
Online DPPH Assay Coupled with LC-QTOF-MS for Rapid Identification of Antioxidant Constituents from Zelkova serrata
Abstract
Zelkova serrata (Thunb.) Makino is a deciduous tree commonly distributed in East Asia. Several investigations have documented the biological activity of extracts of this species, igniting increased interest in its therapeutic potential. However, the phytochemical constituents accountable for these activities remain largely unexplored. In this study, we focused on the antioxidant potential of the leaves and twigs, utilizing bioactivity-guided fractionation combined with LC-QTOF-MS online DPPH screening to systematically identify the active compounds. This approach led to the isolation of eight compounds, three of which (compounds 1, 3, and 7) have not previously been documented in the genus Zelkova. Among the isolates, 4 emerged as the most potent antioxidant, exhibiting significant radical scavenging activity in both DPPH (IC50 = 13.67 ± 1.21 μM) and ABTS (IC50 = 2.69 ± 0.48 μM) assays, exceeding ascorbic acid (IC50 = 61.51 ± 10.22 μM) and trolox (IC50 = 8.22 ± 2.16 μM), respectively. Our findings not only enrich the phytochemical profile of Z. serrata but also highlight the effectiveness of LC-MS-DPPH as an effective approach for rapid antioxidant discovery.
Keywords:
Zelkova serrata, Bioassay-guided isolation, Antioxidant, DPPH, ABTSIntroduction
Oxidative stress represents a significant contributor to numerous pathological conditions, including malignancies, neurodegenerative disorders, and age-associated diseases. The excessive generation of reactive oxygen species (ROS), encompassing free radicals, hydroxyl radicals, hydrogen peroxide, singlet oxygen, and superoxide anions, disrupts cellular homeostasis by damaging critical biomolecules such as proteins, lipids, and nucleic acids.1 These oxidative modifications are implicated in the pathogenesis of Alzheimer’s disease, Parkinson’s disease, diabetes mellitus, and various inflammatory conditions. While endogenous antioxidant defense systems exist to maintain ROS equilibrium, exogenous supplementation becomes imperative when oxidative burden exceeds physiological capacity.2
Although synthetic antioxidants are commercially employed in food preservation and pharmaceutical applications, concerns regarding their potential hepatotoxicity and carcinogenic properties have prompted increased interest in natural alternatives.3,4 Plant-derived polyphenolic compounds, including flavonoids and phenolic acids, have emerged as promising candidates due to their multifaceted mechanisms of action as hydrogen donors, metal chelators, and free radical scavengers.5 Beyond their antioxidant capacity, these phytochemicals demonstrate anti-inflammatory, antiproliferative, antidiabetic, and neuroprotective activities, underscoring their therapeutic potential.3,6,7 Consequently, systematic investigation of plants represents a rational approach for discovering novel natural antioxidants with enhanced safety profiles.
Within this context, Zelkova serrata (Thunb.) Makino, a deciduous broadleaf species belonging to the family Ulmaceae, presents particular interest as a natural resource. This species exhibits widespread distribution across East Asia, including China, Korea, Japan, and Taiwan. The species holds considerable economic and cultural significance, with its timber valued for construction applications, particularly in traditional temple and shrine architecture, as well as high-quality furniture manufacturing. Its adaptability to moderately saline soils has also identified Z. serrata as a candidate for coastal afforestation initiatives.3 Besides, various parts of Z. serrata have been traditionally utilized for both culinary and medicinal purposes. The tender foliage is consumed as a culinary vegetable following preparation, while both bark and leaf tissues serve as medicinal materials in folk remedies.8,9 Notably, extracts derived from the entire plant have found extensive application in anticancer therapy. Extracts from leaves have demonstrated antipyretic and detoxification activities, while twig preparations show antiproliferative and larvicidal effects.3,10 The bark, traditionally employed for its antipyretic, diuretic, and detoxifying properties, has been reported to possess antioxidant capacity; however, the specific chemical entities responsible for these activities remain inadequately characterized.11
