A New Methoxyflavonoid Rutinoside and PTP1B-Inhibitory Phenolic Compounds from the Water Extract of Areca catechu L.
Abstract
Phytochemical investigation of the water extract and alkaloid fraction of Areca catechu L. led to the isolation of one new methoxyflavonoid rutinoside (1), together with eleven known compounds (2−12), five of which (8−12) belong to the pyridine alkaloid class. The structure of the new compound was elucidated through extensive spectroscopic analyses, including 1D and 2D NMR experiments and high-resolution mass spectrometry. Notably, rhamnazin-3-O-rutinoside (2) is reported here for the first time from this species. The isolated compounds were evaluated for their inhibitory activity against protein tyrosine phosphatase 1B (PTP1B) using a p-nitrophenyl phosphate assay. Flavonoids (3 and 4) and stilbene (5) exhibited notable inhibitory activity, with IC50 values of 13.30 ± 0.71, 13.44 ± 0.84, and 8.68 ± 0.12 μM, respectively. In contrast, methoxyflavonoid rutinosides (1 and 2) and pyridine alkaloids (8−12) did not show significant inhibitory activity against PTP1B. These findings provide additional chemical insights into A. catechu and its constituents in relation to PTP1B inhibition.
Keywords:
Areca catechu, Arecaceae, Areca nut, Methoxyflavonoid, Rutinoside, PTP1BIntroduction
Areca catechu L. (Arecaceae) is a palm species widely cultivated throughout tropical Asia and traditionally used in several Asian medical systems, including traditional Chinese medicine, Ayurveda, and Southeast Asian folk medicine.1,2 Within these systems, the seeds and pericarp of A. catechu have been used to treat parasitic infections, digestive disorders (such as dyspepsia, diarrhea, dysentery, constipation, and abdominal distension), liver-related ailments (including jaundice and other hepatic disorders), as well as abdominal pain and edema.1 Phytochemical investigations of A. catechu have revealed the presence of multiple classes of secondary metabolites, such as alkaloids, flavonoids, tannins, phenolic compounds, triterpenoids, sterols, fatty acids, and polysaccharides.1,2 Extracts, fractions, and isolated constituents of A. catechu have been reported to exhibit a wide range of biological activities, including antioxidant, antimicrobial, cytotoxic, anti-inflammatory, analgesic, antinociceptive, hypoglycemic, lipid-modulating, antiparasitic, immunomodulatory, neuroprotective, digestive, antiviral, cardiovascular, and antiallergic effects.1 Based on the diverse biological activity potential reported for the extracts and isolated compounds from A. catechu, and as part of our ongoing research on medicinal plant-derived natural products,3–5 further investigation of its chemical constituents was undertaken. As a result, one new methoxyflavonoid rutinoside and eleven known compounds were isolated and structurally elucidated from this species. Furthermore, the isolated compounds were evaluated for their inhibitory activity against protein tyrosine phosphatase 1B (PTP1B).
Experimental
General experimental procedures − High-resolution electrospray ionization mass spectrometry (HRESIMS) was measured using a Micromass QTOF2-MS spectrometer. An Autobio PHOmo reader was used for spectrophotometric measurement in the PTP1B inhibitory assay. NMR spectra were recorded using a JEOL ECZ600R/S1 600 MHz spectrometer (JEOL, Tokyo, Japan) and a Bruker Avance Digital 500 MHz NMR spectrometer (Bruker, Karlsruhe, Germany) with tetramethylsilane as the internal standard. Open column chromatography was implemented with silica gel 60 (Merck, 230−400 mesh) and reversed-phase (RP)-C18 silica gel (Merck, 75 mesh). Thin-layer chromatography (TLC) was carried out using Merck precoated silica gel F254 plates and RP-18 F254s plates. Preparative high-performance liquid chromatography (HPLC) was conducted utilizing a Gilson HPLC system; pump: model 306; detector: 156 UV/VIS detector, and a YMC-Pack ODS-A column (25 250 mm, 5 μm particle size, YMC Co. Ltd, Japan).
