Nitric Oxide Production Inhibitory Terpenylated Coumarins from Ailanthus altissima
Abstract
LC-MS/MS-based molecular networking was used to guide the targeted isolation of terpenylated coumarins from the barks of Ailanthus altissima. Five known terpenylated coumarins (1–5), along with two simple coumarins (6 and 7), were isolated from the CH2Cl2-soluble fraction. Their structures were determined using spectroscopic techniques, including 1D and 2D NMR and LC-HR-MS/MS. Notably, altissimacoumarins C (1), F (3), and H (4) demonstrated potent inhibition of LPS-induced nitric oxide production in RAW 264.7 macrophages, with IC50 values of 36.1, 27.9, and 11.2 μM, respectively. These results demonstrate the anti-inflammatory properties of A. altissima, supporting its possible application in treating inflammatory disorders.
Keywords:
Ailanthus altissima, Simaroubaceae, Terpenylated coumarin, Molecular networking, Nitric oxideIntroduction
Ailanthus altissima (Mill.) Swingle, belongs to Simaroubaceae family, is widely distributed in Korea, China, India, and Japan. It has long been used in folkloric medicine as astringent, antidiarrheal, antiparasitic, antihemorrhagic, and anti-inflammatory agents.1 In Korea, the dried bark of A. altissima has been employed to treat various inflammatory conditions, particularly chronic cervicitis, endometritis, ulcerative colitis, and vaginal mucositis.2 Previous phytochemical investigations on A. altissima have revealed the presence of alkaloids, coumarins, phenylpropanoids, quassinoids, and triterpenoids.1,2 Among these, quassinoids and indole alkaloids are characteristic components and the most widely studied components in pharmacology, with antitumor,3–7 antimalarial,8 antifeedant,9 anti-aging,10 and anti-inflammatory activities.11-14 The ethanol extracts of A. altissima leaves have been reported to possess anti-inflammatory activity in LPS-stimulated primary astrocytes,15 as well as in bone marrow-derived mast cells and in models of ovalbumin-induced lung inflammation.16 The methanolic extract of A. excelsa stem bark was also shown to possess significant anti-inflammatory activity in rat model.17 In addition, an aqueous extract of A. altissima bark demonstrated anti-inflammatory effects in dextran sulfate sodium-induced acute ulcerative colitis mouse model, by inhibiting the activation of the PI3K/AKT pathway.18 The nitric oxide inhibitory activity of several canthinone-type alkaloids including canthin-6-one, 9-hydroxycanthin-6-one, 10-hydroxycanthin-6-one, and 5-(1-hydroxyethyl)-canthin-6-one and the quassinoid, ailanthone, from A. altissima have been reported.11-14 However, investigations of other anti-inflammatory constituents are still limited.
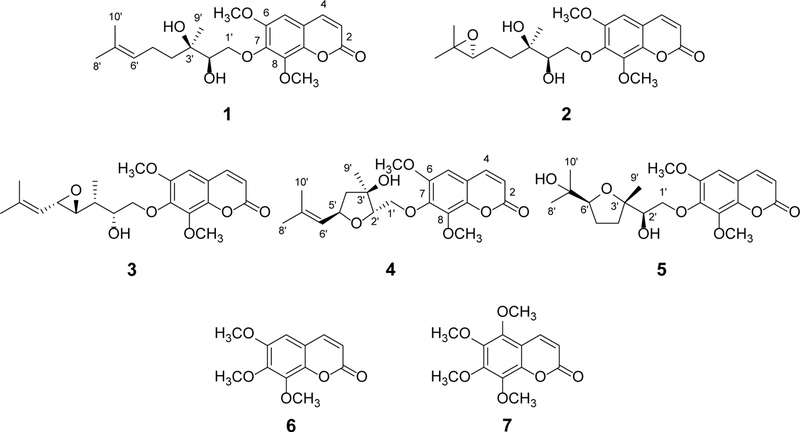
Recently, molecular networking on the Global Natural Products Social (GNPS) platform, based on LC-MS/MS fragment similarities, has proven to be a powerful strategy for the identification of bioactive compounds from natural resources.19–21 Accordingly, LC-MS/MS-based molecular networking was employed to discover anti-inflammatory terpenylated coumarins from the CH2Cl2-soluble fraction of the bark of A. altissima. As a result, seven coumarins (1–7) (Fig. 1) including five terpenylated coumarins (1–5) were isolated and identified.
Experimental
General experimental procedures – UV spectra were recorded on a JASCO UV-550 spectrophotometer (JASCO, Tokyo, Japan). 1D and 2D NMR spectra were recorded on Bruker AVANCE 400 and 900 MHz spectrometer (Bruker, MA, USA). LC-HRMS/MS analyses were performed using an Orbitrap Exploris 120 mass spectrometer, connected to a Vanquish UHPLC system and diode array detector (Thermo Fisher Scientific, MA, USA). MPLC was performed using the Biotage Isolera Prime chromatography system. Semi-preparative HPLC was carried out using a Waters HPLC system, equipped with two Waters 515 pumps, a 2996 photodiode array detector, and three preparative columns, such as YMC J’sphere ODS-H80 (250 × 20 mm, 4 μm, flow rate 10 mL/min), YMC J’sphere ODS-H80 (150 × 20 mm, 4 μm, flow rate 6 mL/min), and YMC J’sphere ODS-H80 (150 × 10 mm, 4 μm, flow rate 3 mL/min). TLC was conducted on silica gel 60 F254 plates (0.25 mm, Merck, Darmstadt, Germany), and the spots were detected using a spray reagent composed of a 10% solution of vanillin solution in H2SO4.