Given the established antioxidant properties of Z. serrata tissues and the current emphasis on natural antioxidant discovery, the present investigation aims to identify and characterize the bioactive compounds from Z. serrata leaves and twigs and evaluate their antioxidant potential.
Experimental
General experimental procedures – A Bruker Avance III 600 MHz (Karlsruhe, Germany) spectrometer was used to acquire the NMR spectra. Mass spectral analysis was performed on an Agilent 1260 system coupled with an Agilent 6530 Q-TOF mass spectrometer (Santa Clara, USA). Thin-layer chromatography (TLC) was performed using RP-18 F254s and Merck precoated silica gel F254 plates. Preparative HPLC was carried out using a YL9101 Plus HPLC system with a Phenomenex Luna C18(2) column (250 × 10 mm, 5 μm). Open column chromatographic separation was achieved using Merck (RP)-C18 (75 mesh) for reverse-phase or 230–400 mesh silica gel 60 for the normal phase.
Plant material – The leaves, twigs, and roots of Z. serrata were obtained from the Medicinal Herb Garden, College of Pharmacy, Seoul National University, Seoul, Korea, and identified by Professor Ki Yong Lee, Korea University. The voucher specimens (KUP-HD133 and KUP-HD134) were deposited at the Herbarium of Natural Product Laboratory, College of Pharmacy, Korea University.
Extraction and isolation – The dried leaves and twigs of Z. serrata (3.0 kg) were chopped and extracted three times with 80% MeOH via sonication (60 min each). The combined extracts were filtered and concentrated using a vacuum rotary evaporator to obtain the MeOH crude extract, which was suspended in distilled water, and partitioned with n-hexane, ethyl acetate (EtOAc), and n-butanol (n-BuOH), respectively. The EtOAc extract (19.2 g) was subjected to silica gel CC eluted with n-hexane:EtOAc (5:1 → 1:1, v/v) and CH2Cl2:MeOH (10:1 → 1:1, v/v) to obtain 10 fractions (E1–E10). E6 (2.3 g) was separated on a silica gel column using n-hexane:acetone (2:1 → 1:5, v/v) to obtain 14 subfractions (E6.1−E6.14). E6.13 was purified by preparative HPLC (MeCN:H2O, 35:65 → 50:50, v/v) to yield 5 (2.0 mg, tR = 15.3 min) and 6 (1.2 mg, tR = 20.2 min). E8 (4.8 g) was chromatographed over an open column eluting with CH2Cl2:MeOH:H2O (90:20:1 → 30:10:1, v/v) to yield 19 subfractions (E8.1−E8.19). E8.6 (560.0 mg) was chromatographed over an RP-C18 column eluting with MeOH:H2O (3:1 → 1:1) to obtain 7 (27.5 mg) and 8 (16.3 mg). E8.9 (1.3 g) was purified by preparative HPLC (MeCN:H2O, 40:60 → 90:10, v/v) to yield 1 (550.3 mg, tR = 33.0 min) and 3 (430.0 mg, tR = 33.9 min). E9 (2.1 g) was subjected to RP-C18 eluted with MeOH:H2O (50:50 → 60:40) to afford 2 (2.3 mg) and 4 (15.8 mg).
Catechin-7-O-β-D-xylopyranoside (1) − ESI-MS m/z 421.1142 [M−H]− (calcd for C20H21O10, 421.1135); 1H-NMR (600 MHz, CD3OD): δ 6.84 (1H, d, J = 2.0 Hz, H-2′), 6.77 (1H, d, J = 8.1 Hz, H-5′), 6.73 (1H, dd, J = 8.1, 2.0 Hz, H-6′), 6.18 (1H, d, J = 2.3 Hz, H-6), 6.12 (1H, d, J = 2.3 Hz, H-8), 4.77 (1H, d, J = 7.2 Hz, H-1′′), 4.61 (1H, d, J = 7.4 Hz, H-2), 4.01 (1H, td, J = 7.4, 5.4 Hz, H-3), 3.31–3.91 (5H, overlap, H-2′′, H-3′′, H-4′′, H-5′′), 2.87 (1H, dd, J = 16.3, 5.4 Hz, H-4a), 2.55 (1H, dd, J = 16.3, 8.0 Hz, H-4b); 13C-NMR (150 MHz, CD3OD): δ 157.1 (C-7), 156.2 (C-5), 155.4 (C-9), 144.9 (C-3′, C-4′), 130.7 (C-1′), 118.6 (C-6′), 114.7 (C-5′), 113.8 (C-2′), 102.3 (C-10), 101.5 (C-1′′), 96.0 (C-6), 95.6 (C-8), 81.5 (C-2), 76.4 (C-3′′), 73.3 (C-2′′), 69.7 (C-4′′), 67.2 (C-3), 65.5 (C-5′′), 27.1 (C-4).