Chemicals and Reagents − Protein tyrosine phosphatase 1B (PTP1B, human recombinant) and dithiothreitol (DTT) were purchased from NKMAX Co., Ltd. (Seongnam, Korea) and Bio-Rad Laboratories (Hercules, CA, USA), respectively. Reagents utilized in the enzyme inhibitory assay were procured from Sigma-Aldrich Co. (St. Louis, MO, U.S.A.), including p-nitrophenyl phosphate (p-NPP), ethylene diamine tetraacetic acid (EDTA), sodium citrate, sodium hydroxide, ursolic acid, and dimethyl sulfoxide (DMSO). The organic solvents were purchased from Duksan Pharmaceutical Co. (Seoul, Korea).
Plant material − The seeds of A. catechu were purchased from the Traditional Market, Wansan-gu, Jeonju-si, Jeonbuk State, Republic of Korea, in May 2023, and authenticated by one of the authors, Professor Byung Sun Min. The voucher specimen (CUD-3183) was stored at the Laboratory of Pharmacognosy in the College of Pharmacy, Daegu Catholic University, Republic of Korea.
Extraction and isolation − The dried nut of A. catechu (part I, 10.5 kg) was crushed and then extracted three times (3 h × 10 L) with methanol. The extract was concentrated under reduced pressure by rotary evaporation to give a crude extract. The crude extract (1.3 kg) was suspended in distilled water (2 L), and the pH was adjusted to 2 with 2% HCl solution, then extracted with CH2Cl2 (AC1, 5 × 5 L). The acidic aqueous layer was adjusted to alkaline pH (pH 10) with ammonia solution and then extracted with CH2Cl2 (AC2, 5 × 4 L). The CH2Cl2 extract (AC2) was concentrated to produce the crude alkaloid extract (ACA). ACA (37 g) was chromatographed on a silica gel column using a stepwise gradient of CH2Cl2-MeOH (50:1 to 0:1, v/v) to yield ten fractions (C1–C10) based on TLC analysis. Fraction C3 (3.5 g) was subjected to column chromatography (CC) on silica gel eluted with a solvent system of CH2Cl2-EtOAc (40:1, v/v) to yield 8 (350 mg). Fraction C5 (1.1 g) was subjected to CC on silica gel (CH2Cl2-EtOAc, 20:1, v/v) to give five fractions (C5.1–C5.5). Compound 11 (5 mg) was obtained from fraction C5.4 (157 mg) by CC on silica gel (CH2Cl2-acetone, 15:1, v/v). Fraction C7 (3.6 g) was chromatographed on a silica gel column using a stepwise gradient of CH2Cl2-MeOH (30:1 to 0:1, v/v) to yield six fractions (C7.1–C7.6). Fraction C7.5 (1.6 g) was further purified on a silica gel column (n-hexane-CH2Cl2-EtOAc, 5:1:1, v/v) to give 10 (10.0 mg), and 9 (100 mg). Compound 12 (4 mg) was obtained from fraction C7.6 (225 mg) by RP-C18 CC (MeOH-H2O, 2:1, v/v).