Plant materials – The dried bark of A. altissima was purchased from the Kyungdong herbal market (Seoul, Republic of Korea) in September 2022. A voucher specimen (CBNU-2022-09-AA) was identified by professor B. Y. Hwang and deposited at the herbarium of the College of Pharmacy, Chungbuk National University, Republic of Korea.
Extraction and isolation – The dried and powdered bark of A. altissima (3.0 kg) was extracted with MeOH (18 L × 2) at room temperature for 3 days. The extract was filtered and evaporated under reduced pressure, and the resulting residue (220.0 g) was suspended in water and sequentially partitioned with n-hexane (4 L × 1), CH2Cl2 (4 L × 1), EtOAc (4 L × 1) and n-BuOH (4 L × 1). The CH2Cl2-soluble fraction (31.8 g) was fractionated by MPLC using Biotage Sfär Silica (Uppsala, Sweden) with a CH2Cl2-MeOH step gradient system (100:0 to 0:100) to obtain 9 fractions, (AAC1–AAC9). AAC4 (2.8 g) was fractionated by MPLC using Biotage Sfär Silica (Uppsala, Sweden) with n-hexane-Acetone gradient system (100:0 to 0:100) to obtain 6 subfractions, (AAC4.1–AAC4.6). AAC4.5 (1.2 g) was separated by MPLC using Biotage Sfär C18 D (Uppsala, Sweden) with a H2O-MeOH gradient system (100:0 to 0:100), and 11 subfractions (AAC4.5.1–4.5.11) were obtained. AAC4.5.5 (36.0 mg) was separated using preparative HPLC [YMC J’sphere ODS-H80 (150 × 10 mm, 4 μm, flow rate 3 mL/min)] using a H2O-MeOH isocratic system (70:30) to afford compound 6 (1.7 mg; tR = 15.0 min). AAC4.6 (456.0 mg) was separated by MPLC using Biotage Sfär C18 D (Uppsala, Sweden) with a H2O-MeOH gradient system (20:80 to 0:100), and five subfractions (AAC4.6.1–AAC4.6.5) were obtained. AAC4.6.4 (32.0 mg) was further purified by prep HPLC [YMC J’sphere ODS-H80 (150 × 20 mm, 4 μm, flow rate 6 mL/min)] using a H2O-CH3CN isocratic system (45:55) to afford compound 4 (0.6 mg; tR = 11.0 min). AAC6 (10.0 g) was fractionated by MPLC using Biotage Sfär Silica (Uppsala, Sweden) with a CH2Cl2-MeOH gradient system (100:0 to 0:100), and nine subfractions (AAC6.1–AAC6.9) were obtained. AAC6.2 (312.0 mg) was separated using prep HPLC [YMC J’sphere ODS-H80 (250 × 20 mm, 4 μm, flow rate 10 mL/min)] and fraction collector with a H2O-CH3CN gradient system (63:37 to 67:33), and 14 subfractions (AAC6.2.1–AAC6.2.14) were obtained. AAC6.2.9 (39.9 mg) purified by preparative HPLC [YMC J’sphere ODS-H80 (150 × 20 mm, 4 μm, flow rate 6 mL/min)] using a H2O-CH3CN isocratic system (68:32) to afford compounds 2 (0.6 mg; tR = 26.8 min), and 5 (1.6 mg; tR = 23.5 min). AAC6.2.12 (11.1 mg) purified by preparative HPLC [YMC J’sphere ODS-H80 (150 × 20 mm, 4 μm, flow rate 6 mL/min)] using a H2O-CH3CN isocratic system (68:32) to afford compound 7 (1.3 mg; tR = 14.2 min). AAC6.3 (272.0 mg) was fractionated by MPLC using Biotage Sfär Silica (Uppsala, Sweden) with CH2Cl2-Acetone gradient system (100:0 to 0:100), and six subfractions (AAC6.3.1–AAC6.3.6) obtained include compound 1 (5.9 mg). AAC6.3.4 (31.3 mg) purified by preparative HPLC [YMC J’sphere ODS-H80 (150 × 10 mm, 4 μm, flow rate 3 mL/min)] using a H2O-MeOH isocratic system (35:65) to afford compound 3 (0.6 mg; tR = 15.0 min).
Altissimacoumarin C (1) – Yellow oil. + 8.8 (c 0.1, MeOH); 1H-NMR (400 MHz, CDCl3): δ 7.61 (1H, d, J = 9.5 Hz, H-4), 6.69 (1H, s, H-5), 6.36 (1H, d, J = 9.5 Hz, H-3), 5.11 (1H, m, H-6′), 4.56 (1H, dd, J = 10.5, 2.0 Hz, H-1′α), 4.06 (3H, s, 8-OCH3), 4.02 (1H, dd, J = 10.5, 8.0 Hz H-1′β), 3.91 (3H, s, 6-OCH3), 3.70 (1H, dd, J = 7.8, 2.4 Hz, H-2′), 2.13 (1H, m, H-5′α), 2.05 (1H, m, H-5′β), 1.67 (3H, s, H-8′), 1.62 (1H, m, H-4′α), 1.61 (3H, s, H-10′), 1.40 (1H, m, H-4′β), 1.25 (3H, s, H-9′); 13C-NMR (100 MHz, CDCl3): δ 160.2 (C-2), 149.7 (C-6), 144.7 (C-7), 143.4 (C-4), 143.0 (C-9), 141.1 (C-8), 132.0 (C-7′), 124.2 (C-6′), 115.6 (C-3), 114.8 (C-10), 103.8 (C-5), 76.2 (C-1′), 75.1 (C-2′), 73.4 (C-3′), 62.0 (8-OCH3), 56.3 (6-OCH3), 37.9 (C-4′), 25.7 (C-8′), 23.3 (C-9′), 22.1 (C-5′), 17.7 (C-10′); HR-ESI-MS m/z 415.1727 [M + Na]+ (calcd. for C21H28O7Na, 415.1727).