Aromadendrin-6-C-β-D-glucopyranoside (2) − ESI-MS m/z 449.1092 [M−H]− (calcd for C21H21O11, 449.1084); 1H-NMR (600 MHz, CD3OD): δ 7.35 (2H, d, J = 8.6 Hz, H-2′, H-6′), 6.84 (2H, d, J = 8.6 Hz, H-3′, H-5′), 5.96 (1H, s, H-8), 5.00 (1H, d, J = 11.5 Hz, H-2), 4.81 (1H, d, J = 9.9 Hz, H-1′′), 4.56 (1H, d, J = 11.5 Hz, H-3), 4.13 (1H, m, H-2′′), 3.87 (1H, m, H-6′′a), 3.72 (1H, m, H-6′′b), 3.46 (1H, m, H-4′′), 3.45 (1H, m, H-3′′), 3.40 (1H, m, H-5′′); 13C-NMR (150 MHz, CD3OD): δ 198.9 (C-4), 167.6 (C-7), 164.1 (C-5), 163.9 (C-9), 159.2 (C-4′), 130.3 (C-2′, C-6′), 129.1 (C-1′), 116.2 (C-3′, C-5′), 106.3 (C-6), 101.7 (C-10), 96.4 (C-8), 85.0 (C-2), 82.5 (C-5′′), 80.2 (C-3′′), 75.1 (C-1′′), 73.6 (C-3), 72.6 (C-2′′), 71.8 (C-4′′), 62.9 (C-6′′).
Catechin-7-O-β-D-apiofuranoside (3) − ESI-MS m/z 421.1139 [M−H]− (calcd for C20H21O10, 421.1135); 1H-NMR (600 MHz, CD3OD): δ 6.84 (1H, d, J = 2.0 Hz, H-2′), 6.77 (1H, d, J = 8.1 Hz, H-5′), 6.72 (1H, dd, J = 8.1, 2.0 Hz, H-6′), 6.14 (1H, d, J = 2.3 Hz, H-6), 6.09 (1H, d, J = 2.3 Hz, H-8), 5.49 (1H, d, J = 3.0 Hz, H-1′′), 4.61 (1H, d, J = 7.4 Hz, H-2), 4.14 (1H, d, J = 3.0 Hz, H-2′′), 4.09 (1H, d, J = 9.7 Hz, H-4′′a), 4.00 (1H, td, J = 7.4, 5.4 Hz, H-3), 3.85 (1H, d, J = 9.7 Hz, H-4′′b), 3.64 (1H, d, J = 11.5 Hz, H-5′′a), 3.61 (1H, d, J = 11.5 Hz, H-5′′b), 2.86 (1H, dd, J = 16.3, 5.4 Hz, H-4a), 2.55 (1H, dd, J = 16.3, 8.0 Hz, H-4b); 13C-NMR (150 MHz, CD3OD): δ 156.8 (C-7), 156.2 (C-5), 155.5 (C-9), 144.9 (C-3′), 144.8 (C-4′), 130.7 (C-1′), 118.6 (C-6′), 114.7 (C-5′), 113.8 (C-2′), 107.3 (C-1′′), 101.8 (C-10), 95.9 (C-6), 95.4 (C-8), 81.5 (C-2), 78.9 (C-3′′), 76.9 (C-2′′), 74.0 (C-4′′), 67.2 (C-3), 63.5 (C-5′′), 27.0 (C-4).
Catechin (4) − ESI-MS m/z 289.0718 [M−H]− (calcd for C15H13O6, 289.0712); 1H-NMR (600 MHz, CD3OD): δ 6.84 (1H, d, J = 2.0 Hz, H-2'), 6.76 (1H, d, J = 8.1 Hz, H-5′), 6.72 (1H, dd, J = 8.1, 2.0 Hz, H-6′), 5.92 (1H, d, J = 2.3 Hz, H-6), 5.85 (1H, d, J = 2.3 Hz, H-8), 4.56 (1H, d, J = 7.5 Hz, H-2), 3.97 (1H, td, J = 7.5, 5.4 Hz, H-3), 2.85 (1H, dd, J = 16.1, 5.4 Hz, H-4a), 2.50 (1H, dd, J = 16.1, 8.2 Hz, H-4b); 13C-NMR (150 MHz, CD3OD): δ 157.8 (C-7), 157.6 (C-9), 156.9 (C-5), 146.3 (C-3′), 146.2 (C-4′), 132.2 (C-1′), 120.0 (C-6′), 116.1 (C-5′), 115.3 (C-2′), 100.8 (C-10), 96.3 (C-6), 95.5 (C-8), 82.9 (C-2), 68.8 (C-3), 28.5 (C-4).
4-Hydroxybenzoic acid (5) − ESI-MS m/z 137.0238 [M−H]− (calcd for C7H5O3, 137.0239); 1H-NMR (600 MHz, CD3OD): δ 7.87 (2H, d, J = 8.2 Hz, H-2, H-6), 6.80 (2H, d, J = 8.2 Hz, H-3, H-5); 13C-NMR (150 MHz, CD3OD): δ 169.7 (C-7), 162.7 (C-4), 132.8 (C-2, C-6), 122.9 (C-1), 115.8 (C-3, C-5).
Vanillic acid (6) − ESI-MS m/z 167.0355 [M−H]− (calcd for C8H7O4, 167.0344); 1H-NMR (600 MHz, CD3OD): δ 7.56 (1H, d, J = 1.9 Hz, H-2), 7.54 (1H, dd, J = 8.2, 1.9 Hz, H-6), 6.82 (1H, d, J = 8.2 Hz, H-5), 3.89 (3H, s, OCH3); 13C-NMR (150 MHz, CD3OD): δ 170.1 (C-7), 152.6 (C-3), 148.6 (C-4), 125.2 (C-1), 123.2 (C-6), 115.8 (C-2), 113.8 (C-5), 56.4 (OCH3).
Hemiphloin (7) − ESI-MS m/z 433.1148 [M−H]− (calcd for C21H21O10, 433.1135); 1H-NMR (600 MHz, CD3OD): δ 7.31 (2H, d, J = 8.6 Hz, H-2′, H-6′), 6.82 (2H, d, J = 8.6 Hz, H-3′, H-5′), 5.97 (1H, s, H-8), 5.34 (1H, dd, J = 12.7, 3.0 Hz, H-2), 4.80 (1H, d, J = 9.9 Hz, H-1′′), 4.13 (1H, m, H-2′′), 3.87 (1H, dd, J = 12.0, 2.5 Hz, H-6′′a), 3.72 (1H, dd, J = 12.0, 6.0 Hz, H-6′′b), 3.45 (2H, m, H-3′′, H-4′′), 3.39 (1H, td, J = 6.5, 6.0, 2.5 Hz, H-5′′), 3.12 (1H, dd, J = 17.1, 12.7 Hz, H-3a), 2.73 (1H, dd, J = 17.1, 3.0 Hz, H-3b); 13C-NMR (150 MHz, CD3OD): δ 198.1 (C-4), 167.2 (C-7), 164.2 (C-5, C-9), 159.0 (C-4′), 130.9 (C-1′), 129.0 (C-2′, C-6′), 116.3 (C-3', C-5'), 105.9 (C-6), 103.2 (C-10), 96.4 (C-8), 82.5 (C-5′′), 80.4 (C-3′′), 80.2 (C-2), 75.2 (C-1′′), 72.6 (C-2′′), 71.8 (C-4′′), 62.9 (C-6′′), 43.9 (C-3).