The dried nuts of A. catechu (part II, 5 kg) were extracted with water (10 L) three times, each extraction lasting 4 h. The combined aqueous extracts were filtered and concentrated under reduced pressure until the volume was reduced to approximately 2 L, which was then successively partitioned with methylene chloride and ethyl acetate, to afford the methylene chloride (4.6 g) and ethyl acetate (92.8 g) fractions, together with the remaining aqueous layer (ACW). The ethyl acetate (92.8 g) fraction was chromatographed by silica gel CC eluted with CH2Cl2:MeOH solvent system (50:1 → 100% MeOH, stepwise) to afford 11 fractions (ACE1–ACE11). Fraction ACE7 (180 mg) was separated by RP-C18 CC (MeOH-H2O, 1:1, v/v) to yield compounds 3 (4 mg) and 4 (5 mg). Fraction ACE5 (1.6 g) was chromatographed on a silica gel column (CH2Cl2-MeOH, 20:1, v/v) to yield six subfractions (ACE5.1-ACE5.6). Sub-fraction ACE5.4 (120 mg) was separated using an RP-C18 column (MeOH-H2O, 1.5:1, v/v) to give compounds 7 (5 mg), 5 (5 mg), and 6 (7 mg). The H2O fraction (ACW) was separated using a Diaion HP-20 column, eluting with a gradient solvent mixture of MeOH-H2O (gradient 25:75, 50:50, 75:25, and 100:0, v/v, stepwise) to yield four fractions (ACW-1 to ACW-4), based on TLC analysis. Fraction ACW3 (1.2 g) was chromatographed on a silica gel column (CH2Cl2-MeOH, 8:1, v/v) to yield five subfractions (ACW3.1–ACW3.5). Fraction ACW3.4 (112 mg) was purified using a preparative Gilson HPLC system (isocratic eluent, 55% MeOH in H2O + 0.1% TFA, 5 mL/min, 60 min) to yield compounds 1 (2 mg) and 2 (3 mg).
5′-O-Methylrhamnazin-3-O-rutinoside (1) − Yellow amorphous powder. 1H-NMR (600 MHz, CD3OD): δ 7.61 (2H, s, H-2′, H-6′), 6.67 (1H, d, J = 2.4 Hz, H-8), 6.36 (1H, d, J = 2.4 Hz, H-6), 5.31 (1H, d, J = 7.8 Hz, H-1′′), 4.55 (1H, d, J = 1.8 Hz, H-1′′′), 3.95 (6H, s, 3′-OCH3, 5′-OCH3), 3.91 (3H, s, 7-OCH3), 3.83 (1H, dd, J = 11.4, 1.8 Hz, H-6′′a), 3.60 (1H, m, H-3′′′), 3.50 (1H, m, H-2′′), 3.47 (1H, m, H-5′′), 3.46 (1H, m, H-6′′b), 3.45 (1H, m, H-2′′′), 3.40 (1H, m, H-5′′′), 3.39 (1H, m, H-3′′), 3.26 (1H, m, H-4′′), 3.24 (1H, m, H-4′′′), 1.10 (3H, d, J = 6.0 Hz, H-6′′′); 13C-NMR (150 MHz, CD3OD): δ 179.3 (C-4), 167.4 (C-7), 162.7 (C-5), 159.2 (C-2), 158.3 (C-8a), 149.0 (C-3′, C-5′), 141.8 (C-4′), 135.6 (C-3), 120.8 (C-1′), 108.5 (C-2′, C-6′), 106.5 (C-4a), 104.2 (C-1′′), 102.6 (C-1′′′), 99.2 (C-6), 93.3 (C-8), 78.1 (C-5′′), 77.4 (C-3′′), 75.9 (C-2′′), 73.8 (C-4′′′), 72.2 (C-2′′′), 72.1 (C-3′′′), 71.7 (C-4′′), 69.8 (C-5′′′), 68.5 (C-6′′), 57.1 (3′-OCH3, 5′-OCH3), 56.5 (7-OCH3), 17.8 (C-6′′′); HR-ESI-MS m/z 691.1852 [M + Na]+ (calcd for C30H36O17Na, 691.1850) and 669.2031 [M + H]+ (calcd for C30H37O17, 669.2031).