Altissimacoumarin E (2) – Yellow oil. + 7.6 (c 0.1, MeOH); 1H-NMR (400 MHz , CDCl3): δ 7.61 (1H, d, J = 9.5 Hz, H-4), 6.69 (1H, s, H-5), 6.37 (1H, d, J = 9.5 Hz, H-3), 4.49 (1H, dd, J = 9.6, 1.5 Hz, H-1′α), 4.06 (3H, s, 8-OCH3), 3.91 (1H, m, H-1β), 3.90 (3H, s, 6-OCH3), 3.88 (1H, m, H-2′), 3.85 (1H, m, H-6′), 2.23 (1H, m, H-4′α), 1.98 (1, m, H-5′α), 1.91 (1H, m, H-5′β), 1.55 (1H, m, 4′β), 1.25 (3H, s, H-8′) 1.19 (3H, s, H-9′), 1.09 (3H, s, H-10′); 13C-NMR (100 MHz, CDCl3): δ 160.2 (C-2), 150.0 (C-6), 144.6 (C-7), 143.4 (C-4), 142.5 (C-9), 141.3 (C-8), 115.6 (C-3), 114.9 (C-10), 103.9 (C-5), 85.3 (C-6′), 83.8 (C-3′), 76.4 (C-1′), 75.3 (C-2′), 71.6 (C-7′), 62.0 (8-OCH3), 56.4 (6-OCH3), 33.2 (C-4′), 27.7 (C-8′), 26.4 (C-5′), 24.9 (C-10′), 23.4 (C-9′); HR-ESI-MS m/z 431.1673 [M + Na]+ (calcd. for C21H28O8Na, 431.1676).
Altissimacoumarin F (3) – Yellow oil. + 136.0 (c 0.1, MeOH); 1H-NMR (400 MHz, CDCl3): δ 7.61 (1H, d, J = 9.5 Hz, H-4), 6.67 (1H, s, H-5), 6.35 (1H, d, J = 9.5 Hz, H-3), 5.19 (1H, m, H-6′), 4.86 (1H, m, H-5′), 4.44 (1H, dd, J = 10.4, 4.4 Hz, H-1′α), 4.10 (1H, dd, J = 10.4, 8.0 Hz, H-1′β), 4.05 (1H, m, H-4′), 4.02 (3H, s, 8-OCH3), 3.88 (3H, s, 6-OCH3), 2.20 (1H, m, H-3′), 1.72 (1H, s, H-8′), 1.71 (3H, s, H-10′), 1.47 (3H, s, H-9′); 13C-NMR (100 MHz, CDCl3): δ 160.4 (C-2), 149.5 (C-6), 144.7 (C-7), 143.4 (C-4), 143.2 (C-9), 140.7 (C-8), 137.0 (C-7′), 125.3 (C-6′), 115.3 (C-3), 114.4 (C-10), 103.9 (C-5), 84.6 (C-4′), 79.4 (C-2′), 74.3 (C-5′), 73.1 (C-1′), 62.0 (8-OCH3), 56.2 (6-OCH3), 47.5 (C-3′), 25.9 (C-8′), 23.3 (C-9′), 18.2 (C-10′); HR-ESI-MS m/z 413.1570 [M + Na]+ (calcd. for C21H26O7Na, 413.1571).
Altissimacoumarin H (4) – Yellow oil. + 23.9 (c 0.1, MeOH); 1H-NMR (900 MHz, CDCl3): δ 7.60 (1H, d, J = 9.5 Hz, H-4), 6.67 (1H, s, H-5), 6.35 (1H, d, J = 9.5 Hz, H-3), 5.35 (1H, m, H-6), 4.77 (1H, m, H-5′), 4.63 (1H, m, H-1′β), 4.11 (1H, dd, J = 9.1, 4.8 Hz, H-2′), 4.03 (3H, s, 8-OCH3), 4.01 (1H, m, H-1′α), 3.88 (3H, s, 6-OCH3), 2.17 (1H, m, H-4′α), 2.01 (1H, m, H-4′β), 1.74 (3H, d, J = 1.2 Hz, H-8′), 1.68 (3H, d, J = 1.2 Hz, H-10′), 1.40 (3H, s, H-9′); 13C-NMR (225 MHz, CDCl3): δ 160.3 (C-2), 149.3 (C-6), 144.6 (C-7), 143.4 (C-4), 143.2 (C-9), 140.3 (C-8), 136.3 (C-7′), 126.2 (C-6′), 115.3 (C-3), 114.2 (C-10), 103.8 (C-5), 81.6 (C-2′), 78.1 (C-3′), 74.0 (C-5′), 73.1 (C-1′), 61.9 (8-OCH3), 56.2 (6-OCH3), 47.2 (C-4′), 25.9 (C-8′), 22.0 (C-9′), 18.1 (C-10′); HR-ESI-MS m/z 413.1572 [M + Na]+ (calcd. for C21H26O7Na, 413.1571).
Altissimacoumarin K (5) – Yellow oil. + 33.8 (c 0.1, MeOH); 1H-NMR (400 MHz , CDCl3): δ 7.62 (1H, d, J = 9.5 Hz, H-4), 6.70 (1H, s, H-5), 6.37 (1H, d, J = 9.5 Hz, H-3), 4.53 (1H, dd, J = 10.1, 2.1 Hz, H-1′), 4.07 (3H, s, 8-OCH3), 3.91 (3H, s, 6-OCH3), 3.85 (1H, dd, J = 9.0, 2.1 Hz, H-2′), 3.78 (1H, t, J = 7.8 Hz, H-6′), 2.14 (1H, m, H-4′α), 1.86 (2H, m, H-5′), 1.74 (1H, m, H-4′β), 1.21 (3H, s, H-8′), 1.19 (3H, s, H-9′), 1.12 (3H, s, H-10′); 13C-NMR (100 MHz, CDCl3): δ 160.3 (C-2), 150.0 (C-6), 145.0 (C-7), 143.4 (C-4), 143.0 (C-9), 141.3 (C-8), 115.5 (C-3), 114.8 (C-10), 103.8 (C-5), 87.0 (C-6′), 83.4 (C-3′), 76.7 (C-1′), 75.5 (C-2′), 70.6 (C-7′), 62.0 (8-OCH3), 56.4 (6-OCH3), 35.3 (C-4′), 27.6 (C-8′), 26.2 (C-5′), 24.0 (C-10′), 22.5 (C-9′); HR-ESI-MS m/z 431.1672 [M + Na]+ (calcd. for C21H28O8Na, 431.1676).
6,7,8-Trimethoxycoumarin (6) – Yellow solid. 1H-NMR (400 MHz, CDCl3): δ 7.61 (1H, d, J = 9.5 Hz, H-4), 6.66 (1H, s, H-5), 6.35 (1H, d, J = 9.5 Hz, H-3), 4.04 (3H, s, 8-OCH3), 4.00 (3H, s, 7-OCH3), 3.90 (3H, s, 6-OCH3); 13C-NMR (100 MHz, CDCl3): δ 160.5 (C-2), 150.1 (C-9), 146.0 (C-8), 143.4 (C-4), 143.1 (C-7), 141.2 (C-6), 115.3 (C-3), 114.3 (C-10), 103.7 (C-5), 61.9 (8-OCH3), 61.5 (7-OCH3), 56.3 (6-OCH3). HR-ESI-MS m/z 237.0756 [M + H]+ (calcd. for C12H13O5, 237.0758).
5,6,7,8-Tetramethoxycoumarin (7) – Yellow solid. 1H-NMR (400 MHz, CDCl3): δ 7.94 (1H, d, J = 9.7 Hz, H-4), 6.30 (1H, d, J = 9.7 Hz, H-3), 4.04 (3H, s, 8-OCH3), 3.98 (3H, s, 7-OCH3), 3.97 (3H, s, 6-OCH3), 3.90 (3H, s, 5-OCH3); 13C-NMR (100 MHz, CDCl3): δ 160.4 (C-2), 150.7 (C-9), 145.1 (C-8), 144.3 (C-7), 142.4 (C-5), 138.7 (C-4), 136.9 (C-6), 114.1 (C-3), 109.5 (C-10), 62.1 (8-OCH3), 61.9 (7-OCH3), 61.7 (6-OCH3), 61.5 (5-OCH3); HR-ESI-MS m/z 267.0860 [M + H]+ (calcd. for C13H15O6, 267.0863).