Isoquercetin (8) − ESI-MS m/z 463.0892 [M−H]− (calcd for C21H19O12, 463.0877); 1H-NMR (600 MHz, CD3OD): δ 7.72 (1H, d, J = 2.2 Hz, H-2′), 7.59 (1H, d, J = 8.5 Hz, H-6′), 6.88 (1H, d, J = 8.5 Hz, H-5′), 6.39 (1H, d, J = 2.5 Hz, H-8), 6.20 (1H, d, J = 2.5 Hz, H-6), 5.26 (1H, d, J = 7.7 Hz, H-1′′), 3.73 (1H, m, H-6′′a), 3.59 (1H, m, H-6′′b), 3.50 (1H, m, H-2′′), 3.45 (1H, m, H-3′′), 3.37 (1H, m, H-4′′), 3.24 (1H, m, H-5′′), 13C-NMR (150 MHz, CD3OD): δ 179.6 (C-4), 166.1 (C-7), 163.2 (C-5), 159.2 (C-9), 158.6 (C-2), 150.0 (C-3′), 146.0 (C-4′), 135.8 (C-3), 123.3 (C-1′), 123.2 (C-6′), 117.7 (C-5′), 116.1 (C-2′), 105.8 (C-10), 104.5 (C-1′′), 100.0 (C-6), 94.9 (C-8), 78.5 (C-5′′), 78.2 (C-3′′), 75.9 (C-2′′), 71.3 (C-4′′), 62.7 (C-6′′).
LC-QTOF-MS conditions − LC-QTOF-MS was conducted according to previously described methodologies.12 Chromatographic analysis utilized an Agilent 1260 HPLC system equipped with a C18 analytical column (Shiseido CapCell PAK, 150 × 4.6 mm i.d., 5 μm). Gradient elution was performed using 0.1% aqueous formic acid (eluent A) and acetonitrile (eluent B) as follows: 5% B for 5 min, followed by a linear increase to 95% B over 25 min, at a flow rate of 0.6 mL/min. Chromatograms were recorded at 280 nm. Mass spectrometry was carried out using an Agilent 6530 quadrupole time-of-flight instrument (Santa Clara, CA, USA) operated in negative electrospray ionization mode, acquiring data across m/z 50–1700.
DPPH radical scavenging assay − The antioxidant capacity was evaluated using the DPPH radical scavenging assay following the method previously described with modifications.13,14 For the assay, LC-MS-derived fractions were reconstituted in ethanol (300 μL total volume). Each sample (10 μL) was combined with 190 μL of DPPH reagent (15 μM in ethanol) and incubated for 30 min under ambient conditions. Spectrophotometric measurement at 517 nm quantified the extent of radical neutralization, with decreased absorbance indicating greater antioxidant potential. The inhibition rate (%), which represents inhibitory activity, was determined using the following formula:
where S0 and C0 are the absorbances of the control and inhibitor in ethanol without DPPH solution, and S and C are the absorbances of the control and inhibitor after 30 min.
ABTS radical scavenging assay − ABTS•+ radical cation was produced by reacting ABTS with potassium persulfate following the described protocol with modification.15 Before usage, this stock solution is diluted with distilled water to achieve an optical density between 0.7 and 0.8. Next, aliquot 20 µL of the sample into a well plate, followed by the addition of 180 µL of the diluted ABTS working solution. After a 7-minute incubation period in the dark at room temperature, the absorbance is measured at 718 nm.
Statistical analysis − All experimental bioassay data were performed in triplicate. Statistical analyses were achieved utilizing Student’s t-test in GraphPad Prism version 6.0 software (San Diego, CA, USA). Statistical significance is indicated by p-values (*p < 0.05, **p < 0.01, and ***p < 0.005).
Results and Discussion
The leaves and twigs (ZSAP) and roots (ZSRT) of Z. serrata were initially screened to guide tissue selection for subsequent investigation. Both tissues demonstrated potent radical scavenging activities in DPPH and ABTS assays (Fig. 1A and B). HPLC profiling revealed that while both tissues share similar major constituents, ZSAP displayed a more diverse profile of minor metabolites. Although ZSRT was characterized by a high concentration of specific major compounds, ZSAP was prioritized for further isolation to better evaluate the effectiveness of the LC-MS-DPPH approach in identifying a broader range of bioactive constituents, including