Rhamnazin-3-O-rutinoside (2) − Yellow amorphous powder. 1H-NMR (500 MHz, CD3OD): δ 7.95 (1H, d, J = 2.0 Hz, H-2′), 7.67 (1H, dd, J = 8.5, 2.0 Hz, H-6′), 6.91 (1H, d, J = 8.5 Hz, H-5′), 6.60 (1H, d, J = 2.0 Hz, H-8), 6.33 (1H, d, J = 2.0 Hz, H-6), 5.26 (1H, d, J = 7.5 Hz, H-1′′), 4.52 (1H, d, J = 2.0 Hz, H-1′′′), 3.94 (3H, s, 3′-OCH3), 3.88 (3H, s, 7-OCH3), 3.82 (1H, m, H-6′′a), 3.60 (1H, m, H-3′′′), 3.48 (1H, m, H-2′′), 3.47 (1H, m, H-5′′), 3.45 (1H, m, H-6′′b), 3.45 (1H, m, H-2′′′), 3.40 (1H, m, H-5′′′), 3.39 (1H, m, H-3′′), 3.25 (1H, m, H-4′′), 3.24 (1H, m, H-4′′′), 1.09 (3H, d, J = 6.0 Hz, H-6′′′); 13C-NMR (125 MHz, CD3OD): δ 179.4 (C-4), 167.4 (C-7), 162.7 (C-5), 159.2 (C-2), 158.4 (C-8a), 151.1 (C-4′), 148.3 (C-3′), 135.6 (C-3), 124.1 (C-6′), 122.7 (C-1′), 116.1 (C-5′), 114.5 (C-2′), 106.5 (C-4a), 104.2 (C-1′′), 102.5 (C-1′′′), 99.2 (C-6), 93.2 (C-8), 78.1 (C-5′′), 77.3 (C-3′′), 75.9 (C-2′′), 73.8 (C-4′′′), 72.2 (C-2′′′), 72.1 (C-3′′′), 71.6 (C-4′′), 69.7 (C-5′′′), 68.5 (C-6′′), 56.7 (7-OCH3), 56.5 (3′-OCH3), 17.9 (C-6′′′); HR-ESI-MS m/z 661.1744 [M + Na]+ (calcd for C29H34O16Na, 661.1745) and 639.1924 [M + H]+ (calcd for C29H35O16, 639.1925).
Arecoline (8) − Oil. 1H-NMR (500 MHz, CDCl3): δ 6.91 (1H, m, H-4), 3.65 (3H, s, 7-OCH3), 3.07 (2H, m, H-2), 2.43 (2H, t, J = 5.5 Hz, H-6), 2.33 (3H, s, 1-NCH3), 2.29 (2H, m, H-5); 13C-NMR (125 MHz, CDCl3): δ 166.1 (CO-7), 137.5 (C-4), 128.7 (C-3), 53.1 (C-2), 51.5 (7-OCH3), 50.7 (C-6), 45.6 (1-NCH3), 26.4 (C-5).
Arecaidine (9) − White amorphous powder. 1H-NMR (500 MHz, CD3OD): δ 6.74 (1H, m, H-4), 3.79 (2H, m, H-2), 3.21 (2H, t, J = 6.0 Hz, H-6), 2.88 (3H, s, 1-NCH3), 2.54 (2H, m, H-5); 13C-NMR (125 MHz, CD3OD): δ 171.3 (CO-7), 131.7 (C-4), 131.2 (C-3), 53.7 (C-2), 50.8 (C-6), 43.3 (1-NCH3), 23.9 (C-5).
Guavacine (10) − White amorphous powder. 1H-NMR (500 MHz, D2O): δ 6.83 (1H, m, H-4), 3.86 (2H, m, H-2), 3.32 (2H, t, J = 6.0 Hz, H-6), 2.56 (2H, m, H-5); 13C-NMR (125 MHz, D2O): δ 171.4 (CO-7), 131.4 (C-4), 127.6 (C-3), 41.1 (C-2), 38.9 (C-6), 20.6 (C-5).