LC-MS/MS analysis and molecular networking – Fragmentation analysis was conducted using LC-HR-MS/MS in data-dependent MSn mode to produce an MS2 spectrum comprising the top four intense ions. A dynamic exclusion filter was utilized to hinder the recurring fragmentation of ions within a 2.5-second window following the acquisition of the MS2 spectrum.21 The LC-HR-MS/MS data were converted to the mzXML format using MS Convert software and subsequently uploaded to the GNPS web platform. Molecular networking was conducted using the GNPS data analysis workflow, which employs a spectral clustering algorithm with a cosine score threshold of 0.6 and requires a minimum of five matched peaks. The resulting molecular networks were visualized with Cytoscape software, version 3.8.2
Measurement of LPS-induced NO production and cell viability – RAW 264.7 cells (American Type Culture Collection, Manassas, VA, USA) were cultured in Dulbecco’s Modified Eagle’s medium (DMEM, Gibco-BRL. Louis, MO, USA) containing 10% heat-inactivated fetal bovine serum, 100 U/mL penicillin, and 100 μg/mL streptomycin, in a humidified atmosphere with 5% CO2 at 37°C. The cells were seeded in a 96-well plate (2 × 105 cells/well) and cultured for 24 h. Subsequently, the cells were expose to the test compounds and extracts, which were initially dissolved in dimethyl sulfoxide (DMSO) and then diluted with DMEM to create a variety of concentrations (final concentration range: Aminoguanidine = 0.5–100 μM, and isolated compounds = 1–200 μM). Subsequently, the cells were stimulated with LPS (1 μg/mL) to induce NO production and incubated for 24 h at 37°C. After incubation, 100 μL of cell-free supernatant was mixed with an equal volume of Griess reagent containing 2% (w/v) sulfanilamide in 5% (w/v) phosphoric acid and 0.2 (w/v) of N-(1-naphthyl) ethylenediamine. Nitrite levels were determined by measuring the absorbance at 550 nm using a sodium nitrite standard calibration curve. Cell viability was determined using the 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl tetrazolium bromide (MTT) colorimetric assay (Sigma-Aldrich, St. Louis, MO, USA).14
Results and Discussion
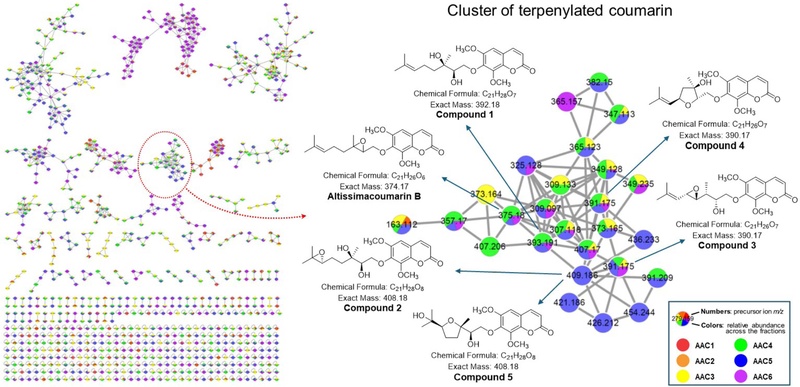
Recently, a naphthoquinone-containing terpenylated coumarin, ailancoumaquinone A, from A. altissima, has been reported to exhibit significant anti-inflammatory activity via inhibition of NO production in LPS-stimulated RAW 264.7 macrophages.22 LC-MS/MS-based molecular networking combined with GNPS spectral library analysis was applied to further explore terpenylated coumarins from the bark of A. altissima. In the CH2Cl2-soluble fraction, one node at m/z 375.180 within cluster A was annotated as altissimacoumarin B, indicating that other nodes in this cluster corresponded to structurally related terpenylated coumarins. Accordingly, molecular networking-guided isolation and further fractionation of the CH2Cl2-soluble-fraction by column chromatography and semipreparative HPLC afforded seven known coumarins (1–7), including five terpenylated coumarins (1–5). On the basis of NMR and MS spectral analysis and comparison with reported literature data, compounds (1–7) were identified as altissimacoumarins C (1),10 E (2),10 F (3),10 H (4),23 and K (5),6 along with 6,7,8-trimethoxycoumarin (6)24 and 5,6,7,8-tetramethoxycoumarin (7).24
All isolated compounds (1–7) were assessed for their anti-inflammatory activity by measuring inhibition of LPS-induced NO production in RAW 264.7 macrophage cells, with aminoguanidine used as a positive control (IC50: 19.8 μM). The IC50 values for NO inhibition and CC50 values in RAW 264.7 cells were presented in Table 1. Notably, the terpenylated coumarins altissimacoumarins C (1), F (3), and H (4) exhibited significant NO inhibitory activity, with IC50 values of 36.1, 27.9 and 11.2 μM, respectively, whereas the two simple coumarins were inactive (IC50 > 50 μM). These results, together with previously findings on prenylated flavonoids,25 suggest that the terpenyl moiety may increases the lipophilicity of coumarins, thereby enhancing their ability to penetrate cell membranes. Although the most active compounds 1, 3, and 4 contain a monoterpenenoid moiety with a double bond between C-6′ and C-7′, however, the role of this structural feature in activity are not clear because only a limited number of compounds were isolated.