those present in lower concentrations. Consequently, the ZSAP methanol extract was partitioned sequentially with n-hexane, EtOAc, and n-BuOH. Both EtOAc and BuOH fractions exhibited pronounced antioxidant activities (Fig. 1C and D). Comparative LC-MS analysis (Fig. 1E) showed that the EtOAc fraction contained peaks spanning a wide retention time range, whereas the BuOH fraction displayed a narrower distribution concentrated in the polar region. As both fractions demonstrated comparable bioactivity, but the EtOAc fraction possessed superior chemical diversity, it was prioritized for subsequent compound isolation.
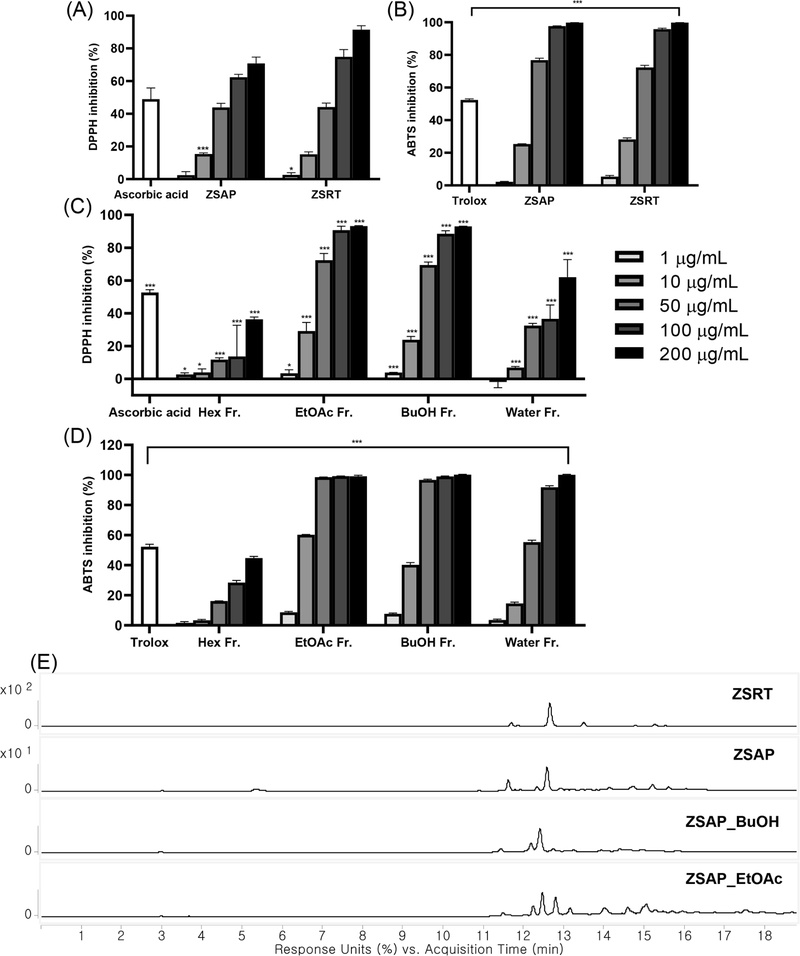
Antioxidant activities and HPLC profiles guide tissue and fraction selection for bioactivity-guided isolation. (A) DPPH and (B) ABTS radical scavenging activities of leaves and twigs (ZSAP) and roots (ZSRT). (C) DPPH and (D) ABTS activities of n-hexane (Hex Fr.), ethyl acetate (EtOAc Fr.), n-butanol (BuOH Fr.), and water (Water Fr.) fractions from ZSAP. (E) HPLC chromatographic profiles (280 nm) showing chemical composition of ZSRT, ZSAP, and the most active fractions (BuOH and EtOAc). Data represent mean ± SD (n = 3). *p < 0.05, **p < 0.01, ***p < 0.005 vs. control. Ascorbic acid and trolox were used as positive controls.
An online DPPH assay coupled with LC-QTOF-MS was performed to rapidly identify antioxidant-active compounds in the EtOAc fraction. The chromatographic separation was monitored by MS detection to obtain the chemical profile (Fig. 2, upper panel). Simultaneously, the column eluent was collected in 30-second intervals into a 96-well microplate for post-column DPPH radical scavenging assessment. The DPPH activity trace revealed multiple negative deflections, corresponding to compounds with free radical scavenging capacity (Fig. 2, lower panel). Analysis of the activity profile indicated that peaks between 12−16 min exhibited significant DPPH scavenging activity and were responsible for the antioxidant properties of the EtOAc fraction. These bioactive peaks were therefore prioritized for isolation and structural characterization.
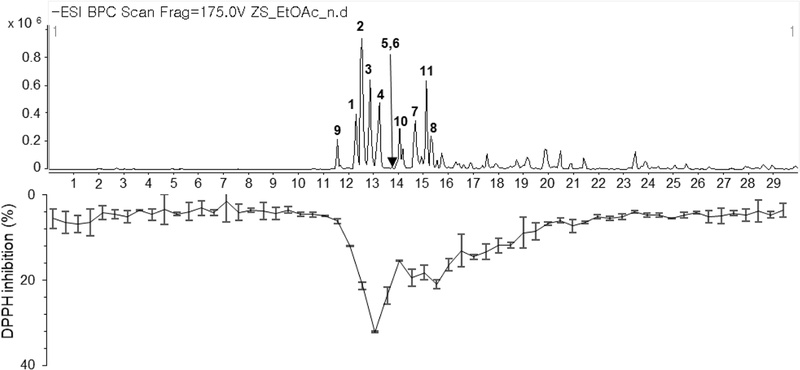
Online DPPH screening coupled with LC-QTOF-MS analysis of the EtOAc fraction from Z. serrata leaves and twigs. Base peak chromatogram (upper panel) and DPPH radical scavenging trace (lower panel, negative peaks indicate antioxidant activity) of the EtOAc fraction.
To facilitate targeted isolation and predict the chemical nature of bioactive constituents, high-resolution mass spectrometry coupled with tandem MS/MS fragmentation analysis was performed on the active peaks identified in the LC-MS-DPPH screening. HRESIMS data provided molecular formulas for the detected compounds, revealing that the majority were flavonoid derivatives (Fig. 3). MS/MS fragmentation patterns revealed distinct behaviors based on glycosylation type (Fig. 3). O-Glycosylated flavonoids (compounds 1, 3, 8, and 10) displayed characteristic neutral losses corresponding to sugar moieties—132 Da for pentoses (apiose, xylose) and 162 Da for hexoses (glucose)—followed by retro-Diels-Alder fragmentations of the aglycone. C-Glycosylated compounds (compounds 2, 7, 9, 11) exhibited diagnostic fragmentations at [M−H−90]⁻ and [M−H−120]⁻, arising from cross-ring cleavages of the C-linked sugar moiety. These fragmentations are characteristic of C-glycosides and readily distinguish them from O-glycosidic isomers.16,17 Based on MS/MS fragmentation patterns and comparison with literature data, tentative identifications were proposed (Table 1). These MS/MS-guided predictions directed subsequent preparative isolation efforts toward confirmation of the predicted structures.
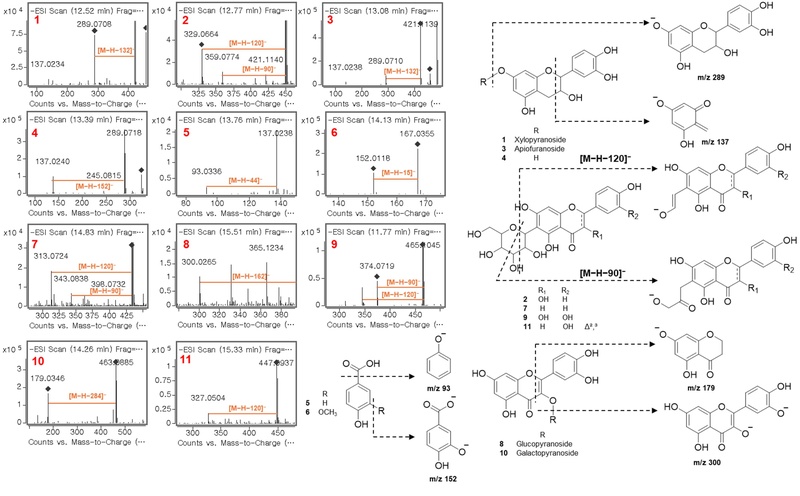
MS/MS fragmentation spectra and diagnostic fragmentation patterns of compounds detected in the EtOAc fraction.