Guvacoline (11) − White amorphous powder. 1H-NMR (500 MHz, D2O): δ 7.22 (1H, m, H-4), 3.93 (2H, m, H-2), 3.80 (3H, s, 7-OCH3), 3.35 (2H, t, J = 6.0 Hz, H-6), 2.64 (2H, m, H-5); 13C-NMR (125 MHz, D2O): δ 178.4 (CO-7), 138.6 (C-4), 123.4 (C-3), 52.6 (7-OCH3), 40.6 (C-2), 39.6 (C-6), 21.8 (C-5).
Arecatemine C (12) − Yellow amorphous powder. 1H-NMR (500 MHz, CD3OD/CDCl3): δ 7.30 (1H, m, H-4), 4.32 (1H, d, J = 16.0 Hz, H-2a), 3.92 (3H, s, 7-OCH3), 3.53 (2H, m, H-6), 3.41 (3H, s, 1-NCH3), 3.19 (1H, d, J = 16.0 Hz, H-2b), 3.04 (1H, m, H-5a), 2.67 (1H, m, H-5b); 13C-NMR (125 MHz, CD3OD/CDCl3): δ 165.5 (CO-7), 137.2 (C-4), 124.7 (C-3), 65.5 (C-2), 62.2 (C-6), 58.6 (1-NCH3), 52.5 (7-OCH3), 23.7 (C-5).
Acid hydrolysis of 1 − The acid hydrolysis of 1 was carried out with some modifications of our previously described method.4,6 Compound 1 (1 mg) was dissolved in 3% HCl (5 mL), and the solutions were stirred at 80°C for 4 h. After cooling, the reaction mixture was extracted with CH2Cl2. The aqueous layer was repeatedly evaporated in vacuo until neutral to yield the sugar residues, which were then subjected to TLC analysis (CH2Cl2:MeOH:H2O, 2:1:0.2, v/v/v). The spots were visualized by spraying with 95% EtOH-H2SO4-anisaldehyde (9/0.5/0.5, v/v), then heated at 200°C for 5 min. The monosaccharide residues, glucose and rhamnose, were identified by their TLC behavior through direct comparison with authentic standards.
PTP1B inhibitory assay − The inhibition assay against recombinant human PTP1B enzyme was conducted using p-nitrophenyl phosphate (p-NPP) as the substrate, according to our previously published protocols.3,7 The inhibitory activity of PTP1B was evaluated via an in vitro assay conducted on a 96-well plate. Samples were prepared at varying concentrations (1–100 μM) by diluting them in a reaction buffer containing 50 mM citrate buffer (pH 6.0, 0.1 M NaCl, 1 mM EDTA, and 1 mM DTT). Next, 10 μL of the test samples were added to each well, followed by 30–40 μL of reaction buffer and 10 μL of PTP1B enzyme (5 μg/mL). The mixture was incubated at 37°C for 10 minutes before adding 50 μL of 2.0 mM p-NPP substrate. Then, the reaction was mixed gently and incubated again at 37°C for 20 minutes. After 20 minutes, 10 µL of 10 M NaOH was added to stop the reaction. Under alkaline conditions, the dephosphorylation of p-NPP produced the yellow p-nitrophenolate anion, allowing spectrophotometric determination of PTP1B activity, which could be quantified at 405 nm. The percent inhibition (%) was obtained by the following equation:
where Ac is the absorbance of the control, and As is the absorbance in the presence of the test sample. Ursolic acid was used as a positive control in this experiment.
Statistical analysis − Statistical analyses were performed by one-way analysis of variance (ANOVA), followed by the Fisher least significant difference test. p<0.05 was considered statistically significant. The results are presented as the mean±standard error of the mean (SEM).