Cytotoxicity and inhibitory effects of compounds 1–7 on LPS-induced NO production in RAW 264.7 macrophage cellsa
Previous studies also reported that terpenylated coumarins isolated from the leaves of Zanthoxylum schinifolium exhibit anti-inflammatory activity by inhibiting LPS-induced NO production in RAW 264.7 macrophages. Furthermore, the presence of a double bond in the terpenoid moiety was found to be important for NO inhibition.26
Although the NO inhibitory effects of canthinone-type alkaloids and quassinoids from A. altissima have been previously reported, our results suggest that terpenylated coumarins may also contribute to the anti-inflammatory activity of A. altissima. Moreover, LC-MS/MS-based molecular networking approach enabled comprehensive chemical profiling, facilitating the identification of the terpenylated coumarin scaffold in A. altissima. Taken together, this study provides initial evidence that terpenylated coumarins isolated from A. altissima exhibit anti-inflammatory activity, supporting the notion that this plant may serve as a valuable source of anti-inflammatory agents.
Acknowledgments
The authors acknowledge the Center for Research Instruments and Experimental Facilities, Chungbuk National University and Korea Basic Science Institute for the NMR spectroscopic measurements.
Conflicts of Interest
The authors declare that they have no conflicts of interest.
References
- Tang, W.; Eisenbrand, B. Handbook of Chinese medicinal plants: Chemistry, pharmacology, toxicology; Wiley-VCH; Weinheim, 2011; p 45.
-
Li, X.; Li, Y.; Ma, S.; Zhao, Q.; Wu, J.; Duan, L.; Xie, Y.; Wang, S. J. Ethnopharmacol. 2021, 275, 114121.
[https://doi.org/10.1016/j.jep.2021.114121]