Identification of chemical constituents from the EtOAc fraction of Z. serrata leaves and twigs using LC-QTOF-MS/MS
The EtOAc fraction was separated over silica gel and prep-HPLC to obtain eight known compounds (1‒8), which were determined to be catechin-7-O-β-D-xylopyranoside (1),18 aromadendrin-6-C-β-D-glucopyranoside (2),18 catechin-7-O-β-D-apiofuranoside (3),19 catechin (4),19 4-hydroxybenzoic acid (5),20 vanillic acid (6),21 hemiphloin (7),18 and isoquercetin (8)22 by analyzing and comparing their spectroscopic data with those reported in the literature. Notably, compounds 1, 3, and 7 are reported herein from this genus for the first time.
Compound 1 was obtained as a yellow amorphous powder. The HRESIMS data exhibited molecular ion peaks at m/z 421.1142 [M−H]−, consistent with the molecular formula C20H22O10. The 1H-NMR spectrum of 1 exhibited an ABX coupling system [δH 6.84 (1H, d, J = 2.0 Hz, H-2′), 6.77 (1H, d, J = 8.1 Hz, H-5′), 6.73 (1H, dd, J = 8.1, 2.0 Hz, H-6′)], two meta-coupled doublets at δH 6.18 (1H, d, J = 2.3 Hz, H-6) and 6.12 (1H, d, J = 2.3 Hz, H-8), two hydroxylated methine at δH 4.61 (1H, d, J = 7.4 Hz, H-2) and 4.01 (1H, td, J = 7.4, 5.4 Hz, H-3), one methylene at δH 2.86 (1H, dd, J = 16.3, 5.4 Hz) and 2.55 (1H, dd, J = 16.3, 8.0 Hz), and one anomeric proton at δH 4.77 (1H, d, J = 7.2 Hz, H-1′′). These spectroscopic data were in complete agreement with those reported for catechin-7-O-β-D-xylopyranoside previously isolated from U. macrocarpa.18 Accordingly, the structure of 1 was unambiguously assigned as catechin-7-O-β-D-xylopyranoside.
Compound 3 displayed spectroscopic features characteristic of a catechin moiety similar to those of compound 1, with the exception of the sugar moiety signals. The 1H NMR spectrum exhibited resonances at δH 5.49 (1H, d, J = 3.0Hz), 4.14 (1H, d, J = 3.0 Hz), 4.09 (1H, d, J = 9.7 Hz), 3.85 (1H, d, J = 9.7 Hz), 3.64 (1H, d, J = 11.5 Hz), 3.61 (1H, d, J = 11.5 Hz), indicative of an apiofuranoside unit rather than the xylopyranoside moiety present in 1. Comparison with literature data confirmed compound 3 as catechin-7-O-β-D-apiofuranoside.19
Compound 7 was assigned the molecular formula C21H22O10 based on HRESIMS analysis. The 1H-NMR spectrum revealed an A2B2 coupling pattern with signals at δH 7.31 (2H, d, J = 8.6 Hz, H-2′, H-6′), 6.82 (2H, d, J = 8.6 Hz, H-3′, H-5′), characteristic of a para-disubstituted B-ring. Additional resonances included an aromatic singlet at δH 5.97 (1H, s, H-8), an oxygenated methine proton at δH 5.34 (1H, dd, J = 12.7, 3.0 Hz), a methylene group at δH 3.12 (1H, dd, J = 17.1, 12.7 Hz) and 2.73 (1H, dd, J = 17.1, 3.0 Hz), and an anomeric proton at δH 4.80 (1H, d, J = 9.9 Hz, H-1′′). The spectroscopic features of 7 were consistent with those reported for naringenin-6-C-β-Dglucopyranoside, leading to its structural assignment as this known flavonoid glycoside.18
To validate the online DPPH screening and evaluate the contribution of individual compounds to the EtOAc fraction's antioxidant activity, all isolated compounds were assessed using both DPPH and ABTS radical scavenging assays (Table 2).
In the online LC-MS-DPPH screening, the most prominent negative peaks corresponded to the retention time regions of compounds 3 and 4, reflecting their superior intrinsic radical scavenging capacity rather than their relative abundance in the fraction, as confirmed by individual IC50 determination (Table 2). Compound 4 exhibited the most potent DPPH radical scavenging activity (IC50 = 13.67 ± 1.21 μM), significantly surpassing the positive control ascorbic acid (IC50 = 61.51 ± 10.22 μM). Compound 3 also displayed notable activity (IC50 = 56.36 ± 9.62 μM), comparable to ascorbic acid. Compounds 1 and 8 showed moderate activity with IC50 values of 126.1 ± 15.15 μM and 165.95 ± 2.55 μM, respectively. The remaining compounds showed no significant DPPH activity (IC50 > 200 μM), consistent with the weak negative deflections observed in their corresponding retention time regions during online screening.
In the ABTS assay, compound 4 again demonstrated the strongest activity (IC50= 2.69 ± 0.48 μM), which was approximately 3-fold more potent than Trolox (IC50 = 8.22 ± 2.16 μM). Notably, compounds 6 and 8 exhibited strong ABTS radical scavenging activity (IC50 = 8.79 ± 0.92 μM and 9.74 ± 2.68 μM, respectively), comparable to Trolox. All tested compounds showed measurable activity in the ABTS assay, with IC50 values ranging from 2.69 to 98.91 μM.
The radical scavenging activities of compounds 1–3 and 5–8 were also in good agreement with those previously reported for their respective compound classes, including catechin glycosides (1 and 3),23C-glycosylated flavanonols and flavanones (2 and 7),24,25 hydroxybenzoic acid derivatives (5 and 6),26 and isoquercetin (8).27
The discrepancies in IC50 values between the two assays (Table 2) stem from different structural requirements. DPPH scavenging is more restrictive, often requiring specific features like catechol groups (Bors criteria), which explains the weak activity of 5 and 6. Conversely, the methoxy group in 6 notably enhanced its ABTS scavenging potency. Furthermore, steric hindrance from bulky sugar moieties in glycosides 7 and 8 likely obstructed access to the large DPPH radical, whereas the less hindered ABTS radical reacted more effectively with these structures.28–30
The potent activity of 4 in both assays identifies it as the principal antioxidant, corroborating the significant negative peak observed during LC-MS-DPPH screening. These results validate the LC-MS-DPPH hyphenated approach for efficient identification of bioactive antioxidants in Z. serrata leaves and twigs.
In conclusion, through systematic bioactivity-guided investigation, this study identified the antioxidant constituents of Z. serrata leaves and twigs. Comparative tissue screening, solvent fractionation, and LC-MS-DPPH online analysis enabled efficient isolation of eight compounds, among which catechin (4) was identified as the primary antioxidant. Notably, compounds 1, 3, and 7 are reported from the genus Zelkova for the first time. These findings demonstrate the effectiveness of the LC-MS-DPPH hyphenated approach for natural product discovery and establish Z. serrata as a promising source of bioactive antioxidants with potential therapeutic applications.
Acknowledgments
This research was supported by National Research Foundation of Korea grants funded by the Korean Government (RS-2019-NR040070 and RS-2021-NR058871) and ITRC support program supervised by the IITP and funded by the MSIT (IITP-2026-RS-2023-00258971, 30%).
Conflicts of Interest
The authors declare that they have no conflicts of interest.
References
-
Di Carlo, E.; Sorrentino, C. Antioxidants 2024, 13, 1109.
[https://doi.org/10.3390/antiox13091109]

