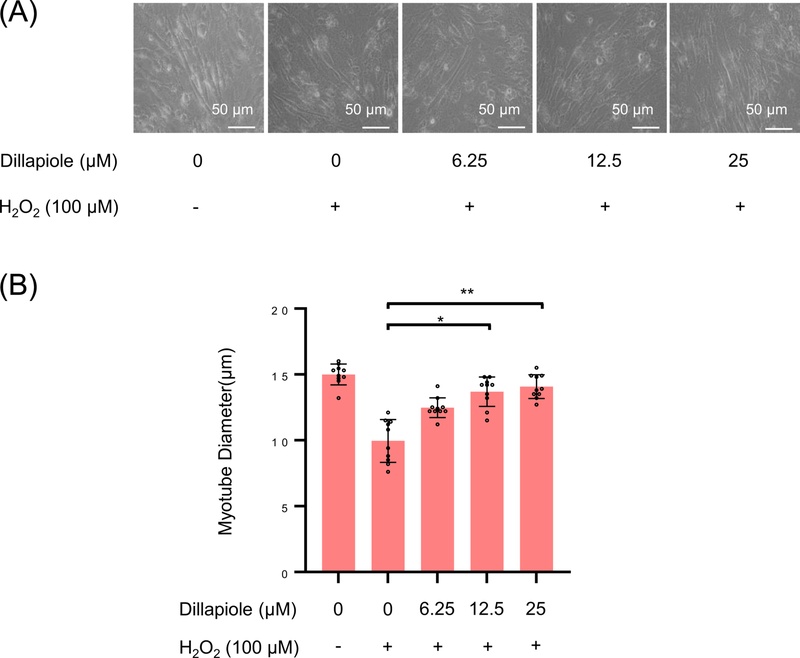
Dillapiole restores myotube diameter in C2C12 cells under atrophic conditions. (A) C2C12 myotubes were exposed to H2O2 (100 µM) in the absence or presence of increasing concentrations of dillapiole (0–25 μM). Representative microscopic images show that dillapiole treatment visibly preserved myotube morphology and thickness compared with atrophy-induced cells. (B) Myotube diameter was quantified from randomly selected fields and expressed as the mean diameter for each treatment group. Control vs. Dose-dependent treatment with dillapiole: *p < 0.05, **p < 0.01.
In this study, dillapiole was identified as a potential protective agent against oxidative stress–induced skeletal muscle atrophy in C2C12 myotubes. Within a non-cytotoxic concentration range, dillapiole reduced H2O2-induced ROS accumulation, suppressed the protein and mRNA expression of the atrophy-related E3 ubiquitin ligases MuRF1 and Atrogin-1, and ultimately preserved myotube diameter under atrophic conditions. These findings indicate that dillapiole exerts both antioxidative and anti-atrophic effects in skeletal muscle cells and may serve as a promising candidate for the development of interventions targeting muscle wasting.
In the initial cytotoxicity assessment, dillapiole concentrations up to 25 μM did not significantly reduce C2C12 cell viability, whereas a concentration of 50 μM produced marked cytotoxic effects. This allowed the establishment of an effective concentration range in which pharmacological activities could be evaluated without interference from toxicity. Defining this therapeutic window is essential for subsequent studies aiming to apply dillapiole as a safe bioactive compound for skeletal muscle protection.
One of the key observations of this study is that dillapiole strongly inhibited H2O2-induced ROS generation. Because excessive ROS production is known to promote muscle atrophy through direct macromolecular damage and activation of proteolytic signaling pathways, the reduction in intracellular ROS levels provides a mechanistic basis for the observed changes in atrophy markers. The results demonstrate that dillapiole can alleviate oxidative stress in myotubes, in agreement with previously reported antioxidant effects in other cell types. Although the precise molecular targets responsible for this antioxidant activity have not yet been identified, possible mechanisms include enhancement of endogenous antioxidant defenses or direct scavenging of ROS.
Consistent with this ROS-lowering effect, dillapiole treatment significantly attenuated the H2O2-induced upregulation of MuRF1 and Atrogin-1, two muscle-specific E3 ubiquitin ligases that play central roles in proteasome-mediated protein degradation during muscle atrophy. The concordant reductions at both the protein and mRNA levels suggest that dillapiole regulates these genes at the transcriptional level or further upstream. Overall, the present findings provide novel evidence that dillapiole can mitigate oxidative stress-induced skeletal muscle atrophy by decreasing ROS levels, suppressing MuRF1 and Atrogin-1 expression, and preserving myotube morphology. These data highlight dillapiole as a promising plant-derived candidate for the prevention or attenuation of muscle wasting associated with oxidative and inflammatory conditions, and underscore the need for further mechanistic and in vivo studies to fully establish its therapeutic potential in sarcopenia and other muscle atrophy-related disorders.
Acknowledgments
This work was supported by Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (RS2023-00239974) and This study was supported by Yong In University’s 2023 academic research grant.
Conflicts of Interest
The authors declare that they have no conflicts of interest.
References
-
Chen, P.; Jia, F.; Wang, M.; Yang, S. Front Physiol. 2025, 16, 1533394.
[https://doi.org/10.3389/fphys.2025.1533394]