-
Bailly, C. Phytother. Res. 2020, 34, 2203–2213.
[https://doi.org/10.1002/ptr.6681]

- Wang, R.; Lu, Y.; Li, H.; Sun, L.; Yang, N.; Zhao, M.; Zhang, M.; Shi, Q. Oncol. Lett. 2018, 15, 6022–6028.
-
Wang, Y.; Wang, W.-J.; Su, C.; Zhang, D.-M.; Xu, L.-P.; He, R.-R.; Wang, L.; Zhang, J.; Zhang, X.-Q.; Ye, W.-C. Bioorg. Med. Chem. Lett. 2013, 23, 654–657.
[https://doi.org/10.1016/j.bmcl.2012.11.116]

-
Yan, Z.-Y.; Lv, T.-M.; Wang, Y.-X.; Shi, S.-C.; Chen, J.-J.; Lin, B.; Liu, Q.-B.; Huang, X.-X.; Song, S.-J. Phytochemistry 2020, 175, 112361.
[https://doi.org/10.1016/j.phytochem.2020.112361]

-
Duan, Z.-K.; Lin, B.; Du, Y.-Q.; Li, C.; Yu, X.-B.; Xue, X.-B.; Liu, Q.-B.; Song, S.-J.; Huang, X.-X. New J. Chem. 2021, 45, 1100–1108.
[https://doi.org/10.1039/D0NJ04872K]