-
Srivilai, J.; Nuengchamnong, N.; Khorana, N.; Wisidsri, N.; Thungmungmee, S.; Aoonboontum, P.; Sasea, K.; Suksathan, P.; Rungsang, T.; Ingkaninan, K. Phytochem. Anal. 2025, 36, 704–717.
[https://doi.org/10.1002/pca.3461]

-
Chang, T.-C.; Chang, S.-T.; Cheng, S.-S. J. Wood Chem. Technol. 2022, 42, 265–273.
[https://doi.org/10.1080/02773813.2022.2085745]

-
Yumita, A.; Hanani, E.; Agustina, A.; Damayanti, F.; Priani, K. N.; Fadila, S. i. N. Nat. Prod. Sci. 2023, 29, 24–30.
[https://doi.org/10.20307/nps.2023.29.1.24]

-
Scalbert, A.; Manach, C.; Morand, C.; Rémésy, C.; Jiménez, L. Crit. Rev. Food Sci. Nutr. 2005, 45, 287–306.
[https://doi.org/10.1080/1040869059096]

-
Vu, N. K.; Le, T. T.; Tran, T. T.; Ha, M. T.; Kim, J. A.; Min, B. S. Nat. Prod. Sci. 2025, 31, 43–48.
[https://doi.org/10.20307/nps.2025.31.1.43]

-
Park, S. Y.; Han, Y. K.; Vinh, L. B.; Lee, K. Y. Nat. Prod. Sci. 2023, 29, 242–250.
[https://doi.org/10.20307/nps.2023.29.4.242]