Results and Discussion
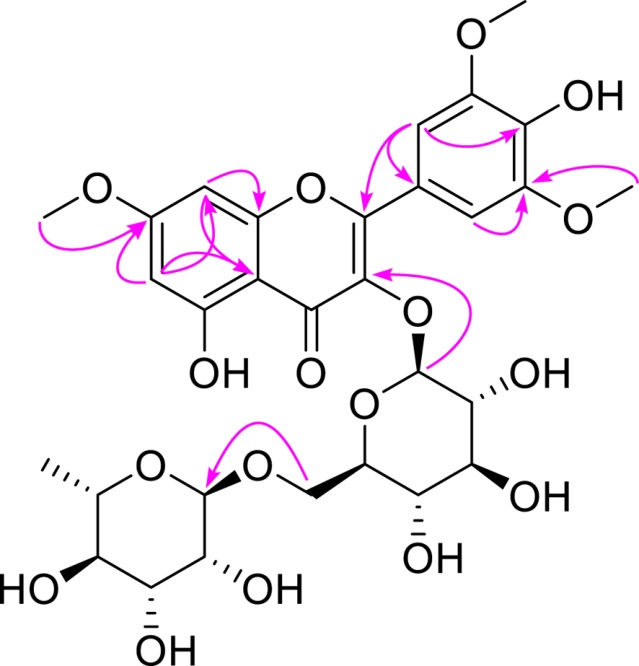
Compound 1 was isolated as a yellow, amorphous powder. Its molecular formula was determined as C30H36O17 based on the HRESIMS data, which showed a sodium adduct ion at m/z 691.1852 ([M + Na]+, calcd for C30H36O17Na, 691.1850) and protonated molecular ion at m/z 669.2031 ([M + H]+, calcd for C30H37O17, 669.2031). The 1H and 13C NMR spectra of 1 displayed characteristic signals of a flavonoid glycoside including one carbonyl carbon at δC 179.3 (C-4); an AX coupling system at δH/δC 6.67 (1H, d, J = 2.4 Hz, H-8)/93.3 and 6.36 (1H, d, J = 2.4 Hz, H-6)/99.2; a symmetrically substituted B ring, as evidenced by two equivalent aromatic proton signals at δH 7.61 (2H, s, H-2′, H-6′) together with the corresponding carbon resonance at δC 108.5; and two anomeric proton signals at δH 5.31 (1H, d, J = 7.8 Hz, H-1′′) and 4.55 (1H, d, J = 1.8 Hz, H-1′′′). In addition, three methoxy groups were observed at δH/δC 3.95 (6H, s, 3′, 5′-OCH3)/57.1 and 3.91 (3H, s, 7-OCH3)/56.5, and their positions were established by the HMBC correlations (Fig. 2). The presence of a rhamnopyranosyl unit was indicated by the characteristic methyl signal at δH/δC 1.10 (3H, d, J = 6.0 Hz, H-6′′′)/17.8 and the small coupling constant of its anomeric proton at δH 4.55 (1H, d, J = 1.8 Hz, H-1′′′)/ 102.6. In contrast, the large coupling constant of the remaining anomeric proton at δH 5.31 (1H, d, J = 7.8 Hz, H-1′′) together with a set of six sugar carbon signals at δC 104.2 (C-1′′), 78.1 (C-5′′), 77.4 (C-3′′), 75.9 (C-2′′), 71.7 (C-4′′), and 68.5 (C-6′′), suggested the presence of a β-glucopyranosyl unit. The 1→6 linkage between the rhamnopyranosyl and glucopyranosyl units, forming a rutinoside moiety, was established by the HMBC correlation from the methylene protons at δH 3.83 and 3.46 (H2-6′′) to the anomeric carbon at δC 102.6 (C-1′′′), together with the downfield shift of C-6′′ (δC 68.5) in the 13C NMR spectrum. The rutinoside moiety of 1 was determined to be attached at C-3 of the aglycone as demonstrated by a key HMBC correlation between H-1′′ (δH 5.31) and C-3 (δC 135.6) (Fig. 2). Detailed analysis of the 1D and 2D NMR data revealed that compound 1 is closely related to 2 (rhamnazin-3-O-rutinoside), differing only by the presence of an additional methoxy group at C-5′. Acid hydrolysis of compound 1 afforded glucose and rhamnose, which were identified by TLC comparison with authentic standards. Based on biosynthetic considerations and comparison with reported data, the sugar units were tentatively suggested as D-glucose and L-rhamnose. Accordingly, compound 1 was identified as 5′-O-methylrhamnazin-3-O-rutinoside, representing a new methoxyflavonoid rutinoside.