-
Okunade, A. L.; Bikoff, R. E.; Casper, S. J.; Oksman, A.; Goldberg, D. E.; Lewis, W. H. Phytother. Res. 2003, 17, 675–677.
[https://doi.org/10.1002/ptr.1336]

-
Duan, Z.-K.; Wang, X.; Lian, M.-Y.; Guo, S.-S.; Gao, Z.-H.; Bai, M.; Huang, X.-X.; Song, S.-J. J. Agric. Food Chem. 2024, 72, 10958–10969.
[https://doi.org/10.1021/acs.jafc.4c01049]

-
Dao, T.-T.; Tran, T.-L.; Kim, J.; Nguyen, P.-H.; Lee, E.-H.; Park, J.; Jang, I.-S.; Oh, W.-K. J. Nat. Prod. 2012, 75, 1332–1338.
[https://doi.org/10.1021/np300258u]

-
Kim, H. M.; Kim, S. J.; Kim, H.-Y.; Ryu, B.; Kwak, H.; Hur, J.; Choi, J.-H.; Jang, D. S. Bioorg. Med. Chem. Lett. 2015, 25, 1017–1020.
[https://doi.org/10.1016/j.bmcl.2015.01.034]

-
Kim, H. M.; Lee, J. S.; Sezirahiga, J.; Kwon, J.; Jeong, M.; Lee, D.; Choi, J.-H.; Jang, D.S. Molecules 2016, 21, 642.
[https://doi.org/10.3390/molecules21050642]

-
Cho, S.-K.; Jeong, M.; Jang, D. S.; Choi, J.-H. Planta Med. 2018, 84, 527–535.
[https://doi.org/10.1055/s-0043-123349]

-
Xue, J. F.; Zhao, C.-G.; Pan, H.; Duan, J.-J.; Jia, Y.-Y.; Chen, H.; Feng, W.-S.; Xue, G.-M. J. Asian Nat. Prod. Res. 2024, 26, 1247–1253.
[https://doi.org/10.1080/10286020.2024.2360047]

-
Kim, S. R.; Park, Y.; Li, M.; Kim, Y. K.; Lee, S.; Son, S. Y.; Lee, S.; Lee, J. S.; Lee, C. H.; Park, H. H.; Lee, J.-Y.; Hong, S.; Cho, Y.-C.; Kim, J.-W.; Yoo, H. M.; Cho, N.; Lee, H.-S.; Lee, S. H. J. Ethnopharmacol. 2022, 286, 114258.
[https://doi.org/10.1016/j.jep.2021.114258]

-
Jin, M. H.; Yook, J.; Lee, E.; Lin, C. X.; Quan, Z.; Son, K. H.; Bae, K. H.; Kim, H. P.; Kang, S. S.; Chang, H. W. Biol. Pharm. Bull. 2006, 29, 884–888.
[https://doi.org/10.1248/bpb.29.884]

-
Sapkal, P. R.; Tatiya, A. U.; Firke, S. D.; Redasani, V. K.; Gurav, S. S.; Ayyanar, M.; Jamkhande, P. G.; Surana, S. J.; Mutha, R. E.; Kalaskar, M. G. Heliyon 2023, 9, e15952.
[https://doi.org/10.1016/j.heliyon.2023.e15952]

-
Ma, S.; Wang, Q.; Wang, H.; Yang, Q.; Li, C.; Yu, Y.; Xie, Y.; Shi, X.; Wang, S. J. Ethnopharmacol. 2024, 337, 118916.
[https://doi.org/10.1016/j.jep.2024.118916]