-
Kang, H.-J.; Jang, Y.-J. Int. J. Oral Sci. 2012, 4, 78–84.
[https://doi.org/10.1038/ijos.2012.14]

- Orwa, C.; Mutua, A.; Kindt, R.; Jamnadass, R.; Anthony, S. Agroforestree Database: A Tree Reference and Selection Guide, version 4.0; World Agroforestry Centre: Nairobi, Kenya, 2009.
- Cheng, S.-S.; Lin, C.-Y.; Liu, S.-L.; Chang, S.-T.; Tsai, K.-H. Jour. Exp. For. Natl. Taiwan Univ. 2021, 35, 25–38.
- Yen, P.-L.; Cheng, S.-S.; Chang, S.-T. Jour. Exp. For. Natl. Taiwan Univ. 2016, 30, 247–258.
-
Kim, Y. J.; Jung, M.; Vinh, L. B.; Kim, J. A.; Kim, H.-J.; Lee, M. K.; Lee, K. Y. Food Chem. X 2025, 31, 103107.
[https://doi.org/10.1016/j.fochx.2025.103107]

-
Han, Y. K.; Vinh, L. B.; Nam, M.-H.; Lee, K. Y. Appl. Biol. Chem. 2023, 66, 53.
[https://doi.org/10.1186/s13765-023-00814-1]

-
Shin, H.; Chung, H.; Park, B.; Lee, K. Y. Nat. Prod. Sci. 2016, 22, 64–69.
[https://doi.org/10.20307/nps.2016.22.1.64]

-
Vu, N. K .; K im, C. S.; Ha, M. T.; Ngo, Q.-M. T.; Park, S. E.; Kwon, H.; Lee, D.; Choi, J. S.; Kim, J. A.; Min, B. S. J. Agric. Food. Chem. 2020, 68, 8797–8811.
[https://doi.org/10.1021/acs.jafc.0c02122]

-
Sobeh, M.; Rezq, S.; Sabry, O. M.; Abdelfattah, M. A. O.; El Raey, M. A.; El-Kashak, W. A.; El-Shazly, A. M.; Mahmoud, M. F.; Wink, M. Biomed. Pharmacother. 2019, 115, 108882.
[https://doi.org/10.1016/j.biopha.2019.108882]

-
Patras, M. A.; Davalos, J. Z.; Kuhnert, N. J. Mass Spectrom. 2023, 58, e4972.
[https://doi.org/10.1002/jms.4972]

-
Wang, W.; Jeong, C.; Lee, Y.; Park, C.; Oh, E.; Park, K.-H.; Cho, Y.; Kang, E.; Lee, J.; Cho, Y.-J. Park, J. H. Y.; Son, Y.-J.; Lee, K. W.; Kang, H. ACS omega#160;2022, 7, 4840–4849.
[https://doi.org/10.1021/acsomega.1c05305]

-
Jung, M. J.; Heo, S.-I.; Wang, M.-H. Food Chem. 2008, 108, 482–487.
[https://doi.org/10.1016/j.foodchem.2007.10.081]

-
Zhang, H.; Conte, M. M.; Huang, X.-C.; Khalil, Z.; Capon, R. J. Org. Biomol. Chem. 2012, 10, 2656–2663.
[https://doi.org/10.1039/c2ob06747a]

-
Refaat, J.; Samy, M. N.; Desoukey, S. Y.; Ramadan, M. A.; Sugimoto, S.; Matsunami, K.; Kamel, M. S. Med. Chem. Res. 2015, 24, 2939–2949.
[https://doi.org/10.1007/s00044-015-1342-8]

-
Kazuma, K.; Noda, N.; Suzuki, M. Phytochemistry 2003, 62, 229–237.
[https://doi.org/10.1016/S0031-9422(02)00486-7]

- Na, M. K.; An, R. B.; Lee, S. M.; Min, B. S.; Kim, Y. H.; Bae, K. H.; Kang, S. S. Nat. Prod. Sci. 2002, 8, 26–29.
- Chang, B.-S.; Kwon, Y.-S.; Kim, C.-M. Kor. J. Pharmacogn. 2004, 35, 80–87.
-
Yuan, X.; Xue, X.; Liang, Z.; He, C. Nat. Prod. Res. 2025, 39, 3504–3511.
[https://doi.org/10.1080/14786419.2024.2340756]

-
Moazzen, A.; Öztinen, N.; Ak-Sakalli, E.; Koşar, M. Heliyon 2022, 8, e10467.
[https://doi.org/10.1016/j.heliyon.2022.e10467]

-
Nile, A.; Nile, S. H.; Shin, J.; Park, G.; Oh, J.-W. Int. J. Mol. Sci. 2021, 22, 10749.
[https://doi.org/10.3390/ijms221910749]

-
Platzer, M.; Kiese, S.; Tybussek, T.; Herfellner, T.; Schneider, F.; Schweiggert-Weisz, U.; Eisner, P. Front. Nutr. 2022, 9, 882458.
[https://doi.org/10.3389/fnut.2022.882458]

-
Platzer, M.; Kiese, S.; Herfellner, T.; Schweiggert-Weisz, U.; Eisner, P. Antioxidants 2021, 10, 811.
[https://doi.org/10.3390/antiox10050811]

-
Platzer, M.; Kiese, S.; Herfellner, T.; Schweiggert-Weisz, U.; Miesbauer, O.; Eisner, P. Molecules 2021, 26, 1244.
[https://doi.org/10.3390/molecules26051244]