-
Fanzani, A.; Conraads, V. M.; Penna, F.; Martinet, W. J. Cachexia Sarcopenia Muscle 2012, 3, 163–179.
[https://doi.org/10.1007/s13539-012-0074-6]

-
Sartori, R.; Romanello, V.; Sandri, M. Nat. Commun. 2021, 12, 330.
[https://doi.org/10.1038/s41467-020-20123-1]

-
Bodine, S. C.; Sinha, I.; Sweeney, H. L. J. Gerontol. A Biol. Sci. Med. Sci. 2023, 78, 14–18.
[https://doi.org/10.1093/gerona/glad023]

-
Ji, Y.; Li, M.; Chang, M.; Liu, R.; Qiu, J.; Wang, K.; Deng, C.; Shen, Y.; Zhu, J.; Wang, W.; Xu, L.; Sun, H. Antioxidants 2022, 11, 1686.
[https://doi.org/10.3390/antiox11091686]

-
Li, H.; Malhotra, S.; Kumar, A. J. Mol. Med. 2008, 86, 1113–1126.
[https://doi.org/10.1007/s00109-008-0373-8]

-
Thoma, A.; Lightfoot, A. P. Adv. Exp. Med. Biol. 2018, 1088, 267–279.
[https://doi.org/10.1007/978-981-13-1435-3_12]

-
Pomiès, P.; Blaquière, M.; Maury, J.; Mercier, J.; Gouzi, F.; Hayot, M. PLoS One 2016, 11, e0160092.
[https://doi.org/10.1371/journal.pone.0160092]

-
Powers, S. K.; Morton, A. B.; Ahn, B.; Smuder, A. J. Free Radic. Biol. Med. 2016, 98, 208–217.
[https://doi.org/10.1016/j.freeradbiomed.2016.02.021]

- Agrawal, S.; Chakole, S.; Shetty, N.; Prasad, R.; Lohakare, T.; Wanjari, M. Cureus 2023, 15, e42178.
-
Xiao, Q.; Sun, C.-C.; Tang, C.-F. Exp. Gerontol. 2023, 184, 112335.
[https://doi.org/10.1016/j.exger.2023.112335]

-
Almeida, R. R. P.; Souto, R. N. P.; Bastos, C. N.; Silva, M. H. L.; Maia, J. G. S. Chem. Biodivers. 2009, 6, 1427–1434.
[https://doi.org/10.1002/cbdv.200800212]

-
Parise-Filho, R.; Pasqualoto, K. F. M.; Magri, F. M. M.; Ferreira, A. K.; Silva, B. A. V. G.; Damião, M. C. F. C. B.; Tavares, M. T.; Azevedo, R. A.; Auada, A. V. V.; Polli, M. C.; Brandt, C. A. Arch. Pharm. 2012, 345, 934–944.
[https://doi.org/10.1002/ardp.201200212]

-
Parise-Filho, R.; Pastrello, M.; Camerlingo, C. E. P.; Silva, G. J.; Agostinho, L. A.; Souza, T.; Magri, F. M. M.; Ribeiro, R. R.; Brandt, C. A.; Polli, M. C. Pharm. Biol. 2011, 49, 1173–1179.
[https://doi.org/10.3109/13880209.2011.575793]

-
Fatima, A.; Ayub, M. A.; Choobkar, N.; Zubair, M.; Thompson, K. D.; Hussain, A. Food Sci. Nutr. 2025, 13, e70089.
[https://doi.org/10.1002/fsn3.70089]

-
Zhang, H.; Qi, G.; Wang, K.; Yang, J.; Shen, Y.; Yang, X.; Chen, X.; Yao, X.; Gu, X.; Qi, L.; Zhou, C.; Sun, H. Biochem. Pharmacol. 2023, 214, 115664.
[https://doi.org/10.1016/j.bcp.2023.115664]