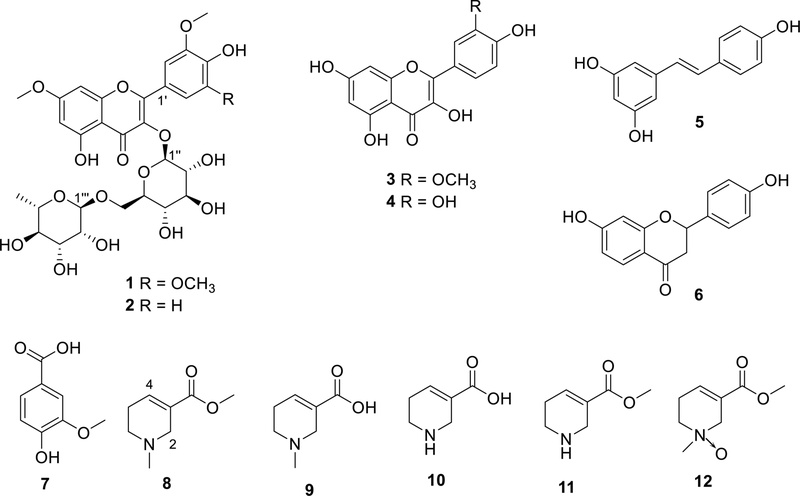
In addition, the known compounds were characterized through spectroscopic analysis and confirmed by comparison with published reference data. The chemical structures of the eleven known compounds were determined as follows: rhamnazin-3-O-rutinoside (2),8 isorhamnetin (3′-O-methylquercetin, 3),9 quercetin (4),10 resveratrol (3,5,4′-trihydroxystilbene, 5),11 naringenin (5,7,4′-trihydroxyflavanone, 6),9 4-hydroxy-3-methoxybenzoic acid (vanillic acid, 7),12 arecoline (8),2,13 arecaidine (9),2,13 guavacoline (10),13 guavacine (11),2,13 and arecatemine C (12)2 (Fig. 1).
The isolated compounds (1−12) were evaluated for their inhibitory activity against PTP1B enzyme using the p-nitrophenyl phosphate assay. As shown in Table 1, compounds 3−5 exhibited notable inhibitory effects with IC50 values of 13.30 ± 0.71, 13.44 ± 0.84, and 8.68 ± 0.12 μM, respectively. Among them, compound 5 showed the most potent activity, which was comparable to the positive control, ursolic acid (IC50 = 9.12 ± 0.34 μM). In contrast, compounds 1−2 and 6−12 did not exhibit significant inhibitory activity (IC50 > 100 μM) under the tested conditions. Pyridine alkaloids are well-known characteristic constituents of A. catechu and have been reported to possess various biological activities.1 However, compounds 8−12 did not exhibit significant inhibitory activity against PTP1B (IC50 > 100 μM) in the present study. This result suggests that the pyridine alkaloid scaffold may not be favorable for PTP1B inhibition.
In summary, phytochemical investigation of A. catechu led to the isolation and structural elucidation of one new methoxyflavonoid rutinoside (1) together with eleven known compounds (2−12). The new compound was characterized by comprehensive spectroscopic analyses, and rhamnazin-3-O-rutinoside was identified for the first time from this species. Biological evaluation demonstrated that the flavonoid (3 and 4) and stilbene (5) derivatives exhibited notable PTP1B inhibitory activity, whereas the methoxyflavonoid rutinosides (1 and 2) and pyridine alkaloids (8−12) showed no significant activity. These findings broaden the chemical knowledge of this species and provide insight into constituents that may contribute to its PTP1B inhibitory potential.