-
Wang, M.; Carver, J. J.; Phelan, V. V.; Sanchez, L. M.; Garg, N.; Peng, Y.; Nguyen, D. D.; Watrous, J.; Kapono, C. A.; Luzzatto-Knaan, T.; Porto, C.; Bouslimani, A.; Melnik, A. V.; Meehan, M. J.; Liu, W.-T.; Crüsemann, M.; Boudreau, P. D.; Esquenazi, E.; Sandoval-Calderón, M.; Kersten, R. D.; Pace, L. A.; Quinn, R. A.; Duncan, K. R.; Hsu, C.-C.; Floros, D. J.; Gavilan, R. G.; Kleigrewe, K.; Northen, T.; Dutton, R. J.; Parrot, D.; Carlson, E. E.; Aigle, B.; Michelsen, C. F.; Jelsbak, L.; Sohlenkamp, C.; Pevzner, P.; Edlund, A.; McLean, J.; Piel, J.; Murphy, B. T.; Gerwick, L.; Liaw, C.-C.; Yang, Y.-L.; Humpf, H.-U.; Maansson, M.; Keyzers, R. A.; Sims, A. C.; Johnson, A. R.; Sidebottom, A. M.; Sedio, B. E.; Klitgaard, A.; Larson, C. B.; Boya P, C. A.; Torres-Mendoza, D.; Gonzalez, D. J.; Silva, D. B.; Marques, L. M.; Demarque, D. P.; Pociute, E.; O'Neill, E. C.; Briand, E.; Helfrich, E. J. N.; Granatosky, E. A.; Glukhov, E.; Ryffel, F.; Houson, H.; Mohimani, H.; Kharbush, J. J.; Zeng, Y.; Vorholt, J. A.; Kurita, K. L.; Charusanti, P.; McPhail, K. L.; Nielsen, K. F.; Vuong, L.; Elfeki, M.; Traxler, M. F.; Engene, N.; Koyama, N.; Vining, O. B.; Baric, R.; Silva, R. R.; Mascuch, S. J.; Tomasi, S.; Jenkins, S.; Macherla, V.; Hoffman, T.; Agarwal, V.; Williams, P. G.; Dai, J.; Neupane, R.; Gurr, J.; Rodríguez, A. M. C.; Lamsa, A.; Zhang, C.; Dorrestein, K.; Duggan, B. M.; Almaliti, J.; Allard, P.-M.; Phapale, P.; Nothias, L.-F.; Alexandrov, T.; Litaudon, M.; Wolfender, J.-L.; Kyle, J. E.; Metz, T. O.; Peryea, T.; Nguyen, D.-T.; VanLeer, D.; Shinn, P.; Jadhav, A.; Müller, R.; Waters, K. M.; Shi, W.; Liu, X.; Zhang, L.; Knight, R.; Jensen, P. R.; Palsson, B. Ø.; Pogliano, K.; Linington, R. G.; Gutiérrez, M.; Lopes, N. P.; Gerwick, W. H.; Moore, B. S.; Dorrestein, P. C.; Bandeira, N. Nat. Biotechnol. 2016, 34, 828–837.
[https://doi.org/10.1038/nbt.3597]

-
Kwon, H.; Kim, J. G.; Oh, J.-J.; Kim, J.-J.; Kim, G.-H.; Hwang, B. Y.; Yim, J. H.; Lee, D. Nat. Prod. Sci. 2020, 26, 340–344.
[https://doi.org/10.20307/nps.2020.26.4.340]

-
Han, J. S.; Kim, J. G.; Le, T. P. L.; Cho, Y. B.; Lee, D.; Hong, J. T.; Lee, M. K.; Hwang, B. Y. Phytochemistry 2023, 206, 113557.
[https://doi.org/10.1016/j.phytochem.2022.113557]

-
Guo, D.-L.; Huang, L.; Zhang, H.-M.; Mu, Y.-T.; Lei, H.-R.; Zhao, L.-L.; Hu, S.; Hu, Y.-J.; Li, C.-C.; Liu, M.-D.; Gu, Y.-C.; Yang, G.-K.; Dong, W.-Z.; Wang, D.; Deng, Y. Phytochemistry 2026, 244, 114747.
[https://doi.org/10.1016/j.phytochem.2025.114747]

-
Ni, J.-C.; Shi, J.-T.; Tan, Q.-W.; Chen, Q.-J. Nat. Prod. Res. 2019, 33, 101–107.
[https://doi.org/10.1080/14786419.2018.1437434]

-
Kayser, O.; Kolodziej, H. Phytochemistry 1995, 39, 1181–1185.
[https://doi.org/10.1016/0031-9422(95)00166-5]

-
Lv, H.-W.; Wang, Q.-L.; Luo, M.; Zhu, M.-D.; Liang, H.-M.; Li, W.-J.; Cai, H.; Zhou, Z.-B.; Wang, H.; Tong, S.-Q.; Li, X.-N. Arch. Pharm. Res. 2023, 46, 207–272.
[https://doi.org/10.1007/s12272-023-01443-4]

-
Nguyen, P.-H.; Zhao, B. T.; Kim, O.; Lee, J. H.; Choi, J. S.; Min, B. S.; Woo, M. H. J. Nat. Med. 2016, 70, 276–281.
[https://doi.org/10.1007/s11418-015-0957-x]