Acknowledgments
This research was supported by the Korea Ministry of Food and Drug Safety (23212KFDA204) and Gyeongsangbuk-do RISE (Regional Innovation System & Education; 2025-RISE-15-107). We thank the Korea Basic Science Institute (KBSI) for mass spectrometric measurements.
Conflicts of Interest
The authors have declared that there are no conflicts of interest.
References
-
Ha, M. T.; Tran, T. H.; Huh, J.; Min, S. E.; Han, K.-H.; Kim, Y.-B.; Kim, J. A.; Min, B. S. Nat. Prod. Sci. 2025, 31, 236–250.
[https://doi.org/10.20307/nps.2025.31.4.236]

-
Tang, S.-N.; Zhang, J.; Liu, D.; Liu, Z.-W.; Zhang, X.-Q.; Ye, W.-C. J. Asian Nat. Prod. Res. 2017, 19, 1155–21159.
[https://doi.org/10.1080/10286020.2017.1307187]

-
Ha, M. T.; Le, T. T.; Tran, T. T.; Tran, T. H.; Kim, J. A.; Min, B. S. Nat. Prod. Sci. 2025, 31, 129–137.
[https://doi.org/10.20307/nps.2025.31.2.129]

-
Le, T. T.; Ha, M. T.; Lee, G. S.; Nguyen, V. P.; Kim, C. S.; Kim, J. A.; Min, B. S. Phytochemistry 2025, 229, 114269.
[https://doi.org/10.1016/j.phytochem.2024.114269]

-
Ha, M. T.; Le, T. T.; Nguyen, V. T.; Kim, J. A.; Choi, J. S.; Min, B. S. Phytochem. Lett. 2023, 57, 231–238.
[https://doi.org/10.1016/j.phytol.2023.08.020]

-
Ha, M. T.; Gal, M.; Kim, J. A.; Lee, J.-H.; Min, B. S. Bioorg. Chem. 2024, 143, 107066.
[https://doi.org/10.1016/j.bioorg.2023.107066]

-
Pham, T. L.; Ha, M. T.; Min, B. S.; Kim, J. A. Nat. Prod. Sci. 2025, 31, 62–73.
[https://doi.org/10.20307/nps.2025.31.1.62]

- Harput, Ü. Ş.; Saracoğlu, I.; Ogihara, Y. Turk. J. Chem. 2004, 28, 761–766.
-
Kulić, Ž.; Steiner, V. J. N.; Butterer, A. Planta Med. 2026, 92, 58–80.
[https://doi.org/10.1055/a-2706-7473]

-
To, D. C.; Nguyen, H. T.; Hoa, T. T. V.; Thuy, N. T. T.; Tran, M. H.; Nguyen, P. H.; Nguyen, P. D. N.; Nhan, N. T.; Tram, N. T. T. Data Brief. 2023, 51, 109713.
[https://doi.org/10.1016/j.dib.2023.109713]

-
Tisserant, L.-P.; Hubert, J.; Lequart, M.; Borie, N.; Maurin, N.; Pilard, S.; Jeandet, P.; Aziz, A.; Renault, J.-H.; Nuzillard, J.-M. J. Nat. Prod. 2016, 79, 2846–2855.
[https://doi.org/10.1021/acs.jnatprod.6b00608]

- Akbar, A.; Ali, M. S.; Zikr-Ur-Rehman, S.; Lateef, M.; Saeed, Z. Int. J. Biol. Biotech. 2020, 17, 9–15.
-
Huang, J. L.; McLeish, M. J. J. Chromatogr. A 1989, 475, 447–450.
[https://doi.org/10.1016/S0021-9673(01)89702-8]